Abstract
The amount of sequence information in public repositories is growing at a rapid rate. Although these data are likely to contain clinically important information that has not yet been uncovered, our ability to effectively mine these repositories is limited. Here we introduce Sequence Bloom Trees (SBTs), a method for querying thousands of short-read sequencing experiments by sequence, 162 times faster than existing approaches. The approach searches large data archives for all experiments that involve a given sequence. We use SBTs to search 2,652 human blood, breast and brain RNA-seq experiments for all 214,293 known transcripts in under 4 days using less than 239 MB of RAM and a single CPU. Searching sequence archives at this scale and in this time frame is currently not possible using existing tools.
Similar content being viewed by others
Main
The National Institutes of Health (NIH) Sequence Read Archive (SRA)1 contains ∼3 petabases of sequence information that can be used to answer biological questions that single experiments do not have the power to address. However, searching the entirety of such a database for a sequence has not been possible in reasonable computational time.
Some progress has been made toward enabling sequence searches on large databases. The NIH SRA provides a sequence search functionality2; however, the search is restricted to a limited number of experiments. Existing full-text indexing data structures such as Burrows-Wheeler transform3, FM-index4 or others5,6,7 are currently unable to mine data of this scale. Word-based indices8,9, such as those used by internet search engines, are not appropriate for edit-distance-based biological sequence searches. The sequence-specific solution caBLAST and its variants10,11,12 require an index of known genomes, genes or proteins, and so cannot search for novel sequences. Further, none of these existing approaches are able to match a query sequence q that spans many short reads.
Here, we use an indexing data structure, Sequence Bloom Tree (SBT), to identify all experiments in a database that contain a given query sequence q. A query is an arbitrary sequence, such as a transcript. The SBT index is independent of eventual queries, so the approach is not limited to searching for known sequences, and the index can be efficiently built and stored in limited additional space. It also does not require retaining the original sequence files and can be distributed separately from the data. SBTs are dynamic, allowing insertions and deletions of new experiments. A coarse-grained version of an SBT can be downloaded and subsequently refined as more specific results are needed. They can be searched using low memory for the existence of arbitrary query sequences. We show that SBTs can search large collections of RNA-seq experiments for a given transcript orders of magnitude faster than existing approaches.
Results
Application of SBT to sequence searching
SBTs create a hierarchy of compressed bloom filters13,14, which efficiently store a set of items. Each bloom filter contains the set of k-mers (length-k subsequences) present within a subset of the sequencing experiments. SBTs are binary trees in which the sequencing experiments are associated with leaves, and each node v of the SBT contains a bloom filter that contains the set of k-mers present in any read in any experiment in the subtree rooted at v (Supplementary Fig. 1). We reduced the space usage by using bloom filters that are compressed by the RRR15 compression scheme (Online Methods). Hierarchies of bloom filters have been used for data management on distributed systems16. However, they have not previously been applied to sequence search, and we find that this allows us to tune the bloom filter error rate much higher than in other contexts (Theorem 2, Online Methods), vastly reducing the space requirements. Bloom filters have also been used for storing implicit de Bruijn graphs17,18, and one view of SBTs is as a generalization of this to multiple graphs.
We used SBTs to search RNA-seq experiments for expressed isoforms. We built an SBT on 2,652 RNA-seq experiments in the SRA for human blood, breast and brain tissues (Supplementary Table 1). The entire SBT required only 200 GB (2.3% of the size of the original sequencing data) (Supplementary Table 2). For these data, construction of the tree took ≈2.5 min per file (Supplementary Table 3).
These experiments could be searched for a single transcript query in, on average, 20 min (Fig. 1), using less than 239 MB of RAM with a single thread (Online Methods). We estimate the comparable search time using SRA-BLAST2 or mapping by STAR19 to be 2.2 d and 921 d, respectively (Online Methods), though SRA-BLAST and STAR return alignments whereas SBT does not. However, even a very fast aligner such as STAR cannot identify query-containing experiments as fast as SBT. We also tested batches of 100 queries and found SBT was an estimated 4,056 times faster than a batched version of the mapping approach (Supplementary Fig. 2). These queries were performed over varying sensitivity threshold θ (the minimum fraction of query k-mers that must exist in order to return a 'hit') as well as the transcripts per million (TPM) threshold used to select the query set (Supplementary Figs. 3 and 4). For approximately half of the queries, the upper levels of the SBT hierarchy provided substantial benefit, particularly on queries that were not expressed in any experiment (Supplementary Fig. 5 and Supplementary Table 4).
