Abstract
Gene expression is controlled by enhancers that activate transcription from the core promoters of their target genes. Although a key function of core promoters is to convert enhancer activities into gene transcription, whether and how strongly they activate transcription in response to enhancers has not been systematically assessed on a genome-wide level. Here we describe self-transcribing active core promoter sequencing (STAP-seq), a method to determine the responsiveness of genomic sequences to enhancers, and apply it to the Drosophila melanogaster genome. We cloned candidate fragments at the position of the core promoter (also called minimal promoter) in reporter plasmids with or without a strong enhancer, transfected the resulting library into cells, and quantified the transcripts that initiated from each candidate for each setup by deep sequencing. In the presence of a single strong enhancer, the enhancer responsiveness of different sequences differs by several orders of magnitude, and different levels of responsiveness are associated with genes of different functions. We also identify sequence features that predict enhancer responsiveness and discuss how different core promoters are employed for the regulation of gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Banerji, J., Rusconi, S. & Schaffner, W. Expression of a β-globin gene is enhanced by remote SV40 DNA sequences. Cell 27, 299–308 (1981).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Roeder, R.G. The role of general initiation factors in transcription by RNA polymerase II. Trends Biochem. Sci. 21, 327–335 (1996).
Kadonaga, J.T. Perspectives on the RNA polymerase II core promoter. Wiley Interdiscip. Rev. Dev. Biol. 1, 40–51 (2012).
Core, L.J. et al. Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat. Genet. 46, 1311–1320 (2014).
Kim, T.-K. & Shiekhattar, R. Architectural and functional commonalities between enhancers and promoters. Cell 162, 948–959 (2015).
Spitz, F. & Furlong, E.E.M. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Kvon, E.Z. et al. Genome-scale functional characterization of Drosophila developmental enhancers in vivo. Nature 512, 91–95 (2014).
Juven-Gershon, T., Cheng, S. & Kadonaga, J.T. Rational design of a super core promoter that enhances gene expression. Nat. Methods 3, 917–922 (2006).
Zabidi, M.A. et al. Enhancer-core-promoter specificity separates developmental and housekeeping gene regulation. Nature 518, 556–559 (2015).
Ede, C., Chen, X., Lin, M.-Y. & Chen, Y.Y. Quantitative analyses of core promoters enable precise engineering of regulated gene expression in mammalian cells. ACS Synth. Biol. 5, 395–404 (2016).
Lubliner, S. et al. Core promoter sequence in yeast is a major determinant of expression level. Genome Res. 25, 1008–1017 (2015).
Patwardhan, R.P. et al. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis. Nat. Biotechnol. 27, 1173–1175 (2009).
Arnold, C.D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Duttke, S.H.C. et al. Perspectives on unidirectional versus divergent transcription. Mol. Cell 60, 348–349 (2015).
Andersson, R. et al. Human gene promoters are intrinsically bidirectional. Mol. Cell 60, 346–347 (2015).
Gu, W. et al. CapSeq and CIP-TAP identify Pol II start sites and reveal capped small RNAs as C. elegans piRNA precursors. Cell 151, 1488–1500 (2012).
Nechaev, S. et al. Global analysis of short RNAs reveals widespread promoter-proximal stalling and arrest of Pol II in Drosophila. Science 327, 335–338 (2010).
Ni, T. et al. A paired-end sequencing strategy to map the complex landscape of transcription initiation. Nat. Methods 7, 521–527 (2010).
Ohler, U., Liao, G.-C., Niemann, H. & Rubin, G.M. Computational analysis of core promoters in the Drosophila genome. Genome Biol. 3, RESEARCH0087 (2002).
Saito, K. et al. A regulatory circuit for piwi by the large Maf gene traffic jam in Drosophila. Nature 461, 1296–1299 (2009).
Sienski, G., Dönertas, D. & Brennecke, J. Transcriptional silencing of transposons by Piwi and maelstrom and its impact on chromatin state and gene expression. Cell 151, 964–980 (2012).
Core, L.J. et al. Defining the status of RNA polymerase at promoters. Cell Rep. 2, 1025–1035 (2012).
Pfeiffer, B.D. et al. Tools for neuroanatomy and neurogenetics in Drosophila. Proc. Natl. Acad. Sci. USA 105, 9715–9720 (2008).
Adelman, K. & Lis, J.T. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nat. Rev. Genet. 13, 720–731 (2012).
modENCODE Consortium. et al. Identification of functional elements and regulatory circuits by Drosophila modENCODE. Science 330, 1787–1797 (2010).
Burke, T.W. & Kadonaga, J.T. Drosophila TFIID binds to a conserved downstream basal promoter element that is present in many TATA-box-deficient promoters. Genes Dev. 10, 711–724 (1996).
