Abstract
Many high-throughput experimental technologies have been developed to assess the effects of large numbers of mutations (variation) on phenotypes. However, designing functional assays for these methods is challenging, and systematic testing of all combinations is impossible, so robust methods to predict the effects of genetic variation are needed. Most prediction methods exploit evolutionary sequence conservation but do not consider the interdependencies of residues or bases. We present EVmutation, an unsupervised statistical method for predicting the effects of mutations that explicitly captures residue dependencies between positions. We validate EVmutation by comparing its predictions with outcomes of high-throughput mutagenesis experiments and measurements of human disease mutations and show that it outperforms methods that do not account for epistasis. EVmutation can be used to assess the quantitative effects of mutations in genes of any organism. We provide pre-computed predictions for ∼7,000 human proteins at http://evmutation.org/.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Change history
30 January 2017
In the version of this article initially published online, an equation in the Online Methods section “Inference of epistatic models of biological sequences section” was incorrect: the term “λJ/2” should have been “λJ/2”. Two sentences later, the sentence presenting the next equation was also incorrect: instead of ending with “and is 0 otherwise,” that should read “and is 1 otherwise.” Both errors have been corrected in the print, PDF and HTML versions of the article.
References
Miersch, S. & Sidhu, S.S. Intracellular targeting with engineered proteins. F1000Res. 5 http://dx.doi.org/10.12688/f1000research.8915.1 (2016).
Boeke, J.D., et al. GENOME ENGINEERING. The Genome Project-Write. Science 353, 126–127 (2016).
Ostrov, N. et al. Design, synthesis, and testing toward a 57-codon genome. Science 353, 819–822 (2016).
Romero, P.A., Tran, T.M. & Abate, A.R. Dissecting enzyme function with microfluidic-based deep mutational scanning. Proc. Natl. Acad. Sci. USA 112, 7159–7164 (2015).
Currin, A., Swainston, N., Day, P.J. & Kell, D.B. Synthetic biology for the directed evolution of protein biocatalysts: navigating sequence space intelligently. Chem. Soc. Rev. 44, 1172–1239 (2015).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Roscoe, B.P. & Bolon, D.N. Systematic exploration of ubiquitin sequence, E1 activation efficiency, and experimental fitness in yeast. J. Mol. Biol. 426, 2854–2870 (2014).
Roscoe, B.P., Thayer, K.M., Zeldovich, K.B., Fushman, D. & Bolon, D.N. Analyses of the effects of all ubiquitin point mutants on yeast growth rate. J. Mol. Biol. 425, 1363–1377 (2013).
Melamed, D., Young, D.L., Gamble, C.E., Miller, C.R. & Fields, S. Deep mutational scanning of an RRM domain of the Saccharomyces cerevisiae poly(A)-binding protein. RNA 19, 1537–1551 (2013).
Stiffler, M.A., Hekstra, D.R. & Ranganathan, R. Evolvability as a function of purifying selection in TEM-1 β-lactamase. Cell 160, 882–892 (2015).
McLaughlin, R.N. Jr., Poelwijk, F.J., Raman, A., Gosal, W.S. & Ranganathan, R. The spatial architecture of protein function and adaptation. Nature 491, 138–142 (2012).
Kitzman, J.O., Starita, L.M., Lo, R.S., Fields, S. & Shendure, J. Massively parallel single-amino-acid mutagenesis. Nat. Methods 12, 203–206, 4, 206 (2015).
Melnikov, A., Rogov, P., Wang, L., Gnirke, A. & Mikkelsen, T.S. Comprehensive mutational scanning of a kinase in vivo reveals substrate-dependent fitness landscapes. Nucleic Acids Res. 42, e112 (2014).
Araya, C.L. et al. A fundamental protein property, thermodynamic stability, revealed solely from large-scale measurements of protein function. Proc. Natl. Acad. Sci. USA 109, 16858–16863 (2012).
Firnberg, E., Labonte, J.W., Gray, J.J. & Ostermeier, M. A comprehensive, high-resolution map of a gene's fitness landscape. Mol. Biol. Evol. 31, 1581–1592 (2014).
Starita, L.M. et al. Massively parallel functional analysis of BRCA1 RING domain variants. Genetics 200, 413–422 (2015).