SBTs can speed up existing algorithms
SBTs can speed up the use of algorithms, such as STAR or SRA-BLAST, by first ruling out experiments in which the query sequences are not present. This allows the subsequent processing time to scale with the size of the number of hits rather than the size of the database. We first used SBTs to filter the full dataset consisting of 2,652 human blood, breast and brain RNA-seq experiments. We then compared the performance of STAR or SRA-BLAST on the filtered dataset with the time to process the unfiltered dataset with these algorithms. Using SBTs to first filter the data reduced the overall query time of STAR or SRA-BLAST by a factor of ≈3 (Supplementary Fig. 6).
Measuring the performance of SBT
To analyze the accuracy of the SBT filter, we compared the experiments returned by SBT with those in which the query sequence was estimated to be expressed using Sailfish20. Because it is impractical to use existing tools to estimate expression over the entire set of experiments, we queried the entire tree, but estimated accuracy on a set of 100 random files on which we ran Sailfish (Fig. 2). Three collections of representative queries were constructed using Sailfish, denoted by High, Medium and Low, which included transcripts of length >1,000 nt that were likely to be expressed at a higher, medium or low level in at least one experiment contained in the set of 100 experiments on which Sailfish was run. The High set was chosen to be 100 random transcripts with an estimated abundance of >1,000 TPM in at least one experiment. The Medium and Low query sets were similarly chosen randomly from among transcripts with >500 and >100 TPM, respectively. These Sailfish estimates were taken as the ground truth of expression for the query transcripts.
Both false positives and false negatives can arise from a mismatch between SBT's definition of present (coverage of k-mers over a sufficient fraction of the query) and Sailfish's definition of expressed (as estimated by read mapping and an expectation-maximization inference). These two definitions are related, but not perfectly aligned, resulting in some disagreement that is quantified by the false-positive rates (FPR) and false-negative rates of Figure 2. The observed false negatives are primarily driven by a few outlier queries for which the SBT reports no results but their expression is above the TPM threshold as estimated by Sailfish. This is supported by the fact that the average true-positive rate at θ = 0.7 for queries that return at least one file was 96–100%, and the median true-positive rate across all queries was 100% for all but the strictest θ (Fig. 2).
Discussion
We used SBT to search all blood, brain and breast SRA sequencing runs for the expression of all 214,293 known human transcripts and used these results to identify tissue-specific transcripts (Supplementary Table 5 and Supplementary Fig. 7). This search took 3.3 d using a single thread (Supplementary Fig. 8). There are presently no search or alignment tools that can solve this scale of sequence search problem in a reasonable time frame, but we estimate an equivalent search using Sailfish would take 92 d. The speed and computational efficiency of SBTs will enable both individual laboratories and sequencing centers to support large-scale sequence searches, not just for RNA-seq data, but for genomic and metagenomic collections as well. Researchers could search for conditions from among thousands that are likely to express a given novel isoform or use SBTs to identify metagenomic samples that are likely to contain a particular strain of bacteria. Fast search of this type will be essential to make good use of the ever-growing collection of available sequencing data.
Currently, it is difficult to access all the relevant data relating to a particular research question from available sequencing experiments. Individual hospitals, sequencing centers, research consortia and research groups are collecting data at a rapid pace, and face the same difficulty of not being able to test computational hypotheses quickly or to find the relevant conditions for further study. SBTs enable the efficient mining of these data and could be used to uncover biological insights that can be revealed only through the analysis of multiple data sets from different sources. Furthermore, SBTs do not require prior knowledge about sequences of interest, making it possible to identify, for example, the expression of unknown isoforms or long noncoding RNAs. This algorithm makes it practical to search large sequencing repositories and may open up new uses for these rich collections of data.
Methods
Data Availability.
An open-source prototype implementation of SBT is available at http://www.cs.cmu.edu/~ckingsf/software/bloomtree (Supplementary Software). Testing and analysis scripts, along with their inputs and outputs, are available at https://github.com/Kingsford-Group/sbtappendix.
SBT construction and insertion.
A SBT is a binary tree that is built by repeated insertion of sequencing experiments. Given a (possibly empty) SBT T, a new sequencing experiment s can be inserted into T by first computing the bloom filter b(s) of the k-mers present in s and then walking from the root along a path to the leaves and inserting s at the bottom of T in the following way. When at node u, if u has a single child, a node representing s (and containing b(s)) is inserted as u's second child. If u has two children, b(s) is compared against the bloom filters b(left(u)) and b(right(u)) of the left left(u) and right right(u) children of u. The child with the more similar filter under the Hamming distance between the filters becomes the current node, and the process is repeated. If u has no children, u represents a sequencing experiment s′. In this case, a new union node v is created as a child of u's parent. This new node has two children: u and a new node representing s.