Lim, C.Y. et al. The MTE, a new core promoter element for transcription by RNA polymerase II. Genes Dev. 18, 1606–1617 (2004).
Zeitlinger, J. et al. RNA polymerase stalling at developmental control genes in the Drosophila melanogaster embryo. Nat. Genet. 39, 1512–1516 (2007).
Engström, P.G., Ho Sui, S.J., Drivenes, O., Becker, T.S. & Lenhard, B. Genomic regulatory blocks underlie extensive microsynteny conservation in insects. Genome Res. 17, 1898–1908 (2007).
Ponjavic, J. et al. Transcriptional and structural impact of TATA-initiation site spacing in mammalian core promoters. Genome Biol. 7, R78 (2006).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. B 58, 267–288 (1996).
Kim, T.-K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
De Santa, F. et al. A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol. 8, e1000384 (2010).
Lam, M.T.Y., Li, W., Rosenfeld, M.G. & Glass, C.K. Enhancer RNAs and regulated transcriptional programs. Trends Biochem. Sci. 39, 170–182 (2014).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Scruggs, B.S. et al. Bidirectional transcription arises from two distinct hubs of transcription factor binding and active chromatin. Mol. Cell 58, 1101–1112 (2015).
Hah, N. et al. A rapid, extensive, and transient transcriptional response to estrogen signaling in breast cancer cells. Cell 145, 622–634 (2011).
Tomancak, P. et al. Systematic determination of patterns of gene expression during Drosophila embryogenesis. Genome Biol. 3, RESEARCHH0088 (2002).
Jayaprakash, A.D., Jabado, O., Brown, B.D. & Sachidanandam, R. Identification and remediation of biases in the activity of RNA ligases in small-RNA deep sequencing. Nucleic Acids Res. 39, e141 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Crooks, G.E., Hon, G., Chandonia, J.-M. & Brenner, S.E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Kodzius, R. et al. CAGE: cap analysis of gene expression. Nat. Methods 3, 211–222 (2006).
Batut, P., Dobin, A., Plessy, C., Carninci, P. & Gingeras, T.R. High-fidelity promoter profiling reveals widespread alternative promoter usage and transposon-driven developmental gene expression. Genome Res. 23, 169–180 (2013).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Stampfel, G. et al. Transcriptional regulators form diverse groups with context-dependent regulatory functions. Nature 528, 147–151 (2015).
Adryan, B. & Teichmann, S.A. FlyTF: a systematic review of site-specific transcription factors in the fruit fly Drosophila melanogaster. Bioinformatics 22, 1532–1533 (2006).
Bailey, T.L. & Gribskov, M. Combining evidence using p-values: application to sequence homology searches. Bioinformatics 14, 48–54 (1998).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
R Development Core Team. R: a Language and Environment for Statistical Computing (Vienna, Austria, 2012).
Acknowledgements
We thank L. Cochella and members of the Stark group for comments on the manuscript and Life Science Editors (http://lifescienceeditors.com) for editorial support. We are grateful to P. Heine and E. Jans (MaxCyte) for help setting up efficient plasmid transfection. Deep sequencing was performed at the Vienna Biocenter Core Facilities GmbH (VBCF) Next-Generation Sequencing Unit (http://vbcf.ac.at). The Stark group is supported by the European Research Council (ERC) under the European Union's Horizon 2020 research and innovation programme (grant agreement no. 647320) and by the Austrian Science Fund (FWF, F4303-B09). Basic research at the IMP is supported by Boehringer Ingelheim GmbH and the Austrian Research Promotion Agency (FFG).
Author information
Authors and Affiliations
Contributions
C.D.A., M.A.Z., and A.S. conceived the project. C.D.A., M.P., and M.R. performed the experiments with the help of K.S., and M.A.Z. the computational analyses. T.K. performed the k-mer based predictions. C.D.A., M.A.Z., and A.S. wrote the manuscript. A.S. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Candidate fragment length distribution.
(a) Candidate fragment length distributions of STAP-seqzfh1 and (b) STAP-seqctrl input fragments in bins of 10bp (40 bins in total).
Supplementary Figure 2 Genomic distribution of eTSSs and agreement of eTSS positions with endogenous TSSs and established sequence motifs.
