Abstract
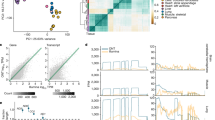
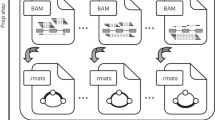
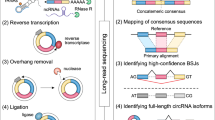
Alternative splicing shapes mammalian transcriptomes, with many RNA molecules undergoing multiple distant alternative splicing events. Comprehensive transcriptome analysis, including analysis of exon co-association in the same molecule, requires deep, long-read sequencing. Here we introduce an RNA sequencing method, synthetic long-read RNA sequencing (SLR-RNA-seq), in which small pools (≤1,000 molecules/pool, ≤1 molecule/gene for most genes) of full-length cDNAs are amplified, fragmented and short-read-sequenced. We demonstrate that these RNA sequences reconstructed from the short reads from each of the pools are mostly close to full length and contain few insertion and deletion errors. We report many previously undescribed isoforms (human brain: ∼13,800 affected genes, 14.5% of molecules; mouse brain ∼8,600 genes, 18% of molecules) and up to 165 human distant molecularly associated exon pairs (dMAPs) and distant molecularly and mutually exclusive pairs (dMEPs). Of 16 associated pairs detected in the mouse brain, 9 are conserved in human. Our results indicate conserved mechanisms that can produce distant but phased features on transcript and proteome isoforms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kornblihtt, A.R. et al. Alternative splicing: a pivotal step between eukaryotic transcription and translation. Nat. Rev. Mol. Cell Biol. 14, 153–165 (2013).
Nilsen, T.W. & Graveley, B.R. Expansion of the eukaryotic proteome by alternative splicing. Nature 463, 457–463 (2010).
Chen, J. & Weiss, W.A. Alternative splicing in cancer: implications for biology and therapy. Oncogene 34, 1–14 (2014).
Bonnal, S., Vigevani, L. & Valcárcel, J. The spliceosome as a target of novel antitumour drugs. Nat. Rev. Drug Discov. 11, 847–859 (2012).
Ben-Dov, C., Hartmann, B., Lundgren, J. & Valcárcel, J. Genome-wide analysis of alternative pre-mRNA splicing. J. Biol. Chem. 283, 1229–1233 (2008).
Fagnani, M. et al. Functional coordination of alternative splicing in the mammalian central nervous system. Genome Biol. 8, R108 (2007).
Johnson, J.M. et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science 302, 2141–2144 (2003).
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320, 1344–1349 (2008).
Wang, E.T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Mortazavi, A., Williams, B.A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat. Methods 5, 621–628 (2008).
Sultan, M. et al. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science 321, 956–960 (2008).
Wilhelm, B.T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453, 1239–1243 (2008).
Modrek, B., Resch, a, Grasso, C. & Lee, C. Genome-wide detection of alternative splicing in expressed sequences of human genes. Nucleic Acids Res. 29, 2850–2859 (2001).
Harrow, J. et al. GENCODE: producing a reference annotation for ENCODE. Genome Biol. 7 (suppl. 1), S4 (2006).
Pan, Q., Shai, O., Lee, L.J., Frey, B.J. & Blencowe, B.J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Bernstein, B.E. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Tilgner, H., Grubert, F., Sharon, D. & Snyder, M.P. Defining a personal, allele-specific, and single-molecule long-read transcriptome. Proc. Natl. Acad. Sci. USA 111, 9869–9874 (2014).
Steijger, T. et al. Assessment of transcript reconstruction methods for RNA-seq. Nat. Methods 10, 1177–1184 (2013).
Cho, H. et al. High-resolution transcriptome analysis with long-read RNA sequencing. PLoS ONE 9, e108095 (2014).
Tilgner, H. et al. Accurate identification and analysis of human mRNA isoforms using deep long read sequencing. G3 3, 387–397 (2013).
Sharon, D., Tilgner, H., Grubert, F. & Snyder, M. A single-molecule long-read survey of the human transcriptome. Nat. Biotechnol. 31, 1009–1014 (2013).
Au, K.F. et al. Characterization of the human ESC transcriptome by hybrid sequencing. Proc. Natl. Acad. Sci. USA 110, E4821–E4830 (2013).
