Abstract
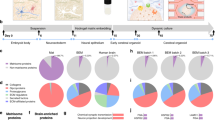
Three-dimensional cell culture models have either relied on the self-organizing properties of mammalian cells1,2,3,4,5,6 or used bioengineered constructs to arrange cells in an organ-like configuration7,8. While self-organizing organoids excel at recapitulating early developmental events, bioengineered constructs reproducibly generate desired tissue architectures. Here, we combine these two approaches to reproducibly generate human forebrain tissue while maintaining its self-organizing capacity. We use poly(lactide-co-glycolide) copolymer (PLGA) fiber microfilaments as a floating scaffold to generate elongated embryoid bodies. Microfilament-engineered cerebral organoids (enCORs) display enhanced neuroectoderm formation and improved cortical development. Furthermore, reconstitution of the basement membrane leads to characteristic cortical tissue architecture, including formation of a polarized cortical plate and radial units. Thus, enCORs model the distinctive radial organization of the cerebral cortex and allow for the study of neuronal migration. Our data demonstrate that combining 3D cell culture with bioengineering can increase reproducibility and improve tissue architecture.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
14 August 2018
In the HTML and PDF versions of this article initially published, μ's appeared as m's throughout the scale bar lengths in the figure legends, as well as in the Methods section in the phrases "added to give a final volume of 150 μl per well," "an average hPSC cell size of 15 μm" and "sectioned on a vibratome to collect 300-μm sections." In the HTML version, μ's appeared as m's in all instances throughout the Methods section except "35 μg/ml of pure laminin" and "35 μl laminin/entactin + 25 μl collagen." The errors have been corrected in the HTML and PDF versions of the article.
21 September 2018
In the PDF version of this article published on 14 August 2018, Figure 2 was replaced by part of Figure 3. The error has been corrected online.
01 October 2018
Nat. Biotechnol. 35, 659–666 (2017); published online 31 May 2017; corrected after print 14 August 2018 In the HTML and PDF versions of this article initially published, μ's appeared as m's throughout the scale bar lengths in the figure legends, as well as in the Methods section in the phrases “added to give a final volume of 150 μl per well,” “an average hPSC cell size of 15 μm” and “sectioned on a vibratome to collect 300-μm sections.
References
Rookmaaker, M.B., Schutgens, F., Verhaar, M.C. & Clevers, H. Development and application of human adult stem or progenitor cell organoids. Nat. Rev. Nephrol. 11, 546–554 (2015).
Fatehullah, A., Tan, S.H. & Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 18, 246–254 (2016).
Yin, X. et al. Engineering stem cell organoids. Cell Stem Cell 18, 25–38 (2016).
Dedhia, P.H., Bertaux-Skeirik, N., Zavros, Y. & Spence, J.R. Organoid models of human gastrointestinal development and disease. Gastroenterology 150, 1098–1112 (2016).
Sasai, Y. Next-generation regenerative medicine: organogenesis from stem cells in 3D culture. Cell Stem Cell 12, 520–530 (2013).
Lancaster, M.A. & Knoblich, J.A. Organogenesis in a dish: modeling development and disease using organoid technologies. Science 345, 1247125–1247125 (2014).
Langer, R. & Vacanti, J. Advances in tissue engineering. J. Pediatr. Surg. 51, 8–12 (2016).
Esch, E.W., Bahinski, A. & Huh, D. Organs-on-chips at the frontiers of drug discovery. Nat. Rev. Drug Discov. 14, 248–260 (2015).
Xinaris, C., Brizi, V. & Remuzzi, G. Organoid models and applications in biomedical research. Nephron 130, 191–199 (2015).
Hynds, R.E. & Giangreco, A. Concise review: the relevance of human stem cell-derived organoid models for epithelial translational medicine. Stem Cells 31, 417–422 (2013).
Lancaster, M.A. & Knoblich, J.A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340 (2014).
Eiraku, M. et al. Self-organized formation of polarized cortical tissues from ESCs and its active manipulation by extrinsic signals. Cell Stem Cell 3, 519–532 (2008).
Mariani, J. et al. Modeling human cortical development in vitro using induced pluripotent stem cells. Proc. Natl. Acad. Sci. USA 109, 12770–12775 (2012).