Rockah-Shmuel, L., Tóth-Petróczy, Á. & Tawfik, D.S. Systematic mapping of protein mutational space by prolonged drift reveals the deleterious effects of seemingly neutral mutations. PLoS Comput. Biol. 11, e1004421 (2015).
Jacquier, H. et al. Capturing the mutational landscape of the beta-lactamase TEM-1. Proc. Natl. Acad. Sci. USA 110, 13067–13072 (2013).
Qi, H. et al. A quantitative high-resolution genetic profile rapidly identifies sequence determinants of hepatitis C viral fitness and drug sensitivity. PLoS Pathog. 10, e1004064 (2014).
Wu, N.C. et al. Functional constraint profiling of a viral protein reveals discordance of evolutionary conservation and functionality. PLoS Genet. 11, e1005310 (2015).
Mishra, P., Flynn, J.M., Starr, T.N. & Bolon, D.N. Systematic mutant analyses elucidate general and client-specific aspects of Hsp90 function. Cell Rep. 15, 588–598 (2016).
Doud, M.B. & Bloom, J.D. Accurate measurement of the effects of all amino-acid mutations to influenza hemagglutinin. bioRxiv 8, E155 (2016).
Deng, Z. et al. Deep sequencing of systematic combinatorial libraries reveals β-lactamase sequence constraints at high resolution. J. Mol. Biol. 424, 150–167 (2012).
Starita, L.M. et al. Activity-enhancing mutations in an E3 ubiquitin ligase identified by high-throughput mutagenesis. Proc. Natl. Acad. Sci. USA 110, E1263–E1272 (2013).
Aakre, C.D. et al. Evolving new protein-protein interaction specificity through promiscuous intermediates. Cell 163, 594–606 (2015).
Julien, P., Miñana, B., Baeza-Centurion, P., Valcárcel, J. & Lehner, B. The complete local genotype-phenotype landscape for the alternative splicing of a human exon. Nat. Commun. 7, 11558 (2016).
Li, C., Qian, W., Maclean, C.J. & Zhang, J. The fitness landscape of a tRNA gene. Science 352, 837–840 (2016).
Fowler, D.M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Gasperini, M., Starita, L. & Shendure, J. The power of multiplexed functional analysis of genetic variants. Nat. Protoc. 11, 1782–1787 (2016).
Sarkisyan, K.S. et al. Local fitness landscape of the green fluorescent protein. Nature 533, 397–401 (2016).
Boucher, J.I., Bolon, D.N. & Tawfik, D.S. Quantifying and understanding the fitness effects of protein mutations: Laboratory versus nature. Protein Sci. 25, 1219–1226 (2016).
Gong, L.I., Suchard, M.A. & Bloom, J.D. Stability-mediated epistasis constrains the evolution of an influenza protein. eLife 2, e00631 (2013).
Kachroo, A.H. et al. Evolution. Systematic humanization of yeast genes reveals conserved functions and genetic modularity. Science 348, 921–925 (2015).
Sim, N.L. et al. SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40, W452 (2012).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Adzhubei, I., Jordan, D.M. & Sunyaev, S.R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 76, 7.20 (2013).
Breen, M.S., Kemena, C., Vlasov, P.K., Notredame, C. & Kondrashov, F.A. Epistasis as the primary factor in molecular evolution. Nature 490, 535–538 (2012).
McCandlish, D.M., Shah, P. & Plotkin, J.B. Epistasis and the dynamics of reversion in molecular evolution. Genetics 203, 1335–1351 (2016).
Ovchinnikov, S., Kamisetty, H. & Baker, D. Robust and accurate prediction of residue-residue interactions across protein interfaces using evolutionary information. eLife 3, e02030 (2014).
Hopf, T.A. et al. Sequence co-evolution gives 3D contacts and structures of protein complexes. eLife 3 http://dx.doi.org/10.7554/eLife.03430 (2014).
Hopf, T.A., et al. Three-dimensional structures of membrane proteins from genomic sequencing. Cell 149, 1607–1621 (2012).
Marks, D.S., Hopf, T.A. & Sander, C. Protein structure prediction from sequence variation. Nat. Biotechnol. 30, 1072–1080 (2012).
Marks, D.S. et al. Protein 3D structure computed from evolutionary sequence variation. PLoS One 6, e28766 (2011).
