Abstract
Whitefly (Bemisia tabaci) damages field crops by sucking sap and transmitting viral diseases. None of the insecticidal proteins used in genetically modified (GM) crop plants to date are effective against whitefly. We report the identification of a protein (Tma12) from an edible fern, Tectaria macrodonta (Fee) C. Chr., that is insecticidal to whitefly (median lethal concentration = 1.49 μg/ml in in vitro feeding assays) and interferes with its life cycle at sublethal doses. Transgenic cotton lines that express Tma12 at ∼0.01% of total soluble leaf protein were resistant to whitefly infestation in contained field trials, with no detectable yield penalty. The transgenic cotton lines were also protected from whitefly-borne cotton leaf curl viral disease. Rats fed Tma12 showed no detectable histological or biochemical changes, and this, together with the predicted absence of allergenic domains in Tma12, indicates that Tma12 might be well suited for deployment in GM crops to control whitefly and the viruses it carries.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Sanahuja, G., Banakar, R., Twyman, R.M., Capell, T. & Christou, P. Bacillus thuringiensis: a century of research, development and commercial applications. Plant Biotechnol. J. 9, 283–300 (2011).
Chougule, N.P. & Bonning, B.C. Toxins for transgenic resistance to hemipteran pests. Toxins (Basel) 4, 405–429 (2012).
Dubey, A.K. & Ko, C.C. Whitefly (Aleyrodidae) host plants list from India. Orient. Insects 42, 49–102 (2008).
Legg, J. in Whitefly and Whitefly-borne Viruses in the Tropics: Building a Knowledge Base for Global Action (eds. Anderson, P.K. & Morales, F.J.) 15–23 (CIAT Publications, 2005).
Navas-Castillo, J., Fiallo-Olivé, E. & Sánchez-Campos, S. Emerging virus diseases transmitted by whiteflies. Annu. Rev. Phytopathol. 49, 219–248 (2011).
Horowitz, A.R., Antignus, Y. & Gerling, D. in The Whitefly, Bemisia tabaci (Homoptera: Aleyrodidae) Interaction with Geminivirus-Infected Host Plants. (ed. Thompson, W.M.O.) 293–322 (Springer, 2011).
Bergé, J.B. & Ricroch, A.E. Emergence of minor pests becoming major pests in GE cotton in China: what are the reasons? What are the alternatives practices to this change of status? GM Crops 1, 214–219 (2010).
Das, S. et al. A mannose binding lectin from leaves of Allium sativum effective against whitefly, and process for its preparation. Indian patent 228783 (2009).
Jin, S., Zhang, X. & Daniell, H. Pinellia ternata agglutinin expression in chloroplasts confers broad spectrum resistance against aphid, whitefly, Lepidopteran insects, bacterial and viral pathogens. Plant Biotechnol. J. 10, 313–327 (2012).
Thakur, N. et al. Enhanced whitefly resistance in transgenic tobacco plants expressing double stranded RNA of V-ATPase A gene. PLoS One 9, e87235 (2014).
Doorn, A.V. & Vos, M.D. Resistance to sap-sucking insects in modern-day agriculture. Front. Plant Sci. 4, 222 (2013).
Hendrix, S.D. An evolutionary and ecological perspective of the insect fauna of ferns. Am. Nat. 115, 171–196 (1980).
Markham, K., Chalk, T. & Stewart, C.N. Jr. Evaluation of fern and moss protein based defenses against phytophagous insects. Int. J. Plant Sci. 167, 111–117 (2006).
Balasubramanian, R., Selvaraj, P. & Sahayaraj, K. Partial purification and characterization of phytoecdysone from Christella parasitica (L.) and screening its pesticidal properties on lepidopteran pests. J. Biopesticides 1, 201–205 (2008).
Sahayaraj, K. & Selvaraj, P. Field evaluation of three fern extracts on Helicoverpa armigera (Hubner) and Spodoptera litura (Fab.) incidence and infestation and groundnut production. Legume Res. 36, 84–86 (2013).
Petterino, C. & Argentino-Storino, A. Clinical chemistry and haematology historical data in control Sprague-Dawley rats from pre-clinical toxicity studies. Exp. Toxicol. Pathol. 57, 213–219 (2006).