Each filter consists of a bit vector of length m and a set of h hash functions h1 : U → [0, m) that map items to bits in the bit vector. Insertion of k ∈ U is performed by setting to 1 the bits specified by hi(k) for i = 1, …, h. Querying for membership of k in b(k) checks these same bits; if they are all 1, the filter is reported to contain k. Because of overlapping hash results, bloom filters have one-sided error: they can report a k-mer k is present when it is not. This error, and its effect on overall query accuracy of SBTs, can be made quite small with the appropriate choice of parameters (see below). Bloom filters have been used in several others contexts in bioinformatics (e.g. refs. 21,22). Hierarchies of Bloom filters have been used in other applications23.
As s is walked down the tree, the filters at the nodes that are visited are unioned with b(s). This unioning process can be made fast (and trivially parallelized for large filters) because the union of two bloom filters can be computed by computing a bit vector with each bit set to 1 exactly when the corresponding bit in either of the two bloom filters is 1. This is particularly beneficial where graphics processing units or vector computations can be used for these single instruction, multiple data (SIMD) operations. SBTs are different than cascading bloom filters24,25, which aim to reduce false-positive rates of a single set query by recursively storing false positives in their own bloom filters. SBT works when word based indices fail26,27.
The insertion process is designed to greedily group together sequencing experiments with similar bloom filters. This is important for two reasons. First, it helps to mitigate the problem of filter saturation. If too many dissimilar experiments are present under a node u, then b(u) tends to have many bits set. In addition, by placing similar experiments in similar subtrees, more subtrees are pruned at an earlier stage of a query, reducing query time.
A primary challenge with scaling SBTs to terabytes of sequence is saturation of the filters at levels of the tree near the root. The filter at any node v is the union of the filters of its children. However, this means as one moves from the leaves to the root, the filters will tend to contain more and more bits set to 1, increasing their false-positive rate. This saturation can be overcome using several techniques: appropriate parameter selection (see “Setting the bloom filter size”), grouping of related experiments during insertion into the tree as above and including only k-mers that have a minimum coverage count (see “Building bloom filters”). Note that filters with poor false-positive rates at high levels of the tree only affect query time: accuracy is governed entirely by the false-positive rate of the leaf filters.
Querying.
Given a query sequence q and a SBT T, the sequencing experiments (at the leaves) that contain q can be found by breaking q into its constituent set of k-mers Kq and then flowing these k-mers over T starting from the root. At each node u, the bloom filter b(u) at that node is queried for each of the k-mers in Kq. If more than θ|Kq| k-mers are reported to be present in b(u), the search proceeds to all of the children of u, where θ is a cutoff between 0 and 1 governing the stringency required of the match. The parameter θ governs a query's tolerance to errors. Ignoring the effects of sequence boundaries, a general SBT query with N k-mers and k-mer size k tolerates at least N (1 − θ)/k k-mer mismatches, between the query and the stored data.
If fewer than that number of k-mers are present, the subtree rooted at u is not searched further (it is pruned). It has been shown that k-mer similarity is highly correlated to the quality of the alignments between sequences28,29,30,31, and SBT guarantees that if the query sequence is present (at sufficient coverage), it will be found.
When a search proceeds to the children, the children are added to a queue for eventual processing. Even though there may be a large frontier of nodes that are currently active, the memory usage for querying is the trivial amount of memory needed to store the tree topology plus the memory needed to store the single current filter. The SBT timings reported here are all for single-threaded operation.
If several queries are to be made, they can be batched together so that a collection C = {Kq1, …, Kqt} of queries starts at the root, and only queries for which |b(u) ∩ Kqi| > θ|Kqi| are propagated to the children. When C becomes empty at a node, the subtree rooted at that node is pruned and not searched further. The main advantage of batching queries in this way is locality of memory references. If b(u) must be loaded from disk, it need be loaded only once per batch C rather than once per query. Batch queries can be parallelized in the same way as nonbatched queries by storing with the nodes on the queue the indices of query sets that remain active at that node. Additionally, batch queries offer an alternative means of parallelization where the query collection C is split evenly among active threads that merge results for the final query results.
Our implementation of SBT allows a user to specify a weight wa between 0 and 1 for each k-mer a in their query Kq. When these weights are specified, a subtree rooted at u is searched if  . That is, a subtree is searched if greater than θ fraction of the possible total k-mer weights are observed. K-mers that the user considers essential to their query (e.g., those spanning an exon junction) can be given higher weight than others. For all experiments here, we use unweighted k-mers (wa = 1 for all a).