Lancaster, M.A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature 472, 51–56 (2011).
Caiazzo, M. et al. Defined three-dimensional microenvironments boost induction of pluripotency. Nat. Mater. 15, 344–352 (2016).
Warmflash, A., Sorre, B., Etoc, F., Siggia, E.D. & Brivanlou, A.H. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Oh, S.K.W. et al. Long-term microcarrier suspension cultures of human embryonic stem cells. Stem Cell Res. (Amst.) 2, 219–230 (2009).
Molander, H., Olsson, Y., Engkvist, O., Bowald, S. & Eriksson, I. Regeneration of peripheral nerve through a polyglactin tube. Muscle Nerve 5, 54–57 (1982).
Chenn, A. & Walsh, C.A. Regulation of cerebral cortical size by control of cell cycle exit in neural precursors. Science 297, 365–369 (2002).
Subramanian, L., Remedios, R., Shetty, A. & Tole, S. Signals from the edges: the cortical hem and antihem in telencephalic development. Semin. Cell Dev. Biol. 20, 712–718 (2009).
Miller, J.A. et al. Transcriptional landscape of the prenatal human brain. Nature 508, 199–206 (2014).
Rakic, P. Specification of cerebral cortical areas. Science 241, 170–176 (1988).
Jones, E.G. & Rakic, P. Radial columns in cortical architecture: it is the composition that counts. Cereb. Cortex 20, 2261–2264 (2010).
Geschwind, D.H. & Rakic, P. Cortical evolution: judge the brain by its cover. Neuron 80, 633–647 (2013).
Kadoshima, T. et al. Self-organization of axial polarity, inside-out layer pattern, and species-specific progenitor dynamics in human ES cell-derived neocortex. Proc. Natl. Acad. Sci. USA 110, 20284–20289 (2013).
Paşca, A.M. et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat. Methods 12, 671–678 (2015).
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Sievers, J., Pehlemann, F.W., Gude, S. & Berry, M. Meningeal cells organize the superficial glia limitans of the cerebellum and produce components of both the interstitial matrix and the basement membrane. J. Neurocytol. 23, 135–149 (1994).
Halfter, W., Dong, S., Yip, Y.-P., Willem, M. & Mayer, U. A critical function of the pial basement membrane in cortical histogenesis. J. Neurosci. 22, 6029–6040 (2002).
Rakic, P. Evolution of the neocortex: a perspective from developmental biology. Nat. Rev. Neurosci. 10, 724–735 (2009).
Wang, X., Tsai, J.-W., LaMonica, B. & Kriegstein, A.R. A new subtype of progenitor cell in the mouse embryonic neocortex. Nat. Neurosci. 14, 555–561 (2011).
Marín, O., Valiente, M., Ge, X. & Tsai, L.-H. Guiding neuronal cell migrations. Cold Spring Harb. Perspect. Biol. 2, a001834 (2010).
Otani, T., Marchetto, M.C., Gage, F.H., Simons, B.D. & Livesey, F.J. 2D and 3D stem cell models of primate cortical development identify species-specific differences in progenitor behavior contributing to brain size. Cell Stem Cell 18, 467–480 (2016).
Matsuda, T. & Cepko, C.L. Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc. Natl. Acad. Sci. USA 101, 16–22 (2004).
Villefranc, J.A., Amigo, J. & Lawson, N.D. Gateway compatible vectors for analysis of gene function in the zebrafish. Dev. Dyn. 236, 3077–3087 (2007).