Morcos, F. et al. Direct-coupling analysis of residue coevolution captures native contacts across many protein families. Proc. Natl. Acad. Sci. USA 108, E1293–E1301 (2011).
Jones, D.T., Buchan, D.W., Cozzetto, D. & Pontil, M. PSICOV: precise structural contact prediction using sparse inverse covariance estimation on large multiple sequence alignments. Bioinformatics 28, 184–190 (2012).
Mann, J.K. et al. The fitness landscape of HIV-1 gag: advanced modeling approaches and validation of model predictions by in vitro testing. PLoS Comput. Biol. 10, e1003776 (2014).
Lapedes, A., Giraud, B. & Jarzynski, C. Using sequence alignments to predict protein structure and stability with high accuracy. Preprint at https://arxiv.org/pdf/1207.2484v1.pdf (2012).
Figliuzzi, M., Jacquier, H., Schug, A., Tenaillon, O. & Weigt, M. Coevolutionary landscape inference and the context-dependence of mutations in beta-lactamase TEM-1. Mol. Biol. Evol. 33, 268–280 (2016).
Sella, G. & Hirsh, A.E. The application of statistical physics to evolutionary biology. Proc. Natl. Acad. Sci. USA 102, 9541–9546 (2005).
Giraud, B.G., Heumann, J.M. & Lapedes, A.S. Superadditive correlation. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 59, 4983–4991 (1999).
Ovchinnikov, S. et al. Large-scale determination of previously unsolved protein structures using evolutionary information. eLife 4, e09248 (2015).
Kosciolek, T. & Jones, D.T. De novo structure prediction of globular proteins aided by sequence variation-derived contacts. PLoS One 9, e92197 (2014).
Besag, J. Statistical analysis of non-lattice data. Statistician 24, 179–195 (1975).
Balakrishnan, S., Kamisetty, H., Carbonell, J.G., Lee, S.I. & Langmead, C.J. Learning generative models for protein fold families. Proteins 79, 1061–1078 (2011).
Kamisetty, H., Ovchinnikov, S. & Baker, D. Assessing the utility of coevolution-based residue-residue contact predictions in a sequence- and structure-rich era. Proc. Natl. Acad. Sci. USA 110, 15674–15679 (2013).
Ekeberg, M., Lövkvist, C., Lan, Y., Weigt, M. & Aurell, E. Improved contact prediction in proteins: using pseudolikelihoods to infer Potts models. Phys. Rev. E 87, 012707 (2013).
Di Nardo, A.A., Larson, S.M. & Davidson, A.R. The relationship between conservation, thermodynamic stability, and function in the SH3 domain hydrophobic core. J. Mol. Biol. 333, 641–655 (2003).
Halabi, N., Rivoire, O., Leibler, S. & Ranganathan, R. Protein sectors: evolutionary units of three-dimensional structure. Cell 138, 774–786 (2009).
Philip, A.F., Kumauchi, M. & Hoff, W.D. Robustness and evolvability in the functional anatomy of a PER-ARNT-SIM (PAS) domain. Proc. Natl. Acad. Sci. USA 107, 17986–17991 (2010).
Bershtein, S., Mu, W. & Shakhnovich, E.I. Soluble oligomerization provides a beneficial fitness effect on destabilizing mutations. Proc. Natl. Acad. Sci. USA 109, 4857–4862 (2012).
Landrum, M.J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D1, D862–D868 (2016).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Capriotti, E., Calabrese, R. & Casadio, R. Predicting the insurgence of human genetic diseases associated to single point protein mutations with support vector machines and evolutionary information. Bioinformatics 22, 2729–2734 (2006).
Grimm, D.G. et al. The evaluation of tools used to predict the impact of missense variants is hindered by two types of circularity. Hum. Mutat. 36, 513–523 (2015).
Bromberg, Y., Yachdav, G. & Rost, B. SNAP predicts effect of mutations on protein function. Bioinformatics 24, 2397–2398 (2008).
van Nimwegen, E. Inferring contacting residues within and between proteins: what do the probabilities mean? PLoS Comput. Biol. 12, e1004726 (2016).
Eddy, S.R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Suzek, B.E., Wang, Y., Huang, H., McGarvey, P.B. & Wu, C.H. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015).