. That is, a subtree is searched if greater than θ fraction of the possible total k-mer weights are observed. K-mers that the user considers essential to their query (e.g., those spanning an exon junction) can be given higher weight than others. For all experiments here, we use unweighted k-mers (wa = 1 for all a).
Setting the bloom filter size.
There are two important parameters that need to be set when constructing the bloom filters contained in a SBT. These are the bloom filter length (m) and the number of hash functions (h) used in the filter. We also must choose the k-mer threshold θ for our queries. We explore below the relationship between m, h, θ and the resulting false-positive rate ξ of the filters.
Let S be a collection of r sequencing experiments with the property that each s ∈ S contains n distinct k-mers. We analyze the behavior of a union of filters under the simplifying assumption that the k-mer overlap between all pairs of experiments in S is uniform. Specifically, assume that the probability that two different experiments si and sj in S share any given k-mer is p. In other words, the expected number of k-mers that appear in sj that do not appear in si is d(1 − p), where d is the number of k-mers in the experiments. We can then estimate the expected number of unique k-mers:
Lemma 1.
Let U = Us ∈ S s be the union of sequencing experiments in S as described above. The expected number of distinct k-mers in the union is n (1 − (1 − p)r)/p.
Proof. We have  . Each k-mer in Si is absent from Uj < iSj independently with probability (1 − p)i−1. Therefore
. Each k-mer in Si is absent from Uj < iSj independently with probability (1 − p)i−1. Therefore  , and we have:
, and we have:

The assumptions of a uniform k-mer count n and uniform overlap probability p do not hold in practice. However, under idealized assumptions, Lemma 1 formalizes the intuition that the expected number of elements in the SBT is the union set of all k-mers. In practice, this allows us to define the size of the bloom filter to be equal to an estimate of the total number of unique k-mers. Under the theoretical assumptions, it also shows that when the overlap is large (p is close to 1), the number of elements of U approaches that of a single experiment. Using this relationship, we can select the optimal number of hash functions for such a union as in Theorem 1.
Theorem 1.
The number of hashes that minimizes the false-positive rate of a union filter U with the expected number of elements is

where load = n/m . Under this setting of h , the FPR of U is

which is at most 1/2 so long as h* ≥ 1.
Proof. Follows directly by treating U as a single filter containing n(1 − (1 − p)r )/p items.
In the case of SBTs, we have an advantage that we are not ultimately interested in a single bloom filter query on a k-mer, but rather a set of queries of the k-mers contained in the longer query string q. Thus, we are concerned mostly with the FPR on queries rather than FPR on k-mers. Theorem 2 explores the connection between the two.
Theorem 2
Let
q
be a query string containing
 distinct k-mers. If we treat the k-mers of q as being independent, the probability that
distinct k-mers. If we treat the k-mers of q as being independent, the probability that
 false-positive k-mers appear in a filter U with FPR ξ is
false-positive k-mers appear in a filter U with FPR ξ is

The above expression is nearly 0 when ξ ≪ θ.
Proof. Treating each k-mer in q independently allows us to model the repeated queries using a binomial distribution, yielding (4). A false positive in q occurs when  false-positive k-mers occur in U. Let X be the number of false-positive k-mers, and let Y be the number of correctly determined k-mers. Then
false-positive k-mers occur in U. Let X be the number of false-positive k-mers, and let Y be the number of correctly determined k-mers. Then  . When θ ≥ ξ, we have
. When θ ≥ ξ, we have  and the following bound holds by Chernoff's inequality:
and the following bound holds by Chernoff's inequality:

In our search application, it is natural to require that at least 1/2 the k-mers of a query are present; if <1/2 are present it is fair to say that the query is not contained within the experiment. Therefore θ will typically be >>0.5. In this case, if we choose the FPR of the bloom filters to be 0.5, by Theorem 2, we will be unlikely to observe >θ fraction of false-positive k-mers in the filter. A bloom filter FPR of 0.5 is much higher than typical applications of bloom filters, in which very low false-positive rates are sought. The above analysis assumes independence of the k-mers, which is, of course, unrealistic. Nevertheless, it formalizes the intuition that choosing a high FPR can still lead to few errors. By choosing such a high filter FPR, we can use smaller filters, limiting the memory footprint of the SBT.