Mátés, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Daza, R.A.M., Englund, C. & Hevner, R.F. Organotypic slice culture of embryonic brain tissue. CSH Protoc. 2007, t4914 (2007).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Roberts, A., Trapnell, C., Donaghey, J., Rinn, J.L. & Pachter, L. Improving RNA-Seq expression estimates by correcting for fragment bias. Genome Biol. 12, R22 (2011).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank members of the Lancaster laboratory for helpful discussion and technical support, especially S. Giandomenico and M. Sutcliffe, as well as A.G. Gianni for sea sponge sample. We also thank members of the Knoblich laboratory for insight and technical help, particularly A. Peer. T.O. was supported by the Cambridge Wellcome Trust PhD program in developmental biology, and F.J.L. is a Wellcome Trust Investigator. M.A.L. was funded by a Marie Curie Postdoctoral fellowship, and work in M.A.L.'s laboratory is supported by the Medical Research Council (MC_UP_1201/9). N.S.C. was funded by an EMBO long-term fellowship and a Deutsche Forschungsgemeinschaft research fellowship (DFG CO 1324/1-1). Work in J.A.K.'s laboratory is supported by the Austrian Academy of Sciences, the Austrian Science Fund (grants I_1281-B19 and Z_153_B09), and an advanced grant from the European Research Council. pCAGEN and pCAG-GFP were gifts from Connie Cepko, Harvard Medical School. pT2/HB was a gift from Perry Hackett, University of Minnesota. pENTR-EGFP2 was a gift from Nathan Lawson, University of Massachusetts Medical School. pCMV(CAT)T7-SB100 was a gift from Zsuzsanna Izsvak, Max Delbruck Center for Molecular Medicine.
Author information
Authors and Affiliations
Contributions
M.A.L. conceived the project, planned and performed experiments, and wrote the manuscript. N.S.C. performed experiments and analyzed data. S.W. and E.H.G. prepared samples and performed various treatments. A.W.P. performed cloning and prepared samples. T.R.B. performed bioinformatics analysis of RNA-seq data. T.O. prepared samples and performed experiments under the supervision of F.J.L. J.A.K. supervised the project, planned and interpreted experiments, and wrote the manuscript.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Corresponding authors
Ethics declarations
Competing interests
M.A.L. and J.A.K. have filed a patent application for use of this technology in future disease modeling and toxicology testing.
Integrated supplementary information
Supplementary Figure 1 Batch-to-batch variability in neuroectoderm formation and testing varying EB size
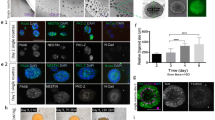
a. Bright field images of 6 spherical EBs made in 96-well U-bottom plate at day 3, from two independent batches, made from H9 cells, showing the homogeneity of EBs early on between EBs and between batches. b. Bright field images of seven independent batches of spherical organoids on the day of transfer to NI (day 10 for batches 4,6; day 11 for batches 1, 2, 3, 5; day 12 for batch 7) showing the degree of variability in generation of polarized neuroectoderm (arrows). c. Mean ratio of EBs per batch (same as in a.) displaying at least some neuroepithelium, recognizable in bright field as optically clear superficial tissue with a clean border and evidence of radial cellular architecture. d. Bright field images of various sized day 10 spherical H9 EBs made with the indicated cell numbers. Note the increase in optically clear neuroectodermal tissue (arrows) relative to the rest of the EB, but the decreased total amount of such tissue in the smaller EBs. e. Immunohistochemical staining of day 10 H9 EBs of varying sizes for the germ layer markers Brachyury for mesoderm, N-Cadherin for neural ectoderm, Sox17 for endoderm, and E-Cadherin for non-neural epithelium. Note the increasing prevalence of mesoderm and endoderm (arrows) with increasing cell number and EB size. Shown at right is quantification of fluorescence staining of the above markers (mean grey value relative to DAPI). Mean shown ± SD. n = 5-10 spheroids (10-day, H9) for each size. f. Bright field images of various sized day 17 spherical EBs showing a decrease in non-neuroepithelial tissues (arrowheads) compared to neuroepithelial buds (arrows). g. Additional bright field images of day 17 spherical EBs made from 100 and 500 cells. Note the increased "purity" of tissues containing primarily neuroepithelial buds (arrows). But note the decreased numbers and smaller sizes of such buds. Also shown is an example of an EB that failed to develop (arrowhead). Scale bars: 200 μm in a., 500 μm in c., d., f., 100 μm in e.. g.
