Abstract
We applied a fusion of CRISPR-Cas9 and activation-induced cytidine deaminase (Target-AID) for point mutagenesis at genomic regions specified by single guide RNAs (sgRNAs) in two crop plants. In rice, we induced multiple herbicide-resistance point mutations by multiplexed editing using herbicide selection, while in tomato we generated marker-free plants with homozygous heritable DNA substitutions, demonstrating the feasibility of base editing for crop improvement.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
05 April 2017
In the version of this article initially published, in the first paragraph after the abstract, a reference for Komor et al. was left out. The reference is now inserted in line 5 as “the base-editing1,18 construct”, replacing “with a construct.” The error has been corrected for the print, PDF and HTML versions of this article.
References
Nishida, K. et al. Science 102, 553–563 (2016).
Mikami, M., Toki, S. & Endo, M. Plant Mol. Biol. 88, 561–572 (2015).
Yu, Q. & Powles, S.B. Pest Manag. Sci. 70, 1340–1350 (2014).
Fauser, F., Schiml, S. & Puchta, H. Plant J. 79, 348–359 (2014).
Ariizumi, T., Shinozaki, Y. & Ezura, H. Breed. Sci. 63, 3–13 (2013).
Fleishon, S., Shani, E., Ori, N. & Weiss, D. New Phytol. 190, 609–617 (2011).
Lu, Y. & Zhu, J.K. Mol. Plant http://dx.doi.org/10.1016/j.molp.2016.11.013 (2016).
Li, J., Sun, Y., Du, J., Zhao, Y. & Xia, L. Mol. Plant http://dx.doi.org/10.1016/j.molp.2016.12.001 (2016).
Terada, R., Urawa, H., Inagaki, Y., Tsugane, K. & Iida, S. Nat. Biotechnol. 20, 1030–1034 (2002).
Ryan, M.D., King, A.M.Q. & Thomas, G.P. J. Gen. Virol. 72, 2727–2732 (1991).
Toki, S. Plant Mol. Biol. Report. 15, 16–21 (1997).
Terada, R., Asao, H. & Iida, S. Plant Cell Rep. 22, 653–659 (2004).
Dang, T.T., Shimatani, Z., Kawano, Y., Terada, R. & Shimamoto, K. Plant Cell Physiol. 54, 2058–2070 (2013).
Sun, H.J., Uchii, S., Watanabe, S. & Ezura, H. Plant Cell Physiol. 47, 426–431 (2006).
Shimatani, Z. et al. Mol. Genet. Genomics 281, 329–344 (2009).
Lei, Y. et al. Mol. Plant 7, 1494–1496 (2014).
Stemmer, M., Thumberger, T., Del Sol Keyer, M., Wittbrodt, J. & Mateo, J.L. PLoS One 10, e0124633 (2015).
Komor, A.C., Kim, Y.B., Packer, M.S., Zuris, J.A. & Liu, D.R. Nature 533, 420–424 (2016).
Acknowledgements
Plant CRISPR vectors pCAS9-TPC and pChimera were kindly provided by F. Fauser, S. Schiml, and H. Puchta at Karlsruhe Institute of Technology via M. Endo and S. Toki. We thank Y. Iida, R. Ohta, Y. Ueke and M. Suzuki for technical assistance. This work was supported by the Cabinet Office, Government of Japan, Cross-ministerial Strategic Innovation Promotion Program (SIP), “Technologies for creating next-generation agriculture, forestry and fisheries” (funding agency: Bio-oriented Technology Research Advancement Institution, NARO) and partly supported by a Special Coordination Fund for Promoting Science and Technology, Creation of Innovative Centers for Advanced Interdisciplinary Research Areas (Innovative Bioproduction Kobe) from the Ministry of Education, Culture, Sports and Technology (MEXT) of Japan, by the Platform Project for Supporting in Drug Discovery and Life Science Research (Platform for Drug Discovery, Informatics, and Structural Life Science) from the Japan Agency for Medical Research and Development (AMED), by JSPS KAKENHI grant numbers 26119710 and 16K14654 and by the New Energy and Industrial Technology Development Organization (NEDO).
