Abstract
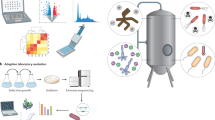
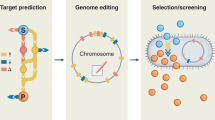
Industrial strain development requires system-wide engineering and optimization of cellular metabolism while considering industrially relevant fermentation and recovery processes. It can be conceptualized as several strategies, which may be implemented in an iterative fashion and in different orders. The key challenges have been the time-, cost- and labor-intensive processes of strain development owing to the difficulties in understanding complex interactions among the metabolic, gene regulatory and signaling networks at the cell level, which are collectively represented as overall system performance under industrial fermentation conditions. These challenges can be overcome by taking systems approaches through the use of state-of-the-art tools of systems biology, synthetic biology and evolutionary engineering in the context of industrial bioprocess. Major systems metabolic engineering achievements in recent years include microbial production of amino acids (L-valine, L-threonine, L-lysine and L-arginine), bulk chemicals (1,4-butanediol, 1,4-diaminobutane, 1,5-diaminopentane, 1,3-propanediol, butanol, isobutanol and succinic acid) and drugs (artemisinin).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hong, K.K. & Nielsen, J. Metabolic engineering of Saccharomyces cerevisiae: a key cell factory platform for future biorefineries. Cell. Mol. Life Sci. 69, 2671–2690 (2012).
Pronk, J.T. et al. How to set up collaborations between academia and industrial biotech companies. Nat. Biotechnol. 33, 237–240 (2015).
Bailey, J.E. Toward a science of metabolic engineering. Science 252, 1668–1675 (1991).
Lee, J.W. et al. Systems metabolic engineering of microorganisms for natural and non-natural chemicals. Nat. Chem. Biol. 8, 536–546 (2012).
Sagt, C.M. Systems metabolic engineering in an industrial setting. Appl. Microbiol. Biotechnol. 97, 2319–2326 (2013).
Park, J.H., Lee, K.H., Kim, T.Y. & Lee, S.Y. Metabolic engineering of Escherichia coli for the production of L-valine based on transcriptome analysis and in silico gene knockout simulation. Proc. Natl. Acad. Sci. USA 104, 7797–7802 (2007).
Park, J.H., Kim, T.Y., Lee, K.H. & Lee, S.Y. Fed-batch culture of Escherichia coli for L-valine production based on in silico flux response analysis. Biotechnol. Bioeng. 108, 934–946 (2011).
Park, J.H., Jang, Y.S., Lee, J.W. & Lee, S.Y. Escherichia coli W as a new platform strain for the enhanced production of L-valine by systems metabolic engineering. Biotechnol. Bioeng. 108, 1140–1147 (2011).
Lee, K.H., Park, J.H., Kim, T.Y., Kim, H.U. & Lee, S.Y. Systems metabolic engineering of Escherichia coli for L-threonine production. Mol. Syst. Biol. 3, 149 (2007).
Lee, J.W. et al. Development of sucrose-utilizing Escherichia coli K-12 strain by cloning β-fructofuranosidases and its application for L-threonine production. Appl. Microbiol. Biotechnol. 88, 905–913 (2010).
Yim, H. et al. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat. Chem. Biol. 7, 445–452 (2011).
Paddon, C.J. et al. High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532 (2013).
Lee, S.J., Song, H. & Lee, S.Y. Genome-based metabolic engineering of Mannheimia succiniciproducens for succinic acid production. Appl. Environ. Microbiol. 72, 1939–1948 (2006).
Becker, J. et al. Systems-wide analysis and engineering of metabolic pathway fluxes in bio-succinate producing Basfia succiniciproducens. Biotechnol. Bioeng. 110, 3013–3023 (2013).
Park, S.H. et al. Metabolic engineering of Corynebacterium glutamicum for L-arginine production. Nat. Commun. 5, 4618 (2014).
Becker, J., Zelder, O., Häfner, S., Schröder, H. & Wittmann, C. From zero to hero—design-based systems metabolic engineering of Corynebacterium glutamicum for L-lysine production. Metab. Eng. 13, 159–168 (2011).