Bonning, B.C. et al. Toxin delivery by the coat protein of an aphid-vectored plant virus provides plant resistance to aphids. Nat. Biotechnol. 32, 102–205 (2013).
Uprety, Y., Poudel, R.C., Asselin, H. & Boon, E. Plant biodiversity and ethnobotany inside the projected impact area of the Upper Seti Hydropower Project, Western Nepal. Environ. Dev. Sustain. 13, 463–492 (2011).
Debendra, S.D. Indigenous vegetables of Nepal for biodiversity and food security. Int. J. Biodivers. Conserv. 5, 98–108 (2013).
Nwosu, M.O. Ethnobotanical studies on some pteridophytes of southern Nigeria. Econ. Bot. 56, 255–259 (2002).
Upadhyay, S.K. et al. RNA interference for the control of whiteflies (Bemisia tabaci) by oral route. J. Biosci. 36, 153–161 (2011).
Upadhyay, S.K. et al. Interaction of Allium sativum leaf agglutinin with midgut brush border membrane vesicles proteins and its stability in Helicoverpa armigera. Proteomics 10, 4431–4440 (2010).
Kumar, M., Shukla, A.K., Singh, H. & Tuli, R. Development of insect resistant transgenic cotton lines expressing cry1EC gene from an insect bite and wound inducible promoter. J. Biotechnol. 140, 143–148 (2009).
Sharma, H.C. Biotechnological Approaches for Pest Management and Ecological Sustainability (CRC Press, 2009).
Kumar, J., Kumar, A., Roy, J.K., Tuli, R. & Khan, J.A. Identification and molecular characterization of begomovirus and associated satellite DNA molecules infecting Cyamopsis tetragonoloba. Virus Genes 41, 118–125 (2010).
Stewart, L. & Rossi, J. Using Cotton Byproducts in Beef Cattle Diets (Bulletin 1311) (University of Georgia Cooperative Extension, 2010).
Petterino, C. & Argentino-Storino, A. Clinical chemistry and haematology historical data in control Sprague-Dawley rats from pre-clinical toxicity studies. Exp. Toxicol. Pathol. 57, 213–219 (2006).
Fu, T.J., Abbott, U.R. & Hatzos, C. Digestibility of food allergens and nonallergenic proteins in simulated gastric fluid and simulated intestinal fluid—a comparative study. J. Agric. Food Chem. 50, 7154–7160 (2002).
Acknowledgements
The authors acknowledge H.K. Yadav, R. Kamal and S. Sharma for help in statistical analysis; M. Lal, Rajesh K. Srivastava, S.H.M. Abidi, A. Bano (deceased) and Rakesh K. Srivastava for technical assistance; and A.C. Little for photography. The authors thank the Council of Scientific and Industrial Research, India, for funding under the Supra Institutional Project (SIP05), the Network Projects (NWP03 and PlaGen BSC0107) and the Council of Scientific and Industrial Research–National Botanical Research Institute for providing research facilities. A.K.S., S.K.U., R.S., M.M., S. Saurabh, H.S., N.T., P.P., A.L.H. and V.R. are supported by CSIR. P.R. and S.K.Y. are supported by the Indian Council of Medical Research. S. Srivastava is supported by the University Grant Commission. R.T. is supported by a Department of Science and Technology (DST) JC Bose National Fellowship.
Author information
Authors and Affiliations
Contributions
A.K.S.: development of cotton transformation protocols, transgenic cotton lines and acquisition of data. S.K.U.: purification of Tma12 and characterization, cloning of tma12. M.M.: MS analysis of Tma12, study on whitefly life cycle on transgenic cotton in clip cage and manuscript preparation. S. Saurabh, R.S., H.S., P.R.: bioprospection of ferns for insecticidal activity, establishment of screening protocol, identification of the potential ferns, execution of experiments and manuscript preparation. N.T.: clip cage assay, analysis of transgenic cotton lines and manuscript preparation. S. Srivastava: analysis of transgenic cotton lines and manuscript preparation. A.L.H., P.P.: study of whitefly life cycle on transgenic cotton in the field. V.R., M.K.S.: evaluation of transgenic cotton in field trials. S.K.Y.: chitin-binding assay, analysis of impact of Tma12 on whitefly at sublethal doses. J.K.: virus studies. K.C.: designing and execution of whitefly bioassays and statistical analysis. P.C.V.: designing and supervision of whitefly life cycle study on transgenic cotton and transgenic cotton development. A.P.S.: collection, identification, mass multiplication and in-house maintenance of ferns. K.N.N.: review of experimental design and writing of manuscript. S.B., M.W., S. Singh, S. Sharma: biosafety experiments. O.: designing and execution of experiments on beneficial insect ladybird beetle. R.S.U.: supervised A.K.S. in cotton tissue culture and field evaluation of transgenic cotton. S.A.R.: critical review of manuscript. R.T.: critical review of results, intellectual input and manuscript editing. P.K.S.: study concept and experimental design, data analysis, study supervision and manuscript writing and editing.