Author information
Authors and Affiliations
Contributions
Z.S., H.I., H.T. and R.T. performed rice experiments. M.T., S.K., T.Y. and H.K. performed tomato experiments. K.N. and T. Arazoe performed NGS analysis. K.N. designed and constructed plasmids. Z.S., T. Ariizumi, R.T. and K.N. designed the experiments. Z.S., K.N., M.T., S.K. and T. Ariizumi, wrote the manuscript. K.N., T. Ariizumi, K.M., H.E. and A.K. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
K.N., Z.S., T. Arazoe and A.K. have filed a patent related to the rice study as PCT/JP2016/085075. S.K., M.T., T.Y., K.M., T. Ariizumi and H.E. have filed a patent related to the tomato study as JP 2017-019921.
Integrated supplementary information
Supplementary Figure 1 Application of Target-AID to rice by EGFP reporter assay and transgenic approach.
(a) Schematics of the switch-mEGFP reporter vector (top) and rice Target-AID (d/nCas9Os-PmCDA1At ) vector (bottom). In the switch module (OFF), a stop codon is inserted immediately downstream of the initiation codon of the EGFP gene, as indicated at the top. LB/RB, left/right border of T-DNA. (b) Fluorescence microscopic images of the transformants of mEGFP marker and Target-AID. Upper panels (Bright field) show shape of calli. Lower panels show GFP signal observed by fluorescence microscopy. Shape of calli are outlined and superimposed. Bar = 1 mm. (c) Frequency of GFP positive calli for each transformant. (d) Sequence alignment of the edited reporter EGFP. The switch module sequence of pRIT3-mEGFP is indicated with the target sequence (red). Mutated positions are shown in blue. Amino acid translation is indicated at the top. (e) Electrophoretogram of CAPS analysis for Fig. 1e and f. 1054 bp PCR product consisting of 1034 bp genomic region and 20 bp primer sequence was digested by PvuII into two fragments (614 bp and 440 bp) in the presence of C-to-T mutation at G590.
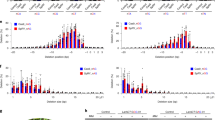
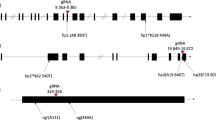
Supplementary Figure 2 Deep sequencing analysis of ALS on- and off-target mutations induced by a series of Target-AID vectors in rice.
Target sequence is highlighted in grey and mismatched bases of the off-target are highlighted in light blue. Mutation frequency is shown and highlighted as indicated at the bottom right. Total indel frequency in the 60 bases (20 bases of target region plus upstream 10 bases and downstream 30 bases) and SNV frequency above 0.1% at each nucleotide position is shown and highlighted.
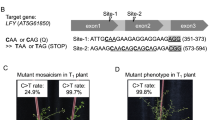
Supplementary Figure 3 Application of Target-AID to tomato.
(a) Engineered Cas9 nuclease expression vectors used for tomato study. Nucleases (Cas9At , nCas9At-PmCDA1Hs , and nCas9At-PmCDA1At ) were under the control of the PcUbi promoter, and transcription was terminated by the Pea3A terminator from P. sativum. sgRNAs were under the control of the Arabidopsis U6 promoter (AtU6). T0 lines were selected on the basis of kanamycin resistance. For construction of ETR1site3 -targeting vector, a backbone of nCas9At-PmCDA1At-2A was used. (b) Targeted mutagenesis frequencies in T0 and T1 tomato lines. ‘No. of T0 lines generated’, number of T0 lines generated for this study; ‘No. of T0 lines sequenced’, number of T0 lines subjected to sequencing analysis; ‘No. of genome-edited T0 lines’, number of transgenic lines with DNA modifications in T0 transgenic lines; ‘No. of T0 lines with indels and no. of lines with substitutions’, number of T0 lines harboring indels and/or substitutions, respectively; ‘No. of T1 lines sequenced’, number of independent T0 generation lines used for sequencing analysis in T1 lines; ‘No. of genome-edited T1 lines’, number of independent T0 generation plants with DNA modifications in T1 lines; ‘No. of T1 lines with indels and no. of T1 lines with substitutions’, number of independent T0 lines harboring indels and substitutions, respectively, in T1 lines. (c) Summary of mutation segregation patterns in T1 offspring plants. Numbers of plants T1 harboring Indel or substitution mutations with each genotype are shown. Genotypes are classified into nine categories including T1 plants with homozygous (Indel/Indel), heterozygous (Indel/WT) or biallelic (Indel/Indel') indels, those with homozygous (S/S), heterozygous (S/WT) or biallelic (S/S' and S/Indel) substitutions, those with WT genotype (WT/WT) and those that contain more than three different sequences (Chimera). Indel' and S' represent different patterns of indel and substitution from those of indel and S, respectively. 167 T1 plants possessed stable DNA substitution, except for T1 plants with chimera genotype.