Supplementary Figure 2 Bioengineered brain organoids display consistent generation of neural ectoderm
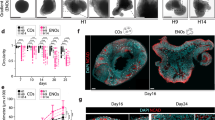
a. Bright field image of the intact braided PLGA fiber. b. Bright field image of isolated microfilaments. c. Hydrated microfilaments within a droplet of EB medium. d. Three examples of microfilament engineered H9 EBs at day 3 of the protocol. Note the elongated and sometimes complex arrangement. e. Bright field image of collagen-type fibers isolated from dried sea sponge used to generate EBs in Figure 1b. f. Bright field imaging of cellulose fibers used to generate EBs in Figure 1b. g. Bright field images of seven independent batches of microfilament organoids on the day of transfer to NI (day 10 for batches 4,6; day 11 for batches 1, 2, 3, 5; day 12 for batch 7), showing reduced variability in generation of polarized neuroectoderm (arrows). h. Mean ratio of EBs per enCOR batch (same as in g.) displaying at least some neuroepithelium, recognizable in bright field as optically clear superficial tissue with a clean border and evidence of radial cellular architecture. i. Quantification of the mean ratio of spheroids or enCORs displaying neuroepithelium. Error bars are S.E.M., note the increased variance in spheroids compared with enCORs. *P<0.05, Student's two-tailed t-test, n = 7 independent batches each. Scale bars: 100 μm in a., 250 μm in b., c., d., g., 500 μm in e., f.
Supplementary Figure 3 Decreased formation of non-neural tissue in enCORs
a. RT-PCR for markers of pluripotency in 20-day H9 microfilament enCORs or spherical organoids. Embryonic stem cells (H9 cells) are positive control and Neg. is the negative water control. Panels at right show the full gel with DNA ladder. b. Full length gels of data shown in Figure 1g with DNA ladder. c.Bright field images several days following Matrigel embedding (day 17) showing the prevalence of non-neural cysts in H9 spherical organoids compared with microfilament organoids, and quantification of the mean ratio of organoids exhibiting non-neural cysts (panel at the right). Error bars are S.E.M. *P<0.05, Student's two-tailed t-test, n=97 organoids from 5 batches for spheroid, n=69 organoids from 5 batches for enCOR. d. Immunohistochemical staining of a H9 spheroid reveals occasional endoderm (Sox17+, inset) and nonneural epithelia (arrow) even at a later time point of 43 days. The same staining of a day 43 H9 enCOR does not exhibit these nonneural identities and instead contains several large lobes of brain tissue (arrows). Scale bars: 200 μm in c., d.
Supplementary Figure 4 Reproducible formation of forebrain tissue in enCORs
a. Bright field image of H9 microfilament organoids after Matrigel embedding (day 19), displaying numerous small buds of neuroepithelium (arrows). b. Embedded H9 microfilament organoids at day 19, following three days of treatment with CHIR. Note the larger, more continuous buds of neuroepithelium (arrows). c. Staining of day 30 H9 spheroids for neural (N-Cadherin or Nestin) and non-neural identities (Sox17 or Brachyury) revealing similar quantities of these identities with and without a three day pulse of CHIR. Boxed regions are shown at higher magnification below panels. d. Quantification of fluorescence staining of the above markers (mean grey value relative to DAPI). Mean values shown ± SD. n = 6 spheroids (day 30, H9) for each condition. e. Representative images of stained sections of day 40 H9 organoids used for quantification shown in Figure 2c. Individual lobes can be recognized as radially oriented, polarized neuroepithelium surrounding a fluid filled ventricle. Serial sections were stained for the indicated markers. Forebrain tissues (white arrows) are Foxg1+ and exhibit fainter Otx2 signal, midbrain tissues (white arrowheads) exhibit strong immunoreactivity for Otx2 and lack other markers, and cerebellar tissue (yellow arrowhead) exhibits staining for both Otx2 and En2, while En2 single positive regions are hindbrain. Foxg1+ regions showing strong Pax6 staining are dorsal forebrain. However, Pax6 also stains other regions such as that marked by the yellow arrow, which is likely spinal chord, as it does not exhibit staining for the other markers. f. Staining for the ventral forebrain marker Gsh2 and the dorsal marker Tbr2 in H9 enCORs at day 40 reveals the presence of both regions of the forebrain. Note that Gsh2 staining on the ventricular surface is nonspecific. g. Staining for the marker of choroid plexus TTR reveals the presence of this region in H9 40-day enCORs h. Staining for the marker of hippocampus Prox1 reveals the presence of this tissue in H9 40-day enCORs. Scale bars: 500 μm in a., b., c., e., 100 μm in f., g., h.