Corresponding author
Ethics declarations
Competing interests
A patent application (WO 2013098858A2; P.K.S., S.K.U., K.C., S. Saurabh, R.S., P.R., H.S., M.M., A.P.S., P.C.V., K.N.N. and R.T. are inventors) based on the Tma12-GM crop strategy for field control of whiteflies is under examination. A.K.S., S.K.U., M.M., S. Saurabh, R.S., H.S., N.T., P.R., P.P., A.L.H., S. Srivastava, V.R., S.K.Y., M.K.S., K.C., P.C.V., A.P.S., K.N.N., R.T., P.K.S., CSIR–National Botanical Research Institute and Council of Scientific and Industrial Research (the funding agency), Ministry of Science and Technology, Government of India, hope to translate the technology jointly with industry to develop whitefly-resistant GM crops.
Integrated supplementary information
Supplementary Figure 1 Overview of the study.
Schematic representation of key steps in the strategy for prospecting ferns for insecticidal proteins and development of transgenic cotton.
Supplementary Figure 2 N-terminal sequencing of Tma12.
Detection of 1st to 12th amino acid sequence ‘HGSMEDPISRVY’ in Tma12 is shown in b to m. Arrows show the probable peak of the identified amino acid residue. The amino acid standards are shown in a. The sequencing was done by Proteomics International, Australia.
Supplementary Figure 3 De novo sequencing of peptides of Tma12.
Schematic representation of fragmentation chemistry of peptides before and after derivatization on MALDI TOF/TOF. Panel (a) represents simultaneous fragmentation of the peptide at both the ends generating b and y ions that made the spectra condensed. The data was analyzed in-silico (DeNovo Explorertm Software) and the top 10 sequences were predicted, as listed above. Panel (b) represents the strategy followed to improve de novo sequencing. The peptide was modified at N-terminus (SPITC modification) and then fragmented. The modification prevented all the b ions (generated from N-terminus fragmentation) from entering into drift tube. This allowed y-ions to produce a clear spectrum with high signal to noise ratio. The series of ions was analyzed manually and amino acid sequence of the peptide was deduced.
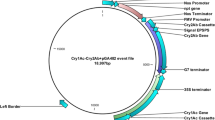
Supplementary Figure 4 Architecture of insecticidal protein-encoding gene.
(a) Structure of tma12 gene. (b) Complete ORF of Tma12 showing signal peptide of 24 amino acids (red fonts) and mature protein of 192 amino acid residues. Protein sequences shown in blue fonts are the three peptides sequenced through N-terminal and de novo sequencing. (c) Conserved domain search of Tma12 showing the best match with bacterial chitin binding domain 3 protein of Herpetosiphon aurantiacus (GI:159899166) with similarity score 55.
Supplementary Figure 5 Characterization of Tma12.
(a) Chitin binding assay and SDS-PAGE analysis showing binding of Tma12 on chitin magnetic beads and its elution. (b) Thermal stability of Tma12; enzymatic activity after incubation at different temperatures for 30 min. Both exo and endo-chitinase activities started declining sharply at 50oC and were lost at 60oC. Chitinolytic activity of Tma12, Michaelis-Menten and Lineweaver-Burk plot are shown for (c) colloidal chitin (d) 4-methylumbelliferyl β-D-N, N’-diacetylchitobioside hydrate and (e) 4-methylumbelliferyl β-D-N, N’, N’’-triacetylchitotriose.
Supplementary Figure 6 Development of putative transgenic cotton lines, insect resistance and generation advancement.