Kind, S. et al. From zero to hero—production of bio-based nylon from renewable resources using engineered Corynebacterium glutamicum. Metab. Eng. 25, 113–123 (2014).
Lee, S.Y., Lee, D.Y. & Kim, T.Y. Systems biotechnology for strain improvement. Trends Biotechnol. 23, 349–358 (2005).
Medema, M.H., van Raaphorst, R., Takano, E. & Breitling, R. Computational tools for the synthetic design of biochemical pathways. Nat. Rev. Microbiol. 10, 191–202 (2012).
Paddon, C.J. & Keasling, J.D. Semi-synthetic artemisinin: a model for the use of synthetic biology in pharmaceutical development. Nat. Rev. Microbiol. 12, 355–367 (2014).
Esvelt, K.M. & Wang, H.H. Genome-scale engineering for systems and synthetic biology. Mol. Syst. Biol. 9, 641 (2013).
Yadav, V.G., De Mey, M., Lim, C.G., Ajikumar, P.K. & Stephanopoulos, G. The future of metabolic engineering and synthetic biology: towards a systematic practice. Metab. Eng. 14, 233–241 (2012).
Cobb, R.E., Si, T. & Zhao, H. Directed evolution: an evolving and enabling synthetic biology tool. Curr. Opin. Chem. Biol. 16, 285–291 (2012).
Lee, S.Y. High cell-density culture of Escherichia coli. Trends Biotechnol. 14, 98–105 (1996).
Lee, B.R., Cho, S., Song, Y., Kim, S.C. & Cho, B.K. Emerging tools for synthetic genome design. Mol. Cells 35, 359–370 (2013).
Song, C.W., Lee, J. & Lee, S.Y. Genome engineering and gene expression control for bacterial strain development. Biotechnol. J. 10, 56–68 (2015).
Zhuang, K.H. & Herrgård, M.J. Multi-scale exploration of the technical, economic, and environmental dimensions of bio-based chemical production. Metab. Eng. 31, 1–12 (2015).
Croughan, M.S., Konstantinov, K.B. & Cooney, C. The future of industrial bioprocessing: batch or continuous? Biotechnol. Bioeng. 112, 648–651 (2015).
Van Dien, S. From the first drop to the first truckload: commercialization of microbial processes for renewable chemicals. Curr. Opin. Biotechnol. 24, 1061–1068 (2013).
Choi, S. et al. Production of 4-hydroxybutyric acid by metabolically engineered Mannheimia succiniciproducens and its conversion to γ-butyrolactone by acid treatment. Metab. Eng. 20, 73–83 (2013).
Anbarasan, P. et al. Integration of chemical catalysis with extractive fermentation to produce fuels. Nature 491, 235–239 (2012).
Guan, Y., Dunham, M., Caudy, A. & Troyanskaya, O. Systematic planning of genome-scale experiments in poorly studied species. PLOS Comput. Biol. 6, e1000698 (2010).
Cobb, R.E., Wang, Y. & Zhao, H. High-efficiency multiplex genome editing of Streptomyces species using an engineered CRISPR/Cas system. ACS Synth. Biol. 4, 723–728 (2015).
Tong, Y., Charusanti, P., Zhang, L., Weber, T. & Lee, S.Y. CRISPR-Cas9 based engineering of actinomycetal genomes. ACS Synth. Biol. 10.1021/acssynbio.5b00038 (25 March 2015).
Shin, J.H., Kim, H.U., Kim, D.I. & Lee, S.Y. Production of bulk chemicals via novel metabolic pathways in microorganisms. Biotechnol. Adv. 31, 925–935 (2013).
Lee, J.W., Kim, T.Y., Jang, Y.S., Choi, S. & Lee, S.Y. Systems metabolic engineering for chemicals and materials. Trends Biotechnol. 29, 370–378 (2011).
Lee, J.W., Kim, H.U., Choi, S., Yi, J. & Lee, S.Y. Microbial production of building block chemicals and polymers. Curr. Opin. Biotechnol. 22, 758–767 (2011).