Supplementary Figure 5 Transcriptome profiling reveals increased neural identity and preferential formation of forebrain
a. Fold enrichment and -log10(P-value) of GO Slim Biological Process terms for genes increased or decreased in enCORs at 20 and 60 days (data from Supplementary Table 1). b. Heatmap of hierarchical clustering of genes according to log2fc values at 20 days and 60 days in enCORs versus spheroids (data from Supplementary Table 2). Clusters are marked and GO biological process term summaries of terms identified (Supplementary Fig. 6) shown at right.
Supplementary Figure 6 GO terms in identified clusters of gene expression
Fold enrichment and -log10(P-value) of GO Biological Process terms for genes in 7 clusters identified by hierarchical clustering according to log2fc value (Supplementary Fig. 5b).
Supplementary Figure 7 Individual marker genes of pluripotency, germ layer identity and brain patterning
a. Screen shots of the IGV view of single gene tracks of markers of pluripotency, neuroectoderm and mesendoderm in spheroid and micropatterned enCORs (labeled uPatterned) at 20 days. b. Single gene tracks of markers of rostral-caudal and dorsal-ventral brain patterning in spheroid and enCORs at 60 days. Schematic of marker expression in the developing brain shown below. c. Heatmap of Spearman correlation coefficients of differentially expressed genes at 60 days in H9 spheroids and enCORs with the Allen BrainSpan transcriptome, sorted by anterior-posterior regional identity and four stages of development.
Supplementary Figure 8 enCORs with ECM addition display organized cortical plate
a. Bright field images of H9 60-day spheroids lacking dissolved ECM and enCORs with dissolved Matrigel showing a band of density (arrow) suggesting the presence of a dense cortical plate. b. Higher magnification of immunostaining shown in Figure 3d for Laminin, MAP2 and Ctip2 showing the radial orientation of neurons that have reached the cortical plate (brackets). c. Side-by-side comparison of CP (arrowheads, brackets) in 60-day H9, 60-day iPSC, and 70-day H1 organoids stained by H&E. d. H&E staining of 60-day H9 enCORs with dissolved Matrigel show a dense band consistent with CP, whereas enCORs instead treated from day 30 with the matrix metalloprotease inhibitor GM6001 show no signs of CP formation. e. H&E staining of 60-day H9 enCORs with dissolved Matrigel compared with dissolved laminin, dissolved laminin/entactin complex, or a mixture of laminin/entactin and collagen I. Note the presence of a dense CP band with Matrigel, while laminin alone or in combination with entactin and collagen I display no clear CP and only occasionally a faint band of cells with laminin/entactin. Scale bars: 200 μm in a., 100 μm in b., c., d., e.
Supplementary Figure 9 Comparison of enCOR method with other 3D organoid methods
a. H&E staining of H9 enCOR, H9 cortical spheroid generated according to Pasca et al. and iPSC SFEBq organoids generated according to Kadoshima et al. SFEBq 1 and 2 were generated in independent laboratories, with SFEBq 1 generated in the Lancaster laboratory and SFEBq 2 generated according to Otani et al. in the Livesey laboratory. All are shown at 60-days of development and at the same magnification. Note the dense CP band (black arrowheads) with the enCOR method and the overall larger size as well as presence of larger lobules surrounding fluid-filled ventricle-like spaces. While the other methods show ventricular zone regions (yellow arrowheads), such a dense organized CP is not visible. Lower panels are higher magnifications of the boxed regions. b. Immunofluorescent staining similarly reveals a Ctip2+ CP (bracket) surrounded by Map2 staining in an iPSC enCOR while other methods do not display this architecture. Laminin staining is detected as expected in enCORs and SFEBq organoids where it can be seen in some SFEBq organoids coating the external surface (white arrowheads) with adjacent formation of a thin band of cells reminiscent of a rudimentary CP (yellow asterisk). c. Staining for neuronal subtypes in developing CP in H9 enCORs at two time points reveals the progressive separation of upper layer (Satb2+ and Cux2+) which labels a broad population, and deep layer (Ctip2+) neurons which label neurons in deeper regions of the CP. Note the progressive thickening of the radially organized CP, and the evident layers exhibiting transitioning organization of neuronal processes (Map2 staining). Scale bars: 500 μm in a. upper panel, 250 μm in a. lower panel, 100 μm and b., c.