Supplementary Figure 4 Target-AID vectors induce amino acid substitutions and phenotype in genome-edited tomato plants.
(a) Changes in amino acid residues in DELLA- or ETR1-targeted T1 plants. Substitutions in amino acid residues were due to corresponding DNA substitutions in T1 plants. Line names (starting with #) correspond to those in Supplementary Figure 6b. The marker gene-free plants are highlighted. (b) Change in amino acid residues in DELLA-targeted T2 (#3_2_4) and T1 (#27_9) plants. (c) Leaflet appearance in DELLA-targeted T2 (#3_2_4) and T1 (#27_9) plants with homozygous 2-bp substitution and 12-bp deletion, respectively. Leaflets are highly serrated in WT (arrows), whereas those in procera mutant and the DNA edited plants were less serrated and looked smoother than those in WT. The procera is a loss-of-function allele of DELLA gene which shows reduced leaf serration and was shown as a control to compare leaf phenotype. Bar = 10 mm.
Supplementary Figure 5 Deep sequencing analysis of on- and off-target sites induced by a series of Target-AID and Cas9 vectors in tomato.
(a) Prediction of DELLA off-target sites in tomato. Potential off-target genomic regions were selected by CCTop (http://crispr.cos.uni-heidelberg.de/index.html), and top one off-target site was highlighted in light blue. Distance to the closest exon: 0 indicates that the target site and exon coordinate overlap; NA indicates that the target sites are farther than 100 kb from the next exon. Location of the target site: E, I, and - indicate exonic, intronic, and intergenic, respectively. (b) DELLA on- and off-target mutation frequencies induced by a series of Target-AID and Cas9 in tomato. Target sequence is highlighted in grey and mismatched bases of the off-target are in light blue. Mutation frequency above 0.1% is shown and highlighted as indicated at the bottom right. Total indel frequency in the 42 bases (20 bases of target region plus upstream 10 bases and downstream 12 bases) and SNV frequency above 0.1% at each nucleotide position is shown and highlighted.
Supplementary Figure 6 Marker gene-free DELLA- or ETR1-targeted transgenic plants with stably inherited DNA mutations.
(a) Presence of the kanamycin-resistance marker gene (NPT II) was tested by PCR analysis in T1 plants harboring DNA modifications. PCR analysis was conducted to confirm the absence of the kanamycin-resistance marker gene (NPT II, upper panel) in the genome of progenies derived from T0 lines with engineered Cas9 vectors. Lower panel indicates PCR amplification of the endogenous ACTIN gene, as a positive control (PC) for genomic DNA extraction. The negative control (NC) represents no DNA template added in PCR reaction mixture to prove that no DNA was contaminated in the PCR reaction. # indicates parental line, whereas following numbers indicate individual offspring. Line #3BC1_6 is an F1 plant derived from a cross between line #3 (T0 line) and the WT. WT, wild-type tomato plants (negative control of PCR for NPT II amplification). (b) The mutation spectrum in marker-gene free plants that correspond to those examined in (a). Primary null-segregant, the first generation that carried no transgene; Zygosity, type of stably inherited mutation; Next generation, self-pollinated siblings of primary null-segregant; No. of plants examined, number of transgenic plants used for calculation of mutation efficiency or heritable frequency; Ratio of segregated mutation, percentage of mutation patterns found in the next generation; N.D., Not determined.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 1396 kb)
Rights and permissions
About this article
Cite this article
Shimatani, Z., Kashojiya, S., Takayama, M. et al. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat Biotechnol 35, 441–443 (2017). https://doi.org/10.1038/nbt.3833
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3833
This article is cited by
-
Engineering APOBEC3A deaminase for highly accurate and efficient base editing
Nature Chemical Biology (2024)
-
LsMybW-encoding R2R3-MYB transcription factor is responsible for a shift from black to white in lettuce seed
Plant Cell Reports (2024)
-
CRISPR/Cas-mediated germplasm improvement and new strategies for crop protection
Crop Health (2024)
-
Crop bioengineering via gene editing: reshaping the future of agriculture
Plant Cell Reports (2024)
-
Fusion of a rice endogenous N-methylpurine DNA glycosylase to a plant adenine base transition editor ABE8e enables A-to-K base editing in rice plants
aBIOTECH (2024)