A total of 16 transgenic plants were developed, 9 of them exhibited high resistance against whitefly. However, eight plants shown in top panels did not flower. Four other plants (left lower panel) developed flowers but did not advance into next generation. Four plants (right lower panel) produced bolls and advanced to next generation. The transgenic line number is given on the pots. Scale bars,10 cm.
Supplementary Figure 7 Expression analysis of transgene in event 14.
(a) qRT-PCR of different parts of transgenic plant, the expression of tma12 in leaf tissue is considered as unity and relative expression of tma12 in all other plant parts is plotted in the graph. The relative expression of tma12 was highest in root followed by stem, fruit wall, leaf, petal, anthers and square. (b) Immuno-blot analysis of the different parts of Event14. Tma12 was purified from stem, leaf, root and seed of the transgenic plant and quantified using purified Tma12 as standard. The expression of Tma12 was reasonably well in leaves, moderate in stem and barely detectable in seeds. There was no detectable expression in root despite of high transcript level.
Supplementary Figure 8 Whitefly resistance in transgenic cotton prevents infection by CLCuV.
Detection of the cotton leaf curl virus (type CLCuKoV-Bu) by PCR. Presence of the virus is seen in non transgenic cotton plants, with the PCR product of ∼2.8 kb DNA representing the viral genome. (b) Amplification of ∼1.3 kb alpha satellite DNA and (c) the ∼1.3 kb beta satellite DNA, associated with CLCuV from non-transgenic cotton plants. (d-f) Transgenic cotton plants showed no amplification of the genome of CLCuV, the alpha satellite or the beta satellite. M is λ DNA/HindIII-EcoRI digested molecular weight markers.
Supplementary Figure 9 Effect of Tma12-transgenic cotton on Helicoverpa armigera.
Leaves of transgenic and control cotton plants were challenged with neonatal larvae of H. armigera. Insect larvae feed on both the leaves. This indicated that Tma12-cotton is not efficacious to H. armigera. Scale bar, 2.5 cm.
Supplementary Figure 10 Effect of Tma12-transgenic cotton on nontarget sucking insect pests.
(a) 4th instar aphids were inoculated on transgenic or control plants (20 per plant) and the population was recorded on 9 plants at days 7 and 22. Data shown as mean ± SD per plant. No significant difference in aphid population was observed between control and transgenic cotton plants (P value >0.05). (b) Representative photograph of transgenic and control leaves infested with aphids. Scale bars, 2.5 cm. (c) Three leaves of transgenic or control cotton plants were challenged with six mature female mealybugs (2 per leaf). The insects were counted on days 6 and 24. Data shown as mean ± SD per plant. (d) Representative photograph of transgenic and control leaves infested with mealybugs. Insect population increased rapidly till day 6, while decreased and stabilized on day 24. Scale bars, 2.5 cm.
Supplementary Figure 11 Field evaluation of Tma12 transgenic cotton in T1 and T2.
(a) Transgenic and (b) Nontransgenic cotton plants in T1 generation. (c) Transgenic and (d) Nontransgenic cotton plants in T2 generation. Scale bars, 1 m.
Supplementary Figure 12 Evaluation of Tma12 transgenic cotton in contained field trials.
(a) Transgenic (b) Nontransgenic and (c) Pesticide-protected nontransgenic cotton plants in T3 generation. (d) Nontransgenic and (e) transgenic cotton plants in T4 generation. Scale bars for a, b & c; 0.5 m and d & e, 1 m.
Supplementary Figure 13 Pepsin sensitivity assay.
SDS-PAGE showing time course digestion of Tma12; the insecticidal protein was digested completely, within 30 sec of incubation with pepsin.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Tables 1–7 and Supplementary Note 1 (PDF 3754 kb)
Rights and permissions
About this article
Cite this article
Shukla, A., Upadhyay, S., Mishra, M. et al. Expression of an insecticidal fern protein in cotton protects against whitefly. Nat Biotechnol 34, 1046–1051 (2016). https://doi.org/10.1038/nbt.3665
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3665
This article is cited by
-
Foliar application of clay-delivered RNA interference for whitefly control
Nature Plants (2022)
-
Advanced genome editing strategies for manipulation of plant specialized metabolites pertaining to biofortification
Phytochemistry Reviews (2022)
-
Dynamic genome evolution in a model fern
Nature Plants (2022)