Supplementary Figure 10 Slice culture and live imaging in enCORs enables visualization of neuronal migration and activity.
a. Nestin staining reveals basal processes (arrowheads) with end feet that terminate at the laminin positive basement membrane (arrows) in a 60-day H9 enCOR. b. Nestin staining in 60-day H9 spheroid reveals the presence of radial glial processes that show disorganization outside the ventricular zone (arrowheads) and terminal end feet within the tissue (arrows). c. Frames from live imaging of a GFP electroporated neuron (Supplementary Video 3) showing the dynamic multipolar phase of migration with multiple transient processes (arrowheads) extending from the main cell body (arrow). Time stamp is hours:minutes. d. Frames from live imaging of a putative tangentially migrating interneuron (Supplementary Video 6). Note the presence of two leading processes, which the cell body alternates between. Before the nucleus (arrow) enters the leading process, a small accumulation can be seen (arrowhead) consistent with the antecedent movement of the centrosome, a behavior observed in vivo (Higginbotham, H. R. & Gleeson, J. G. The centrosome in neuronal development. Trends in Neurosciences 30, 276–283 (2007)). e. A false color heatmap frame of live imaging with the calcium dye Fluo-4 (Supplementary Video 7) and single cell tracings of the indicated cells, labeled regions of interest (ROI), as measured by change in relative fluorescence (ΔF/F = (mean grey value - minimum grey value)/minimum grey value) showing spontaneous calcium surges. Scale bars: 100 μm in a. and b., 20 μm in c., 50 μm in d., e.
Supplementary information
Supplementary Table 1
Upregulated and downregulated genes in enCORs at 20 days and 60 days, and the associated GO term analysis (XLSX 116 kb)
Supplementary Table 2
Hierarchical clusters of up and down regulated genes and the associated GO term analysis (XLSX 193 kb)
Supplementary Video 1
Live imaging of GFP electroporated slice culture of H1 1009 enCOR electroporated with GFP on day 63, followed by vibratome sectioning four 1010 days later and live imaging 24-hours later (MOV 50315 kb)
Supplementary Video 2
Live image of GFP electroporated H9 organoid displaying several labeled radial glial processes and their endfeet (asterisks) as well as several migrating neurons (arrows). (MOV 110588 kb)
Supplementary Video 3
Higher magnification of the multipolar neuron marked by blue arrow in Supplementary Video 2. (MOV 474 kb)
Supplementary Video 4
Live imaging of farnesyl-GFP electroporated cells in slice culture taken 13 days after electroporation (MOV 130051 kb)
Supplementary Video 5
Higher magnification of live imaging of farnesyl-GFP electroporated slice culture shown in Supplementary Video 4 (MOV 190 kb)
Supplementary Video 6
Higher magnification of the neuron marked by the orange arrow in Supplementary Video 2 (MOV 10907 kb)
Supplementary Video 7
Live imaging of spontaneous calcium surges using the calcium dye Fluo-4 in a slice culture taken 13 days after electroporation (MOV 85495 kb)
Rights and permissions
About this article
Cite this article
Lancaster, M., Corsini, N., Wolfinger, S. et al. Guided self-organization and cortical plate formation in human brain organoids. Nat Biotechnol 35, 659–666 (2017). https://doi.org/10.1038/nbt.3906
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3906
This article is cited by
-
A multidimensional atlas of human glioblastoma-like organoids reveals highly coordinated molecular networks and effective drugs
npj Precision Oncology (2024)
-
Single-cell mapping of lipid metabolites using an infrared probe in human-derived model systems
Nature Communications (2024)
-
Innovative explorations: unveiling the potential of organoids for investigating environmental pollutant exposure
Environmental Science and Pollution Research (2024)
-
Patentability of Brain Organoids derived from iPSC– A Legal Evaluation with Interdisciplinary Aspects
Neuroethics (2024)
-
The Case Against Organoid Consciousness
Neuroethics (2024)