Koren, S. et al. Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat. Biotechnol. 30, 693–700 (2012).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Kuleshov, V. et al. Whole-genome haplotyping using long reads and statistical methods. Nat. Biotechnol. 32, 261–266 (2014).
McCoy, R.C. et al. Illumina TruSeq synthetic long-reads empower de novo assembly and resolve complex, highly-repetitive transposable elements. PLoS ONE 9, e106689 (2014).
The External RNA Controls Consortium. The External RNA Controls Consortium: a progress report. Nat. Methods 2, 731–734 (2005).
Chinwalla, A.T. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Wu, T.D. & Watanabe, C.K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Li, S. et al. Multi-platform assessment of transcriptome profiling using RNA-seq in the ABRF next-generation sequencing study. Nat. Biotechnol. 32, 915–925 (2014).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Duret, L., Chureau, C., Samain, S., Weissenbach, J. & Avner, P. The Xist RNA gene evolved in eutherians by pseudogenization of a protein-coding gene. Science 312, 1653–1655 (2006).
Braunschweig, U. et al. Widespread intron retention in mammals functionally tunes transcriptomes. Genome Res. 24, 1774–1786 (2014).
Benjamini, Y. & Yekutieli, D. The control of the false discovery rate in multiple testing under dependency. Ann. Stat. 29, 1165–1188 (2001).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Karolchik, D. et al. The UCSC Genome Browser database: 2014 update. Nucleic Acids Res. 42, D764–D770 (2014).
Cheng, J. et al. Protection from Fas-mediated apoptosis by a soluble form of the Fas molecule. Science 263, 1759–1762 (1994).
Lareau, L.F., Inada, M., Green, R.E., Wengrod, J.C. & Brenner, S.E. Unproductive splicing of SR genes associated with highly conserved and ultraconserved DNA elements. Nature 446, 926–929 (2007).
Sun, S., Zhang, Z., Sinha, R., Karni, R. & Krainer, A.R. SF2/ASF autoregulation involves multiple layers of post-transcriptional and translational control. Nat. Struct. Mol. Biol. 17, 306–312 (2010).
Smith, C.W. & Valcárcel, J. Alternative pre-mRNA splicing: the logic of combinatorial control. Trends Biochem. Sci. 25, 381–388 (2000).
Barash, Y. et al. Deciphering the splicing code. Nature 465, 53–59 (2010).
Tilgner, H. et al. Deep sequencing of subcellular RNA fractions shows splicing to be predominantly co-transcriptional in the human genome but inefficient for lncRNAs. Genome Res. 22, 1616–1625 (2012).
Dujardin, G. et al. Transcriptional elongation and alternative splicing. Biochim. Biophys. Acta 1829, 134–140 (2013).
Carrillo Oesterreich, F., Preibisch, S. & Neugebauer, K.M. Global analysis of nascent RNA reveals transcriptional pausing in terminal exons. Mol. Cell 40, 571–581 (2010).
Vargas, D.Y. et al. Single-molecule imaging of transcriptionally coupled and uncoupled splicing. Cell 147, 1054–1065 (2011).
Acknowledgements
We thank N. Spies and F.A. Bava for a thorough reading of this manuscript and valuable comments and S. Shringarpure, V. Kuleshov, C.S. Foo and H. Tang for valuable comments on statistics. We thank A. Brunet for providing mice and S. Munro for valuable comments on this manuscript. We also thank the Genetics Bioinformatics Service Center at Stanford for providing a well-working computing cluster. M.R. is paid by grant 12-131829 from the Danish Council for Independent Research. This work was supported by grant 5U01HL10739304 (to M.S. as co-PI), 1P50HG007735-01 (to M.S. as co-PI) and 5P01GM09913004 (to M.S.).
Author information
Authors and Affiliations
Contributions
H.T., T.B., F.C. and M.P.S. devised the project. F.J., T.B., E.J., A.M. and M.R. carried out experiments. I.H. euthanized mice and extracted brains. H.T. carried out computational analysis. C.D.B. and M.P.S. supervised the project and provided financial support. H.T. wrote the first version of the manuscript. H.T., F.J., M.R. and M.P.S. wrote the final version of the manuscript with contributions from the other authors.
