Abstract
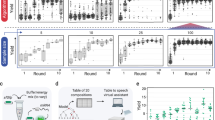
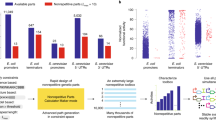
Large microbial gene clusters encode useful functions, including energy utilization and natural product biosynthesis, but genetic manipulation of such systems is slow, difficult and complicated by complex regulation. We exploit the modularity of a refactored Klebsiella oxytoca nitrogen fixation (nif) gene cluster (16 genes, 103 parts) to build genetic permutations that could not be achieved by starting from the wild-type cluster. Constraint-based combinatorial design and DNA assembly are used to build libraries of radically different cluster architectures by varying part choice, gene order, gene orientation and operon occupancy. We construct 84 variants of the nifUSVWZM operon, 145 variants of the nifHDKY operon, 155 variants of the nifHDKYENJ operon and 122 variants of the complete 16-gene pathway. The performance and behavior of these variants are characterized by nitrogenase assay and strand-specific RNA sequencing (RNA-seq), and the results are incorporated into subsequent design cycles. We have produced a fully synthetic cluster that recovers 57% of wild-type activity. Our approach allows the performance of genetic parts to be quantified simultaneously in hundreds of genetic contexts. This parallelized design-build-test-learn cycle, which can access previously unattainable regions of genetic space, should provide a useful, fast tool for genetic optimization and hypothesis testing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Sequence Read Archive
References
Czar, M.J., Anderson, J.C., Bader, J.S. & Peccoud, J. Gene synthesis demystified. Trends Biotechnol. 27, 63–72 (2009).
Gibson, D.G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010).
Kröger, J.D. et al. The transcriptional landscape and small RNAs of Salmonella enterica serovar Typhimurium. Proc. Natl. Acad. Sci. USA 109, E1277–E1286 (2012).
Arnold, W., Rump, A., Klipp, W., Priefer, U.B. & Pühler, A. Nucleotide sequence of a 24,206-base-pair DNA fragment carrying the entire nitrogen fixation gene cluster of Klebsiella pneumoniae. J. Mol. Biol. 203, 715–738 (1988).
Chan, L.Y., Kosuri, S. & Endy, D. Refactoring bacteriophage T7. Mol. Sys. Biol. 1, 2005.0018 (2005).
Temme, K., Zhao, D. & Voigt, C.A. Refactoring the nitrogen fixation gene cluster from Klebsiella oxytoca. Proc. Natl. Acad. Sci. USA 109, 7085–7090 (2012).
Temme, K., Hill, R., Segall-Shapiro, T.H., Moser, F. & Voigt, C.A. Modular control of multiple pathways using engineered orthogonal T7 polymerases. Nucleic Acids Res. 40, 8773–8781 (2012).
Beatty, P.H. & Good, A.G. Future prospects for cereals that fix nitrogen. Science 333, 416–417 (2011).
Arnold, W., Rump, A., Klipp, W., Priefer, U.B. & Pühler, A. Nucleotide sequence of a 24,206-base-pair DNA fragment carrying the entire nitrogen fixation gene cluster of Klebsiella pneumoniae. J. Mol. Biol. 203, 715–738 (1988).
Eady, R.R., Issack, R., Kennedy, C., Postgate, J.R. & Ratcliffe, H.D. Nitrogenase synthesis in Klebsiella pneumonia: comparison of ammonium and oxygen regulation. J. Gen. Microbiol. 104, 277–285 (1978).
Lowe, D.J. & Thorneley, R.N.F. The mechanism of Klebsiella pneumoniae nitrogenase action: pre-steady-state kinetics of H2 formation. Biochem. J. 224, 877–886 (1984).
Dixon, R., Cheng, Q., Shen, G.F., Day, A. & Dowson-Day, M. Nif gene transfer and expression in chloroplasts: prospects and problems. Plant Soil 194, 193–203 (1997).
Dukes, P., Lamken, E. & Wilson, R. Workshop report: Combinatorial design theory (Banff International Research Station meeting 08w5098) https://www.birs.ca/workshops/2008/08w5098/report08w5098.pdf (2008).
Alper, H., Fischer, C., Nevoigt, E. & Stephanopoulos, G. Tuning genetic control through promoter engineering. Proc. Natl. Acad. Sci. USA 102, 12678–12683 (2005).
Beynon, J., Cannon, M., Buchanan-Wollaston, V. & Cannon, F. The nif promoters of Klebsiella pneumonia have a characteristic primary structure. Cell 34, 665–671 (1983).
Bilitchenko, L. et al. Eugene—a domain specific language for specifying and constraining synthetic biological parts, devices, and systems. PLoS ONE 6, e18882 (2011).
Salis, H.M., Mirsky, E.A. & Voigt, C.A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Davis, J.H., Rubin, A.J. & Sauer, R.T. Design, construction and characterization of a set of insulated bacterial promoters. Nucleic Acids Res. 39, 1131–1141 (2011).