To set the bloom filter size, we follow the intuition of Lemma 1, and use an estimate of the total number of unique k-mers across as an estimate of the number of items any individual filter will contain. As it is computationally expensive to quantify this across all 2,652 files, the total was estimated by counting the combined k-mer content of 100 random files using Jellyfish 2.0, yielding an estimate of 1,902,731,933 k-mers. Because we use a filter FPR of 0.5 and h = 1, as suggested by the above theorems, a single element in the SBT has a storage cost of ≤1 bit. Therefore, we set the size m of each bloom filter (in bits) to approximately equal this estimate of the number of k-mers. This offers an approximation that, by undercounting k-mers, sacrifices some accuracy at the highest levels of the tree for a reduced bloom filter size. This value is also substantially higher than the number of k-mers expected in any individual leaf filter and allows leaf filters (where accuracy is most important) to be less saturated and easily compressed. This leads to an uncompressed filter size of 239 MB, and any k-mer of sufficient coverage that is shared between two files will correspond to a shared bit.
Experiments selected for inclusion in the SBT.
A SBT was constructed from 2,652 human, RNA-seq short-read sequencing runs from the NIH SRA. These 2,652 files represented the entire set of publicly available, human RNA-seq runs from blood, brain and breast tissues stored at the SRA at the time of download as determined by keywords in their metadata and excluding files sequenced using the SOLID technology. Files where the metadata was unclear about tissue type or experimental setup were discarded. This tree was used for all experiments described in the manuscript.
Building bloom filters.
The construction of the SBT involves three major tasks: creation of bloom filters for each of the experiments included at its leaves, the construction of the tree and internal bloom filters, and the RRR compression15 of each of the filters. Timing for each stage is given in Supplementary Table 4.
In the experiments here, bloom filters were constructed using the Jellyfish k-mer counting library32 from short-read FASTA files downloaded from the NIH SRA by counting canonical k-mers (the lexicographically smaller k-mer between a k-mer and its reverse complement). We choose k = 20 as these k-mers are reasonably unique within the human genome. Jellyfish was allowed to use 20 threads—all other computation reported here was run with a single thread.
To select only k-mers from sufficiently expressed transcripts and to avoid counting k-mers resulting from sequencing errors, we built trees containing k-mers that occur greater than a file-dependent threshold. This threshold count(si) was determined using the file size of experiment si as follows: count(si) = 1 if si is 300 MB or less, count(si) = 3 for files of size 300–500 MB, count(si) = 10 for files of size 500 MB–1 GB, count(si) = 20 for files between 1 GB and 3 GB, and count(si) = 50 for files > 3 GB or larger FASTA files. These cutoffs were determined by the analysis of a small set of 18 sequence experiments of various sizes and tissue types and were chosen such that at least 60% of the transcripts expressed at a non-zero level in each of these files had an estimated uniform coverage above this number. In practice, we found these thresholds to outperform two naive thresholds (count(si) = 0 and count(si) = 3 for all i) in speed and accuracy. We report only the results from the file-dependent threshold for this reason.
We can use a cutoff based on file size here because all the experiments sequenced the human transcriptome. In a situation where experiments of mixed organism origin are included, a more sophisticated scheme based directly on sequencing coverage would be needed to avoid counting sequencing errors.
After the SBT is built, the filters (both leaf and internal) are compressed using the RRR15 bit vector compression scheme as implemented in the succinct data structures library33. This permits querying a bit without decompression and incurs only a O(log m) factor increase in access time (where m is the size of the bloom filter).
Hardware used for computational experiments.
All times in all experiments reported here, except for SRA-BLAST, were obtained on a shared computer with Intel Xeon 2.60 GHz CPUs using a single thread (or 15 threads in the case of STAR and 20 in the case of Jellyfish). The SBT queries were limited to keeping a single compressed filter in memory at any one time, leading to memory usage of <239 megabytes of RAM. SRA-BLAST queries were executed using its web interface.
Representative query sets and ground truth results.
To determine the accuracy of SBTs, we selected a subset of 100 random read files and used Sailfish20 to quantify the expression of all transcripts in each of these experiments. All SBT queries are queried on the full set of 2,652 files but the accuracy is computed based only on the random subset of files for which we computed expression results from Sailfish.
Note that SBT returns no false negatives in the sense that if a query is covered by k-mers in sufficient depth over θ-fraction of its length then the experiment will be returned. The false negatives in Figure 2 are those experiments where Sailfish indicated the transcript was sufficiently expressed but SBT indicated that it was not sufficiently present. In this context, present and expressed are different concepts: “present” means sufficient coverage of the transcript at a given depth, whereas “expression” is estimated using Sailfish's expectation-maximization approach to allocate mapped reads to isoforms. These are related, but not identical notions. SBT has a 0% false negative rate and a very low false-positive rate identifying present transcripts (false positives identifying present transcripts are due to the one-sided bloom filter errors and k-mer shredding of the reads). Figure 2 shows SBT's false-negative and false-positive rates identifying expressed transcripts, which are partially due to the mismatch between the definitions of present and expressed.