(a) Pie charts of the genomic distribution of all genomic sequences that initiate transcription with tag counts >=5, >=10, >=20, respectively, in response to the zfh1 enhancer compared to the D. melanogaster genome (leftmost pie chart; the sector indicates aTSSs ±50bp that make up 0.32% of the genome). (b) Bar plots visualizing the enrichment of the regions from a, over the genome. (c) Metagene profile of average normalized STAP-seqzfh1 tag counts around aTSSs, including the 5th and 95th percentiles determined by bootstrapping. (d) Metagene profiles of STAP-seqzfh1, scRNA-seq18 and PEAT16 at all eTSSs. (e) As in d, but specifically for proximal, distal, within coding DNA sequence (CDS) and intronic eTSSs. (f) Agreement of STAP-seq eTSSs and embryo-derived TSSs by PEAT16 that are shifted from aTSSs between 1 to 10 nucleotides (each row is scaled to the respective maximum; see Figure 1e for the equivalent comparison with scRNA-seq from S2 cells15). (g) Sequence-logos depicting position-specific nucleotide frequencies for eTSSs that are aTSS-proximal or -distal, within CDS, or intronic. (h) Sequence logos of eTSSs that coincide with aTSS (shift 0, top row, the +1 position of aTSSs is indicated by the arrow) or are proximally-misaligned by 1 to 10 base pairs (bps) from aTSSs (logos are moved closer together but not scaled differently relatively to each other).
Supplementary Figure 3 Reproducibility of STAP-seqzfh1 and comparisons between STAP-seq screens with different enhancers.
(a) Scatterplots comparing focused STAP-seqzfh1 screens with different screens with diverse developmental (dev; top) and housekeeping enhancers (hk; bottom; see Figure 2 for further comparisons; PCC: Pearson correlation coefficient). (b) Scatterplot depicting the similarity between focused and genome-wide STAP-seqzfh1 screens. (c) Scatterplot for two independent biological replicates of focused STAP-seqzfh1 screens, including standard deviations (s.d) calculated across three independent biological replicates (error bars).
Supplementary Figure 4 Induced activities are consistent across cell types.
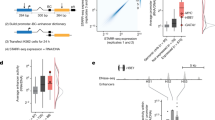
Scatterplot depicting STAP-seq tag counts for STAP-seqzfh1 in S2 cells (x-axis) versus STAP-seqtj in OSCs (y-axis) and their similarity (expressed as PCC: Pearson correlation coefficient). TSSs that are endogenously – as measured by GRO-seq19,20 – and exclusively active in S2 cells or OSCs are labeled blue or red, respectively. Aligned to each axis are the respective cumulative distributions used to assess the difference between TSSs that are endogenously exclusively active in either S2 cells or OSCs by Kolmogorov–Smirnov tests (P values indicated). The scatter plot corresponds to Figure 3a.
Supplementary Figure 5 STAP-seq is complementary to methods that assess endogenous transcription initiation.
(a) Venn diagram depicting the overlap of aTSSs detected by STAP-seqzfh1 and scRNA-seq in S2 cells genome-wide (see Figures 3d and e for an equivalent analysis of the focused STAP-seq screens). (b) Proportion of aTSSs uniquely detected by STAP-seqzfh1 that are also detected during different developmental stages by RAMPAGE44 (left bar) or are in closed chromatin in S2 cells (right bar). (c) aTSSs uniquely detected by scRNA-seq that contain housekeeping core promoter motifs. (d) Gene Ontology analysis of aTSSs uniquely detected by either STAP-seqzfh1 or scRNA-seq. The results from b-d suggest that aTSSs uniquely detected by scRNA-seq are housekeeping-type core promoters (see also Figures 3e and f), while aTSSs uniquely detected by STAP-seqzfh1 are endogenously not active in S2 cells (b, right bar) but in other cell types (b, left bar).
Supplementary Figure 6 A wide range of enhancer-responsiveness.
(a) Scatterplots depicting the range of enhancer-responsiveness as determined by STAP-seqzfh1 over STAP-seqctrl at eTSSs for replicate (rep) 1 versus 2. The distributions of enhancer-responsiveness at the respective TSSs (brown) and at random positions (grey) are shown by density plots along the axes. (b) As in a, but, at all, position-corrected, and TATA box, Initiator (Inr), MTE or DPE-containing and position-corrected aTSSs. (c) TSS strengths are independent of candidate lengths. Boxplots depicting maximal length of candidate STAP-seqzfh1 fragments intersecting positions with the corresponding STAP-seqzfh1 tag counts (left) or eTSSs of different enhancer-responsiveness (middle), and ranks of eTSSs which +1 positions intersect candidate STAP-seqzfh1 fragments of different lengths (right). Center line: median; limits: interquartile range; whiskers: 5th and 95th percentiles. (d) Housekeeping enhancers activate transcription from candidate sequences at a reduced dynamic range compared to developmental enhancers. Scatterplots depicting the range of enhancer-responsiveness of the indicated focused STAP-seq screens at aTSSs in genomic regions covered by the focused candidate libraries for developmental (left and middle) and housekeeping (right) enhancers. To account for the broad nature of initiation at housekeeping core promoters, enhancer-responsiveness in d, is calculated in a window of ±50bp around aTSSs.