Crook, N.C., Freeman, E.S. & Alper, H.S. Re-engineering multicloning sites for function and convenience. Nucleic Acids Res. 39, e92 (2011).
Weber, E., Engler, C., Gruetzner, R., Werner, S. & Marillonnet, S. A modular cloning system for standardized assembly of multigene constructs. PLoS ONE 6, e16765 (2011).
Wang, X. et al. Using synthetic biology to distinguish and overcome regulatory and functional barriers related to nitrogen fixation. PLoS ONE 8, e68677 (2013).
Cannon, F.C., Dixon, R.A. & Postgate, J.R. Derivation and properties of F-prime factors in Escherichia coli carrying nitrogen fixation genes from Klebsiella pneumoniae. J. Gen. Microbiol. 93, 111–125 (1976).
Price, M.N., Huang, K.H., Arkin, A.P. & Alm, E.J. Operon formation is driven by co-regulation and not by horizontal gene transfer. Genome Res. 15, 809–819 (2005).
Lim, H.N., Lee, Y. & Hussein, R. Fundamental relationship between operon organization and gene expression. Proc. Natl. Acad. Sci. USA 108, 10626–10631 (2011).
Liang, L.W., Hussein, R., Block, D.H.S. & Lim, H.N. Minimal effect of gene clustering on expression in Escherichia coli. Genetics 193, 453–465 (2013).
Endy, D., You, L., Yin, J. & Molineux, I. Computation, prediction, and experimental test of fitness for bacteriophage T7 mutants with permuted genomes. Proc. Natl. Acad. Sci. USA 97, 5375–5380 (2000).
von Dassow, G., Meir, E., Munro, E.M. & Odell, G.M. The segment polarity network is a robust developmental module. Nature 406, 188–192 (2000).
Hamilton, T.L. et al. Transcriptional profiling of nitrogen fixation in Azotobacter vinelandii. J. Bacteriol. 193, 4477–4486 (2011).
Yan, Y. et al. Global transcriptional analysis of nitrogen fixation and ammonium repression in root-associated Pseudomonas stutzeri A1501. BMC Genomics 11, 11 (2010).
Poza-Carrión, C., Jiménez-Vicente, E., Navarro-Rodríguez, M., Echavarri-Erasun, C. & Rubio, L.M. Kinetics of nif gene expression in a nitrogen-fixing bacterium. J. Bacteriol. 196, 595–603 (2014).
Jeng, S.T., Gardner, J.F. & Gumport, R.I. Transcription termination by bacteriophage T7 RNA polymerase at rho-independent terminators. J. Biol. Chem. 265, 3823–3830 (1990).
McAllister, W.T. & Morris, C. Utilization of bacteriophage T7 late promoters in recombinant plasmids during infection. J. Mol. Biol. 153, 527–544 (1981).
Cardinale, S. & Arkin, A.P. Contextualizing context for synthetic biology–identifying causes of failure of synthetic biological systems. Biotechnol. J. 7, 856–866 (2012).
Dixon, R.A. & Postgate, J.R. Genetic transfer of nitrogen fixation from Klebsiella pneumoniae to Escherichia coli. Nature 237, 102–103 (1972).
Dixon, R. & Cannon, F. Construction of a P plasmid carrying nitrogen fixation genes from Klebsiella pneumoniae. Nature 260, 268–271 (1976).
Moser, F. et al. Genetic circuit performance under conditions relevant for industrial bioreactors. ACS Synth. Biol. 1, 555–564 (2012).
Gorochowski, T.E., van den Berg, E., Kerkman, R., Roubos, J.A. & Bovenberg, R.A.L. Using synthetic biological parts and microbioreactors to explore the protein expression characteristics of Escherichia coli. ACS Synth. Biol. 3, 129–139 (2014).
Plackett, R.L. & Burman, J.P. The design of optimum multifactorial experiments. Biometrika 33, 305–325 (1946).
May, O., Voigt, C.A. & Arnold, F.H. in Enzyme Catalysis in Organic Synthesis: A Comprehensive Handbook 2nd edn. (eds. Drauz, K. & Waldmann, H.) Ch. 4 (Wiley-VCH Verlag, 2002).
Ran, L. et al. Genome erosion in a nitrogen-fixing vertically transmitted endosymbiotic multicellular cyanobacterium. PLoS ONE 5, e11486 (2010).
Stucken, K. et al. The smallest known genomes of multicellular and toxic cyanobacteria: comparison, minimal gene sets for linked traits and the evolutionary implications. PLoS ONE 5, e9235 (2010).
Endy, D., You, L., Yin, J. & Molineux, I.J. Computation, prediction, and experimental tests of fitness for bacteriophage T7 mutants with permuted genomes. Proc. Natl. Acad. Sci. USA 97, 5375–5380 (2000).