SRA-BLAST.
There are presently no search or alignment tools that can solve the sequence search problem in short-read sequencing files at the scale we attempt here. However, as alignments can be used to determine query coverage and thus the presence of transcripts in sequence files, we compare with SRA-BLAST2. SRA-BLAST has a limitation on the total nucleotide count that can be searched at once and requires specifying SRX (experiment) files rather then SRR (run) files. Because it is impractical to use SRA-BLAST at the SBT scale, we estimated an average SRA-BLAST query time from 100 random queries of a transcript against a single SRX experiment set using the SRA-BLAST webtool2. Specifically, we randomly selected a short read file from the total 2,652 set and a query from the Low representative query set and recorded the time it took SRA-BLAST to return an alignment. Some publicly available SRR files cannot be searched using this webtool and random queries containing these files were discarded. The extrapolated time to process one >1,000-nt query against one megabase of sequence read file was recorded at 0.024 s per megabase per query.
The comparison with SRA-BLAST is not meant to indicate that SBT can provide the same information as SRA-BLAST. In fact, the tools provide complementary information: a list of experiments identified with SBT can be searched for individual read alignments with SRA-BLAST, thereby partially overcoming SRA-BLAST's limitation on the number of experiments that can be searched.
STAR.
We also compare our search times with an alignment-based approach using a read mapping algorithm, STAR20. To do this, we built a separate STAR index for each of the 100 sequence queries in the Low query set using a size-6 pre-index string. Reads from the 100 files analyzed by Sailfish (our ground truth set) were mapped to these indices, allowing zero mismatches during the alignment and a single thread. After 3 d of 15-threaded continuous run-time, only one STAR query had completed searching these 100 files, and from this an average 15-thread STAR query time of 0.708 s per megabase per query was calculated by normalizing the time it takes to perform a STAR alignment against the total size of each sequence file. Single-threaded times were approximated using the CPU clock time, which took 9.9 s per megabase per query. Whereas these single-query indices are more representative of the standard search use case, they represent an index size smaller than is typically used with STAR. To estimate batched times, a single STAR index was built from all 100 sequences queries in the Low query set using a size-11 pre-index string. Six alignments that took longer than 4 h were terminated before completion and their times were discarded as outliers. In this case, an average STAR query time of 0.0110 s per megabase per query was calculated. Note in either case, the time per megabase per query does not include the time to build the STAR index. Although STAR was designed to perform efficient alignments, it represents one of the most competitive existing tools that could be adapted to the general search problem we solve. The comparison with STAR is not intended to indicate that SBT provides the same information as STAR, but rather to show that even a very fast aligner such as STAR cannot identify experiments that contain a query sequence as quickly as SBT.
References
Leinonen, R., Sugawara, H. & Shumway, M. The sequence read archive. Nucleic Acids Res. 39, D19–D21 (2011).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Burrows, M. & Wheeler, D.J. A block sorting lossless data compression algorithm. Technical Report 124 (Digital Equipment Corporation, 1994).
Ferragina, P. & Manzini, G. Indexing compressed text. J. Assoc. Comput. Mach. 52, 552–581 (2005).
Grossi, R. & Vitter, J.S. Compressed suffix arrays and suffix trees with applications to text indexing and string matching. SIAM J. Comput. 35, 378–407 (2005).
Grossi, R., Vitter, J.S. & Xu, B. Wavelet trees: from theory to practice. in Data Compression, Communications and Processing (CCP), 2011 First International Conference on 21–24 June 2011 (pp. 210–221). (IEEE, 2011).
Navarro, G. & Mäkinen, V. Compressed full-text indexes. ACM Comput. Surv. 39, Article No. 2 doi:10.1145/1216370.1216372 (2007).
Ziviani, N., Moura, E., Navarro, G. & Baeza-Yates, R. Compression: a key for next-generation text retrieval systems. IEEE Computer 33, 37–44 (2000).
Navarro, G., Moura, E., Neubert, M., Ziviani, N. & Baeza-Yates, R. Adding compression to block addressing inverted indexes. Inf. Retrieval 3, 49–77 (2000).
Loh, P.-R., Baym, M. & Berger, B. Compressive genomics. Nat. Biotechnol. 30, 627–630 (2012).
Daniels, N.M. et al. Compressive genomics for protein databases. Bioinformatics 29, i283–i290 (2013).
Yu, Y.W., Daniels, N.M., Danko, D.C. & Berger, B. Entropy-scaling search of massive biological data. Cell Syst. 1, 130–140 (2015).