Supplementary Figure 7 Biological significance and sequence properties of eTSSs with different enhancer-responsiveness.
(a) Gene Ontology analysis of the top and bottom 1,000 genes associated with the strongest and weakest eTSSs, respectively (see Figure 4f for an equivalent analysis restricted to eTSSs containing exclusively TATA box, Inr, MTE or DPE). (b-c) Enrichment of single (b) or combinations (c) of core promoter motifs in eTSSs of different ranks compared to random genomic sequences. As has been observed before17, the combination of TATA box and DPE motif is less strongly enriched compared to the TATA box and Inr or Inr and DPE motif pairs and the combination of all three motifs rarely occurs in the same eTSS. (d) Boxplots depicting motif deviation (distance in base pairs, bp) from their consensus positions (determined as the average position across all eTSSs) versus enhancer-responsiveness of eTSSs. Center line: median; limits: interquartile range; whiskers: 5th and 95th percentiles (e) Heatmaps depicting positional occurrences of core promoter motifs around eTSSs. Note that TATA boxes are somewhat less positionally constraint, as previously observed for mammalian core promoters26.
Supplementary Figure 8 Induced and basal activities measured by STAP-seqzfh1 and STAP-seqctrl, respectively, reveal low basal activities for top eTSSs.
(a) Histograms depicting distributions of normalized STAP-seqzfh1 and STAP-seqctrl tag counts at positions that are covered by at least 1 tag (left) or the top 1,000 STAP-seqzfh1 positions (right). The inset on the left shows that STAP-seqzfh1 but not STAP-seqctrl reaches high normalized tag counts. (b) Representative screenshots of the top three eTSSs, depicting high STAP-seqzfh1 and low STAP-seqctrl tag counts.
Supplementary Figure 9 Sequences with the highest basal activities are at housekeeping genes and overlap housekeeping-type enhancers.
(a) Core promoter motif enrichment analysis of candidate sequences with the highest basal (STAP-seqctrl) versus the highest induced (STAP-seqzfh1) activities (top 500 each). NS: not significant. (b) Gene Ontology (GO) analysis for genes associated with eTSSs that show the highest basal (STAP-seqctrl, right) versus the highest induced (STAP-seqzfh1, left) activities. (c) Fraction of candidates as in a, that show STARR-seq enrichment values of at least 3-fold. Developmental and housekeeping STARR-seq data are from ref. 9. (d) As in c, but plotting average STARR-seq signals around the most prominent, or the center of dispersed TSSs.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 1734 kb)
Supplementary Table 1
Enhancers used for STAP-seq screens. Genomic coordinates (dm3) and cloned sequences of S2 cell developmental (zfh1, sgl, ham), OSC developmental (tj) and housekeeping (ncm, ssp3) enhancers used for the respective STAP-seq screens. (XLSX 10 kb)
Supplementary Table 2
BACs that are contained in the focused libraries. Indicated are the coordinates and IDs of the BACs that were used to generate the focused STAP-seq libraries. (XLSX 9 kb)
Supplementary Table 3
Primers used for amplification of the D. pseudoobscura sequences (spike-in controls). Indicated are the primers used to amplify the sequences from the D. pseudoobscura genome, which were used to generate the STAP-seq spike-in control plasmids. (XLSX 9 kb)
Supplementary Table 4
Details on individual candidates for luciferase validations. Primer pairs used to amplify candidates for luciferase validation as well as their genomic coordinates, the luciferase fold change, standard deviations and enhancer-responsiveness are indicated. (XLSX 15 kb)
Supplementary Table 5
Number of mapped reads and eTSSs for STAP-seq screens. Reported are total mapped reads and collapsed fragments (see Methods) for all STAP-seq screens (dm3) and the respective D. pseudoobscura spike-in controls (dp3). (XLSX 9 kb)
Rights and permissions
About this article
Cite this article
Arnold, C., Zabidi, M., Pagani, M. et al. Genome-wide assessment of sequence-intrinsic enhancer responsiveness at single-base-pair resolution. Nat Biotechnol 35, 136–144 (2017). https://doi.org/10.1038/nbt.3739
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3739
This article is cited by
-
Leveraging massively parallel reporter assays for evolutionary questions
Genome Biology (2023)
-
Using Synthetic DNA Libraries to Investigate Chromatin and Gene Regulation
Chromosoma (2023)
-
DeepSTARR predicts enhancer activity from DNA sequence and enables the de novo design of synthetic enhancers
Nature Genetics (2022)
-
Differential cofactor dependencies define distinct types of human enhancers
Nature (2022)
-
Compatibility rules of human enhancer and promoter sequences
Nature (2022)