Choi, Y.J. & Lee, S.Y. Microbial production of short-chain alkanes. Nature 502, 571–574 (2013).
Dellomonaco, C., Clomburg, J.M., Miller, E.N. & Gonzalez, R. Engineered reversal of the β-oxidation cycle for the synthesis of fuels and chemicals. Nature 476, 355–359 (2011).
Yu, J.L., Xia, X.X., Zhong, J.J. & Qian, Z.G. Direct biosynthesis of adipic acid from a synthetic pathway in recombinant Escherichia coli. Biotechnol. Bioeng. 111, 2580–2586 (2014).
Borodina, I. et al. Establishing a synthetic pathway for high-level production of 3-hydroxypropionic acid in Saccharomyces cerevisiae via β-alanine. Metab. Eng. 27, 57–64 (2015).
Jiang, M. & Pfeifer, B.A. Metabolic and pathway engineering to influence native and altered erythromycin production through E. coli. Metab. Eng. 19, 42–49 (2013).
Zhang, K., Li, H., Cho, K.M. & Liao, J.C. Expanding metabolism for total biosynthesis of the nonnatural amino acid L-homoalanine. Proc. Natl. Acad. Sci. USA 107, 6234–6239 (2010).
Smanski, M.J. et al. Functional optimization of gene clusters by combinatorial design and assembly. Nat. Biotechnol. 32, 1241–1249 (2014).
Ling, H., Teo, W., Chen, B., Leong, S.S. & Chang, M.W. Microbial tolerance engineering toward biochemical production: from lignocellulose to products. Curr. Opin. Biotechnol. 29, 99–106 (2014).
Utrilla, J. et al. Engineering and adaptive evolution of Escherichia coli for D-lactate fermentation reveals GatC as a xylose transporter. Metab. Eng. 14, 469–476 (2012).
Dunlop, M.J. et al. Engineering microbial biofuel tolerance and export using efflux pumps. Mol. Syst. Biol. 7, 487 (2011).
Lam, F.H., Ghaderi, A., Fink, G.R. & Stephanopoulos, G. Biofuels. Engineering alcohol tolerance in yeast. Science 346, 71–75 (2014).
Na, D. et al. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat. Biotechnol. 31, 170–174 (2013).
Yoo, S.M., Na, D. & Lee, S.Y. Design and use of synthetic regulatory small RNAs to control gene expression in Escherichia coli. Nat. Protoc. 8, 1694–1707 (2013).
King, Z.A. & Feist, A.M. Optimal cofactor swapping can increase the theoretical yield for chemical production in Escherichia coli and Saccharomyces cerevisiae. Metab. Eng. 24, 117–128 (2014).
Rellos, P., Ma, J. & Scopes, R.K. Alteration of substrate specificity of Zymomonas mobilis alcohol dehydrogenase-2 using in vitro random mutagenesis. Protein Expr. Purif. 9, 83–90 (1997).
Kim, J. & Copley, S.D. Inhibitory cross-talk upon introduction of a new metabolic pathway into an existing metabolic network. Proc. Natl. Acad. Sci. USA 109, E2856–E2864 (2012).
Jantama, K. et al. Eliminating side products and increasing succinate yields in engineered strains of Escherichia coli C. Biotechnol. Bioeng. 101, 881–893 (2008).
Kim, T.Y., Park, J.M., Kim, H.U., Cho, K.M. & Lee, S.Y. Design of homo-organic acid producing strains using multi-objective optimization. Metab. Eng. 28, 63–73 (2015).
Lewis, N.E., Nagarajan, H. & Palsson, B.O. Constraining the metabolic genotype-phenotype relationship using a phylogeny of in silico methods. Nat. Rev. Microbiol. 10, 291–305 (2012).
Wang, H.H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Wang, H.H. et al. Genome-scale promoter engineering by coselection MAGE. Nat. Methods 9, 591–593 (2012).
Warner, J.R., Reeder, P.J., Karimpour-Fard, A., Woodruff, L.B. & Gill, R.T. Rapid profiling of a microbial genome using mixtures of barcoded oligonucleotides. Nat. Biotechnol. 28, 856–862 (2010).