Nawrocki, E.P. et al. Rfam 12.0: updates to the RNA families database. Nucleic Acids Res. 43, D130–D137 (2015).
Jaynes, E.T. Information theory and statistical mechanics. Phys. Rev. 106, 620 (1957).
Dunn, S.D., Wahl, L.M. & Gloor, G.B. Mutual information without the influence of phylogeny or entropy dramatically improves residue contact prediction. Bioinformatics 24, 333–340 (2008).
Toth-Petroczy, A. et al. Structured states of disordered proteins from genomic sequences. Cell 167, 158–170.e12 (2016).
Reva, B., Antipin, Y. & Sander, C. Determinants of protein function revealed by combinatorial entropy optimization. Genome Biol. 8, R232 (2007).
Kosorok, M.R. Brownian distance covariance and high dimensional data. Ann. Appl. Stat. 3, 1266–1269 (2009).
Matthews, B.W. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim. Biophys. Acta 405, 442–451 (1975).
Robin, X. et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinformatics 12, 77 (2011).
Berman, H.M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Pérez, F. & Granger, B.E. IPython: a system for interactive scientific computing. Comput. Sci. Eng. 9, 21–29 (2007).
Van der Walt, S., Colbert, S.C. & Varoquaux, G. The NumPy array: a structure for efficient numerical computation. Comput. Sci. Eng. 13, 22–30 (2011).
Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Acknowledgements
The authors would like to thank A. Lapedes, B. Rost, and members of the Marks laboratory for scientific discussion, and J. Reeb for help with existing mutation prediction software. C.S. was funded by NIGMS (R01GM106303). D.S.M. and T.A.H. were funded by NIGMS (R01GM106303) and the Raymond and Beverley Sackler Foundation. J.B.I. was funded by an NSF Graduate Research Fellowship (DGE1144152).
Author information
Authors and Affiliations
Contributions
D.S.M., T.A.H. and C.S. initiated the project. T.A.H. and J.B.I. developed algorithms and wrote software. T.A.H., J.B.I. and D.S.M. analyzed the data with contributions from M.S. F.J.P. advised on the interpretation of experiments. C.P.I.S. supplied processed human genetic variation data. T.A.H., J.B.I., C.S. and D.S.M. wrote the paper. D.S.M. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Computation of context-dependent mutation effects from the coevolutionary sequence record
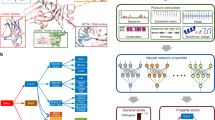
Left: The evolutionary pressure to maintain functional biomolecules leaves a record of amino acid or nucleotide co-conservation in multiple alignments of a sequence family. Middle: A pairwise graphical model learned from the natural sequence variation reveals family-specific constraints between pairs of positions (Jij) as well as at single sites (hi). Each hi is a vector unique to each position in the family that describes the relative favorability of different amino acids or nucleotides at that position, while each Jij is a matrix unique to each pair of positions describing an interaction pattern for the relative favorability of different combinations of amino acids/nucleotides at those positions. The values of these parameters are inferred by maximizing the probability of observing the natural sequences, with additional penalties for model complexity. Right: The inferred probability model can be applied to compute the relative effect of both single and higher-order substitutions. The calculation evaluates how compatible substitutions in the context of the wild-type sequence are with the functional constraints on the family by summing over the changes of couplings to all other sites (Jij), as well as the changes of single-site constraint terms in the changed positions (hi).
Supplementary Figure 2 Evolutionary statistical energy landscapes capture mutational sensitivity of sites
Computed mutational sensitivities per position (average difference in evolutionary statistical energy ΔE across all possible substitutions for each site) based on the epistatic model agree with experimental mutational sensitivities on 20 analyzed single-substitution landscapes for 15 biomolecules as measured by Spearman's rank correlation coefficient ρ. For correlations with the effects of individual substitutions, see Fig. 3a.
Supplementary Figure 3 Effect distributions of experimental mutation scans
The analyzed experimental datasets show considerable differences in the overall shape of their effect distributions. Many of the experiments are biased towards large fractions of neutral or deleterious variants, or have bimodal effect distributions biased towards either end of the effect scale but with little resolution of intermediate effects (here measured by the skewness and normality of the distributions (Online Methods); plots are ordered from negative to positive skew). Summary statistics can also be found in Supplementary Table 4.