Corresponding author
Ethics declarations
Competing interests
A. Moshrefi, E. Jaeger and F. Chen are employees of Illumina. T. Blauwkamp is a former employee of Illumina. M. Snyder is on the scientific advisory board of Personalis, GenapSys and AxioMx. C. Bustamante is a founder of Identify Genomics. He is also on the Scientific Advisory Board of Identify, Etalon, Personalis and Ancestry.com. He is a former member of the advisory board member of InVitae. None of these organizations played a role in the design or conduct of the work presented here.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1 and 2 and Supplementary Results (PDF 5403 kb)
Supplementary Data Set 1
This is a README describing all the supplementary datasets. (ZIP 324 kb)
Supplementary Data Set 2
Human Molecules per Million measurements for spliced genes. See associated README for file format. (ZIP 231 kb)
Supplementary Data Set 3
Mouse Molecules per Million measurements for spliced genes for both mice combined. See associated README for file format. (ZIP 228 kb)
Supplementary Data Set 4
Mouse Molecules per Million measurements for spliced genes for mouse number 2. See associated README for file format. (ZIP 220 kb)
Supplementary Data Set 5
Human Percent-Spliced-In (Psi) measurements for splice-sites. See associated README for file format. (ZIP 4739 kb)
Supplementary Data Set 6
Mouse Percent-Spliced-In (Psi) measurements for splice-sites for both mice combined. See associated README for file format. (ZIP 2622 kb)
Supplementary Data Set 7
Mouse Percent-Spliced-In (Psi) measurements for splice-sites for mouse number 1. See associated README for file format. (ZIP 2236 kb)
Supplementary Data Set 8
Mouse Percent-Spliced-In (Psi) measurements for splice-sites for mouse number 2. See associated README for file format. (ZIP 1686 kb)
Supplementary Data Set 9
Human Percent-Isoforme (Pi) measurements for spliced genes. See associated README for file format. (ZIP 3023 kb)
Supplementary Data Set 10
Mouse Percent-Isoforme (Pi) measurements for spliced genes for both mice combined. See associated README for file format. (ZIP 1100 kb)
Supplementary Data Set 11
Mouse Percent-Isoforme (Pi) measurements for spliced genes for mouse number 1. See associated README for file format. (ZIP 932 kb)
Supplementary Data Set 12
Mouse Percent-Isoforme (Pi) measurements for spliced genes for mouse number 2. See associated README for file format. (ZIP 695 kb)
Supplementary Data Set 13
Human "distant Molecularly Associated Pairs" (dMAPs) of exons and "distant Molecularly and Mutually Exclusive Pairs" (dMEPs) of exons using only human brain RNA. See associated README for file format. (ZIP 6 kb)
Supplementary Data Set 14
Human "distant Molecularly Associated Pairs" (dMAPs) of exons and "distant Molecularly and Mutually Exclusive Pairs" (dMEPs) of exons using human brain RNA and a variety of previously published long read RNA-datasets (Tilgner et al, GGG, 2013; Sharon et al, Nature Biotechnology, 2013; Tilgner et al, PNAS, 2014). See associated README for file format. (ZIP 11 kb)
Supplementary Data Set 15
Mouse "distant Molecularly Associated Pairs" (dMAPs) of exons and "distant Molecularly and Mutually Exclusive Pairs" (dMEPs) of exons using only mouse brain RNA. See associated README for file format. (ZIP 1 kb)
Rights and permissions
About this article
Cite this article
Tilgner, H., Jahanbani, F., Blauwkamp, T. et al. Comprehensive transcriptome analysis using synthetic long-read sequencing reveals molecular co-association of distant splicing events. Nat Biotechnol 33, 736–742 (2015). https://doi.org/10.1038/nbt.3242
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3242
This article is cited by
-
Splicing complexity as a pivotal feature of alternative exons in mammalian species
BMC Genomics (2023)
-
Long-read sequencing reveals the landscape of aberrant alternative splicing and novel therapeutic target in colorectal cancer
Genome Medicine (2023)
-
Full-length transcriptome from different life stages of cobia (Rachycentron canadum, Rachycentridae)
Scientific Data (2023)
-
Single-molecule long-read sequencing reveals the potential impact of posttranscriptional regulation on gene dosage effects on the avian Z chromosome
BMC Genomics (2022)
-
A comparison of mRNA sequencing (RNA-Seq) library preparation methods for transcriptome analysis
BMC Genomics (2022)