Bloom, B.H. Space/time trade-offs in hash coding with allowable errors. Commun. ACM 13, 422–426 (1970).
Broder, A. & Mitzenmacher, M. Network applications of bloom filters: a survey. Internet Math. 1, 485–509 (2005).
Raman, R., Raman, V. & Srinivasa Rao, S. Succinct indexable dictionaries with applications to encoding k-ary trees and multisets. in Proceedings of the Thirteenth Annual ACM-SIAM Symposium on Discrete Algorithms, SODA '02 (233–242) (Society for Industrial and Applied Mathematics, Philadelphia, PA, USA, 2002).
Crainiceanu, A. Bloofi: a hierarchical bloom filter index with applications to distributed data provenance. in Proceedings of the 2nd International Workshop on Cloud Intelligence, article 4. 10.1145/2501928.2501931 (ACM, 2013).
Pell, J. et al. Scaling metagenome sequence assembly with probabilistic de Bruijn graphs. Proc. Natl. Acad. Sci. USA 109, 13272–13277 (2012).
Chikhi, R. & Rizk, G. Space-efficient and exact de Bruijn graph representation based on a Bloom filter. Algorithms Mol. Biol. 8, 22 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Patro, R., Mount, S.M. & Kingsford, C. Sailfish enables alignment-free isoform quantification from RNA-seq reads using lightweight algorithms. Nat. Biotechnol. 32, 462–464 (2014).
Stranneheim, H. et al. Classification of DNA sequences using Bloom filters. Bioinformatics 26, 1595–1600 (2010).
Melsted, P. & Pritchard, J.K. Efficient counting of k-mers in DNA sequences using a bloom filter. BMC Bioinformatics 12, 333 (2011).
Crainiceanu, A. & Lemire, D. Multidimensional bloom filters. Inf. Syst. 54, 311–324 (2015).
Salikhov, K., Sacomoto, G. & Kucherov, G. Using cascading Bloom filters to improve the memory usage for de Brujin graphs. Algorithms Mol. Biol. 9, 2 (2014).
Rozov, R., Shamir, R. & Halperin, E. Fast lossless compression via cascading Bloom filters. BMC Bioinformatics 15 (suppl. 9), S7 (2014).
Witten, I., Moffat, A. & Bell, T. Managing Gigabytes, 2nd edn. (Morgan Kaufmann, 1999).
Baeza-Yates, R. & Ribeiro, B. Modern Information Retrieval (Addison-Wesley, 1999).
Zhang, Q., Pell, J., Canino-Koning, R., Howe, A.C. & Brown, C.T. These are not the k-mers you are looking for: efficient online k-mer counting using a probabilistic data structure. PLoS One 9, e101271 (2014).
Brown, C.T., Howe, A.C., Zhang, Q., Pyrkosz, A.B. & Brom, T.H. A Reference-Free Algorithm for Computational Normalization of Shotgun Sequencing Data. arXiv:1203.4802 [q-bio.GN]. Preprint at http://arxiv.org/abs/1203.4802.
Rasmussen, K.R., Stoye, J. & Myers, E.W. Efficient q-gram filters for finding all ɛ-matches over a given length. J. Comput. Biol. 13, 296–308 (2006).
Philippe, N., Salson, M., Commes, T. & Rivals, E. CRAC: an integrated approach to the analysis of RNA-seq reads. Genome Biol. 14, R30 (2013).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Gog, S., Beller, T., Moffat, A. & Petri, M. in 13th International Symposium on Experimental Algorithms, Copenhagen, 29 June–1 July 2014 (eds. Gudmundsson, J. & Katajainen, J.) 326–337 (Springer, 2014).
Acknowledgements
This research is funded in part by the Gordon and Betty Moore Foundation's Data-Driven Discovery Initiative through grant GBMF4554 to C.K., by the US National Science Foundation (CCF-1256087, CCF-1319998) and by the US National Institutes of Health (R21HG006913, R01HG007104). C.K. received support as an Alfred P. Sloan Research Fellow. B.S. is a predoctoral trainee supported by US National Institutes of Health training grant T32 EB009403 as part of the Howard Hughes Medical Institute (HHMI)–National Institute of Biomedical Imaging and Bioengineering (NIBIB) Interfaces Initiative.” We would like to thank D. Filippova, J. Fong, H. Wang, E. Sefer, G. Johnson, and especially G. Marçais, G. Duggal and R. Patro for sharing their source code, and for valuable discussions and comments on the manuscript.