Supplementary Figure 4 Agreement between evolutionary statistical energies and all features tested in experimental mutation scans
Full set of Spearman's rank correlation coefficients ρ between evolutionary statistical energies ΔE and experimental effects across all tested functional features and conditions in the analyzed mutation scans (e.g. different antibiotic concentrations or number of rounds of selection). Correlation coefficients are provided in Supplementary Table 3.
Supplementary Figure 5 Agreement of ΔE with bacterial fitness depends on strength of antibiotic selection
(a) Many mutations to TEM-1 β-lactamase which are deleterious according to ΔE are only revealed as deleterious in vivo by increasing selective pressure through higher ampicillin concentrations (mutational sensitivity per position (average ΔE); left to right, shades of blue additionally indicate concentration of first significant effect determined by fitting a two-component Gaussian mixture model at 2500 μg/ml). (b) Agreement between ΔE and experimental effects for kanamycin kinase decreases as the concentration of antibiotic is increased to a point where most variants are completely depleted and intermediate fitness effects cannot be resolved anymore (mixture model fitted at 1:8 WT MIC in log space).
Supplementary Figure 6 Prediction of human disease variants
(a) Evolutionary statistical energies ΔE computed using the independent model separate human disease-associated variants from frequent alleles in the population, but not as strongly as the epistatic model (Fig. 3b). The separation increases with the minimum allele frequency (AF) of the variants assumed to be neutral (area under the ROC curve (AUC)=0.88 for AF≥0.1, AUC=0.90 for AF≥0.25, AUC=0.92 for AF≥0.5). (b) The epistatic model outperforms all other tested methods on the HumVar dataset without any training on disease variants, as measured by the area under the ROC curve (colored lines for individual methods; grey line: expectation for random classifier; inset: AUC across the full range of specificities (left) and up to a false positive rate of 20% (right); AUCs of SIFT are < 0.5). Since PolyPhen-2 was trained on HumVar, the results here may overestimate its performance (see Online Methods for explanation). (c) On the subset of "difficult" variants that are predicted differently by SIFT and PolyPhen-2, the epistatic model is more accurate than all other methods but overall AUCs are lower than on the full dataset (figure elements as in b).
Supplementary Figure 7 Epistatic interactions critical for accurate prediction of functional residues
Across the proteins with high-throughput datasets where epistatic interactions lead to better agreement with the experimental data (first column), certain subgroups of substitutions contribute to the overall reduction in error, while others are predicted with comparable or slightly better accuracy by the independent model (for definitions of subgroups, see Online Methods).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 6 (PDF 1504 kb)
Supplementary Table 2
Computed and experimental mutational landscapes. (XLSX 7104 kb)
Supplementary Table 3
Correlations between evolutionary statistical energy differences and experimental data. (XLSX 34 kb)
Supplementary Table 4
Skew and normality of effect distributions of experimental mutation scans. (XLSX 10 kb)
Supplementary Table 5
Comparison of epistatic model to established methods. (XLSX 20 kb)
Supplementary Table 7
Correlations with experiments across varying alignments depths. (XLSX 102 kb)
Supplementary Table 8
Error analysis for individual substitutions. (XLSX 2626 kb)
Supplementary Table 9
Evolutionary couplings for all analyzed biomolecules. (XLSX 17114 kb)
Supplementary Code
Source code for inferring pairwise graphical models from sequence alignments and predicting the effects of mutations. (ZIP 14051 kb)
Rights and permissions
About this article
Cite this article
Hopf, T., Ingraham, J., Poelwijk, F. et al. Mutation effects predicted from sequence co-variation. Nat Biotechnol 35, 128–135 (2017). https://doi.org/10.1038/nbt.3769
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3769
This article is cited by
-
Machine learning for functional protein design
Nature Biotechnology (2024)
-
Evolution shapes interaction patterns for epistasis and specific protein binding in a two-component signaling system
Communications Chemistry (2024)
-
Deep generative design of RNA family sequences
Nature Methods (2024)
-
SESNet: sequence-structure feature-integrated deep learning method for data-efficient protein engineering
Journal of Cheminformatics (2023)
-
Lynch syndrome, molecular mechanisms and variant classification
British Journal of Cancer (2023)