Densmore, D., Kittleson, J.T., Bilitchenko, L., Liu, A. & Anderson, J.C. Rule based constraints for the construction of genetic devices. Proc. 2010 IEEE ISCAS, 10.1109/ISCAS.2010.5537540 (2010).
Suh, M.H., Pulakat, L. & Gavini, N. Functional expression of the FeMo-cofacter-specific biosynthetic genes nifEN as a NifE-N fusion protein synthesizing unit in Azotobacter vinelandii. Biochem. Biophys. Res. Commun. 299, 233–240 (2002).
Fischbach, M. & Voigt, C.A. Prokaryotic gene clusters: a rich toolbox for synthetic biology. Biotechnol. J. 5, 1277–1296 (2010).
Stacy, G.S., Burris, R.H. & Evans, H.J. Biological Nitrogen Fixation (Chapman and Hall, 1992).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Chen, Y.J. et al. Characterization of 582 natural and synthetic terminators and quantification of their design constraints. Nat. Methods 10, 659–664 (2013).
Stewart, W.D., Fitzgerald, G.P. & Burris, R.H. In situ studies on nitrogen fixation with the acetylene reduction technique. Science 158, 536 (1967).
Giannoukos, G. et al. Efficient and robust RNA-seq process for cultured bacteria and complex community transcriptomes. Genome Biol. 13, r23 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Barnett, D.W., Garrison, E.K., Quinlan, A.R., Stromberg, M.P. & Marth, G.T. BamTools: a C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 27, 1691–1692 (2011).
Li, H. et al. 1000 Genome Project Data Processing Subgroup. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Acknowledgements
M.J.S., D.Z., J.C., R.N., D.B.G. and C.A.V. are supported by the US Defense Advanced Research Projects Agency (DARPA) Living Foundries grant HR0011-12-C-0067 and the US National Science Foundation Synthetic Biology Engineering Research Center (SynBERC) through grant SA5284-11210 and are also supported by the Institute for Collaborative Biotechnologies through contract W911NF-09-0001 from the US Army Research Office. The content of the information does not necessarily reflect the position or the policy of the US government, and no official endorsement should be inferred. S.B. and D.D. are supported by DARPA Living Foundries grant HR0011-12-C-0067. M.J.S. is an HHMI Fellow of the Damon Runyon Cancer Research Foundation, DRG-2129-12. Y.P. is supported by the Samsung Scholarship.
Author information
Authors and Affiliations
Contributions
M.J.S., D.Z. and C.A.V. conceived and designed the experiments and wrote the manuscript. M.J.S. performed the nifUSVWZM, monocistronic and RBS library construction and analysis. D.D. and S.B. performed the clustering analysis, wrote the design files and analyzed data. D.Z. constructed and analyzed the nifHDKY, nifENJ and complete cluster library. Y.P., D.B.G., M.B., G.G., R.N. and D.C. performed the RNA-seq experiments and analysis. L.B.A.W. and J.C. performed experiments.
Corresponding author
Ethics declarations
Competing interests
S.B. and D.D. are co-founders of Lattice Automation, Inc., a company that produces biodesign automation software.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–27, Supplementary Tables 1 and 2 and Supplementary Notes 1–11 (PDF 9314 kb)
Supplementary Data File 1.eug
Eugene file for nifUSVWZM libray design (XLS 28 kb)
Supplementary Data File 2.eug
Eugene file for newly identified rules for refactored nif cluster (XLS 42 kb)
Supplementary Data File 3.xlsx
Characterization data for nifUSVWZM library (XLSX 6348 kb)
Supplementary Data File 4.xlsx
Characterization data for nifHDKY, nifENJ, and full cluster libraries (XLSX 33 kb)
Supplementary Data File 5.xlsx
Characterization data for 16-gene monocistronic library (XLSX 69 kb)
Supplementary Data File 6.xlsx
Characterization data for 16-gene RBS swapping library (XLSX 42 kb)
Supplementary Data File 7.eug
Eugene file for 16-gene monocistronic library (XLS 27 kb)
Rights and permissions
About this article
Cite this article
Smanski, M., Bhatia, S., Zhao, D. et al. Functional optimization of gene clusters by combinatorial design and assembly. Nat Biotechnol 32, 1241–1249 (2014). https://doi.org/10.1038/nbt.3063
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3063
This article is cited by
-
Choreographing root architecture and rhizosphere interactions through synthetic biology
Nature Communications (2024)
-
A blueprint for a synthetic genetic feedback optimizer
Nature Communications (2023)
-
Toward improved terpenoids biosynthesis: strategies to enhance the capabilities of cell factories
Bioresources and Bioprocessing (2022)
-
A colorimetric method to measure in vitro nitrogenase functionality for engineering nitrogen fixation
Scientific Reports (2022)
-
A versatile active learning workflow for optimization of genetic and metabolic networks
Nature Communications (2022)