Author information
Authors and Affiliations
Contributions
B.S. and C.K. designed the method, devised the experiments, implemented the software and wrote the manuscript. B.S. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
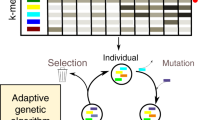
Supplementary Figure 1 Schematic of a Sequence Bloom Tree.
Each node contains a bloom filter that holds the kmers present in the sequencing experiments under it. θ is the fraction of kmers required to be found at each node in order to continue to search its subtree. The SBT returns the experiments that likely contain the query sequence on which further analysis can be performed.
Supplementary Figure 2 Comparison with STAR on batched queries.
STAR was run using an index built from 100 batch-queries and a size 11 pre-index string. Both SBT and STAR were run using one thread and SBT was limited to a single filter in RAM. SBT is an estimated 4056 times faster than STAR under these conditions. STAR times are estimated from extrapolating from querying 100 random SRR files.
Supplementary Figure 3 Average time to query a >1,000-nt sequence across 2,652 SRA files indexed in a Sequence Bloom Tree.
To explore the parameter space of the Sequence Bloom Tree, each of the High, Medium, and Low sets defined in Online Methods were individually queried in the Sequence Bloom Tree for various kmer thresholds θ = 0.7, 0.8, 0.9 and the results averaged. As the threshold increases from 0.7 to 0.9, the time per query decreases while the time per query varies little between changes in TPM.
Supplementary Figure 4 Receiver operating characteristic curve averaged over 100 queries with estimated expression >10 TPM and variable θ.
Solid lines represent mean TP and FP rates, dashed lines represent the median rates on the same experiment. Accuracy is determined by taking Sailfish estimates as ground truth and comparing against SBT estimates on a per-transcript basis.
Supplementary Figure 5 Total number of Sequence Bloom Tree nodes visited as a function of the number of leaf hits when querying 100 random human transcripts in the Low query set.
Number of nodes includes both internal and leaf nodes of the SBT. Each point represents a single query. When a query is found in many of the leaves, the query must also visit a nearly equal number of internal tree nodes, and so the tree structure would not provide any benefit over merely searching all the leaf filters directly. On the other hand, when the query is found in only a few leaves, the total number of nodes visited can be significantly smaller than the number of leaves. For the SBT built here, we find that for queries that are found in 600 or fewer leaves, the tree structure and internal nodes result in an improvement of overall efficiency by visiting fewer than 2652 nodes. A naive approach that did not use the tree would require querying 2652 leaf filters for all queries (denoted by dashed line). Approximately half of the randomly selected queries known to be expressed in the included experiments fall below this threshold.
Supplementary Figure 6 The benefit of SBT as a pre-processing filter.
Estimated time to process the full 2652 dataset using a combination of SBT followed by STAR or SRA-BLAST versus the time to process with STAR or SRA-BLAST without SBT. Dark bars denote the SBT time while the total height represents the total time using conventional search tools on the post-SBT set.
Supplementary Figure 7 Distribution of hit counts (the number of individual SRR files that matched a given query) for all known human transcripts as a function of threshold.
Approximately 83%, 89%, and 95% of all queries (depending on TPM threshold) have fewer than 600 matching files, and the SBT hierarchy would provide significant improvements over a search that used leaf filters only on these queries.
Supplementary Figure 8 Time for querying all known human transcripts.
Total times (single-threaded) for querying all 214,293 human transcripts (in batch mode) against all publicly available blood, breast, and brain RNA-seq experiments in the SRA for θ = 0.7, 0.8, 0.9 as well as the extrapolated time to run Sailfish on the full dataset. Sailfish is significantly faster than nearly all other algorithms for RNA-seq quantification.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 2–5 (PDF 991 kb)
Supplementary Table 1
Supplementary Table 1 (TXT 81 kb)
Supplementary Software zip file
Supplementary Software (ZIP 170 kb)
Rights and permissions
About this article
Cite this article
Solomon, B., Kingsford, C. Fast search of thousands of short-read sequencing experiments. Nat Biotechnol 34, 300–302 (2016). https://doi.org/10.1038/nbt.3442
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3442
This article is cited by
-
Hierarchical Interleaved Bloom Filter: enabling ultrafast, approximate sequence queries
Genome Biology (2023)
-
Sketching and sampling approaches for fast and accurate long read classification
BMC Bioinformatics (2022)
-
Strain level microbial detection and quantification with applications to single cell metagenomics
Nature Communications (2022)
-
Disk compression of k-mer sets
Algorithms for Molecular Biology (2021)
-
Bifrost: highly parallel construction and indexing of colored and compacted de Bruijn graphs
Genome Biology (2020)