Hollinshead, W., He, L. & Tang, Y.J. Biofuel production: an odyssey from metabolic engineering to fermentation scale-up. Front. Microbiol. 5, 344 (2014).
Dahl, R.H. et al. Engineering dynamic pathway regulation using stress-response promoters. Nat. Biotechnol. 31, 1039–1046 (2013).
Galanie, S., Thodey, K., Trenchard, I.J., Filsinger, I.M. & Smolke, C.D. Complete biosynthesis of opioids in yeast. Science 349, 1095–1100 (2015).
Karr, J.R. et al. A whole-cell computational model predicts phenotype from genotype. Cell 150, 389–401 (2012).
Hsiao, T.Y., Glatz, C.E. & Glatz, B.A. Broth recycle in a yeast fermentation. Biotechnol. Bioeng. 44, 1228–1234 (1994).
Yin, J., Fu, X.Z., Wu, Q., Chen, J.C. & Chen, G.Q. Development of an enhanced chromosomal expression system based on porin synthesis operon for halophile Halomonas sp. Appl. Microbiol. Biotechnol. 98, 8987–8997 (2014).
Caspeta, L. et al. Biofuels. Altered sterol composition renders yeast thermotolerant. Science 346, 75–78 (2014).
Bonneau, R. et al. A predictive model for transcriptional control of physiology in a free living cell. Cell 131, 1354–1365 (2007).
Xia, J., Yamaji, N. & Ma, J.F. An appropriate concentration of arginine is required for normal root growth in rice. Plant Signal. Behav. 9, e28717 (2014).
Burgard, A.P., Pharkya, P. & Maranas, C.D. Optknock: a bilevel programming framework for identifying gene knockout strategies for microbial strain optimization. Biotechnol. Bioeng. 84, 647–657 (2003).
Antoniewicz, M.R. 13C metabolic flux analysis: optimal design of isotopic labeling experiments. Curr. Opin. Biotechnol. 24, 1116–1121 (2013).
Trinh, C.T., Wlaschin, A. & Srienc, F. Elementary mode analysis: a useful metabolic pathway analysis tool for characterizing cellular metabolism. Appl. Microbiol. Biotechnol. 81, 813–826 (2009).
Kim, H.U., Kim, T.Y. & Lee, S.Y. Metabolic flux analysis and metabolic engineering of microorganisms. Mol. Biosyst. 4, 113–120 (2008).
Acknowledgements
We thank Seok Hyun Park, Sol Choi, Joungmin Lee, Chan Woo Song, Jae Ho Shin and Won Jun Kim for thoughtful discussions. This work was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries (NRF-2012M1A2A2026556) and by Intelligent Synthetic Biology Center through the Global Frontier Project (2011-0031963) from the Ministry of Science, ICT and Future Planning (MSIP) through the National Research Foundation (NRF) of Korea. This work was also supported by the Novo Nordisk Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–3 (PDF 1931 kb)
Rights and permissions
About this article
Cite this article
Lee, S., Kim, H. Systems strategies for developing industrial microbial strains. Nat Biotechnol 33, 1061–1072 (2015). https://doi.org/10.1038/nbt.3365
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3365
This article is cited by
-
Efficient Synthesis of Chiral Aryl Alcohol with a Novel Kosakonia radicincitans Isolate in Tween 20/L-carnitine: Lysine-Containing Synergistic Reaction System
Applied Biochemistry and Biotechnology (2024)
-
Systemic metabolic engineering of Enterobacter aerogenes for efficient 2,3-butanediol production
Applied Microbiology and Biotechnology (2024)
-
Enhanced production of d-pantothenic acid in Corynebacterium glutamicum using an efficient CRISPR–Cpf1 genome editing method
Microbial Cell Factories (2023)
-
Multivariate modular metabolic engineering for enhanced l-methionine biosynthesis in Escherichia coli
Biotechnology for Biofuels and Bioproducts (2023)
-
Construction of a plasmid-free l-leucine overproducing Escherichia coli strain through reprogramming of the metabolic flux
Biotechnology for Biofuels and Bioproducts (2023)