Abstract
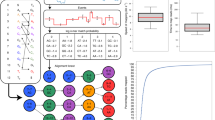
Nanopore sequencing of DNA is a single-molecule technique that may achieve long reads, low cost and high speed with minimal sample preparation and instrumentation. Here, we build on recent progress with respect to nanopore resolution and DNA control to interpret the procession of ion current levels observed during the translocation of DNA through the pore MspA. As approximately four nucleotides affect the ion current of each level, we measured the ion current corresponding to all 256 four-nucleotide combinations (quadromers). This quadromer map is highly predictive of ion current levels of previously unmeasured sequences derived from the bacteriophage phi X 174 genome. Furthermore, we show nanopore sequencing reads of phi X 174 up to 4,500 bases in length, which can be unambiguously aligned to the phi X 174 reference genome, and demonstrate proof-of-concept utility with respect to hybrid genome assembly and polymorphism detection. This work provides a foundation for nanopore sequencing of long, natural DNA strands.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shendure, J. & Lieberman Aiden, E. The expanding scope of DNA sequencing. Nat. Biotechnol. 30, 1084–1094 (2012).
McCarthy, J.J., McLeod, H.L. & Ginsburg, G.S. Genomic medicine: a decade of successes, challenges, and opportunities. Sci. Transl. Med. 5, 189sr184 (2013).
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Shendure, J. & Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 26, 1135–1145 (2008).
Mitra, R.D., Shendure, J., Olejnik, J., Edyta Krzymanska, O. & Church, G.M. Fluorescent in situ sequencing on polymerase colonies. Anal. Biochem. 320, 55–65 (2003).
Levene, M.J. et al. Zero-mode waveguides for single-molecule analysis at high concentrations. Science 299, 682–686 (2003).
Braslavsky, I., Hebert, B., Kartalov, E. & Quake, S.R. Sequence information can be obtained from single DNA molecules. Proc. Natl. Acad. Sci. USA 100, 3960–3964 (2003).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nat. Biotechnol. 26, 1146–1153 (2008).
Kasianowicz, J.J., Brandin, E., Branton, D. & Deamer, D.W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl. Acad. Sci. USA 93, 13770–13773 (1996).
Manrao, E.A. et al. Reading DNA at single-nucleotide resolution with a mutant MspA nanopore and phi29 DNA polymerase. Nat. Biotechnol. 30, 349–353 (2012).
Wanunu, M. Nanopores: a journey towards DNA sequencing. Phys. Life Rev. 9, 125–158 (2012).
Derrington, I.M. et al. Nanopore DNA sequencing with MspA. Proc. Natl. Acad. Sci. USA 107, 16060–16065 (2010).
Wallace, E.V.B. et al. Identification of epigenetic DNA modifications with a protein nanopore. Chem. Commun. (Camb.) 46, 8195–8197 (2010).
Manrao, E.A., Derrington, I.M., Pavlenok, M., Niederweis, M. & Gundlach, J.H. Nucleotide discrimination with DNA immobilized in the MspA nanopore. PLoS ONE 6, e25723 (2011).
Butler, T.Z., Pavlenok, M., Derrington, I.M., Niederweis, M. & Gundlach, J.H. Single-molecule DNA detection with an engineered MspA protein nanopore. Proc. Natl. Acad. Sci. USA 105, 20647–20652 (2008).
Cherf, G.M. et al. Automated forward and reverse ratcheting of DNA in a nanopore at 5-angstrom precision. Nat. Biotechnol. 30, 344–348 (2012).
de Bruijn, N.G. A combinatorial problem. Koninklijke Netherlandse Akademie v. Wetenschappen 49, 758–764 (1946).
Laszlo, A.H. et al. Detection and mapping of 5-methylcytosine and 5-hydroxymethylcytosine with nanopore MspA. Proc. Natl. Acad. Sci. USA 110, 18904–18909 (2013).
Needleman, S.B. & Wunsch, C.D. A general method applicable to search for similarities in amino acid sequence of 2 proteins. J. Mol. Biol. 48, 443–453 (1970).
Durbin, R., Eddy, S., Krogh, A. & Mitchison, G. Biological Sequence Analysis (Cambridge University Press, 2006).
Bashir, A. et al. A hybrid approach for the automated finishing of bacterial genomes. Nat. Biotechnol. 30, 701–707 (2012).
Koren, S. et al. Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat. Biotechnol. 30, 693–700 (2012).
Ribeiro, F.J. et al. Finished bacterial genomes from shotgun sequence data. Genome Res. 22, 2270–2277 (2012).
Shendure, J. et al. Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732 (2005).
Baaken, G., Sondermann, M., Schlemmer, C., Ruhe, J. & Behrends, J.C. Planar microelectrode-cavity array for high-resolution and parallel electrical recording of membrane ionic currents. Lab Chip 8, 938–944 (2008).
Malmstadt, N., Nash, M.A., Purnell, R.F. & Schmidt, J.J. Automated formation of lipid-bilayer membranes in a microfluidic device. Nano Lett. 6, 1961–1965 (2006).
Schibel, A.E., Edwards, T., Kawano, R., Lan, W. & White, H.S. Quartz nanopore membranes for suspended bilayer ion channel recordings. Anal. Chem. 82, 7259–7266 (2010).
Heitz, B.A., Jones, I.W., Hall, H.K. Jr., Aspinwall, C.A. & Saavedra, S.S. Fractional polymerization of a suspended planar bilayer creates a fluid, highly stable membrane for ion channel recordings. J. Am. Chem. Soc. 132, 7086–7093 (2010).
Heitz, B.A. et al. Polymerized planar suspended lipid bilayers for single ion channel recordings: comparison of several dienoyl lipids. Langmuir 27, 1882–1890 (2011).
Jain, T., Guerrero, R.J., Aguilar, C.A. & Karnik, R. Integration of solid-state nanopores in microfluidic networks via transfer printing of suspended membranes. Anal. Chem. 85, 3871–3878 (2013).
Yusko, E.C. et al. Controlling protein translocation through nanopores with bio-inspired fluid walls. Nat. Nanotechnol. 6, 253–260 (2011).
Acknowledgements
Thanks to J.J. Bartlett and B. Tickman for their help in data acquisition. This work was supported by the National Institutes of Health, National Human Genome Research Institutes (NHGRI) $1,000 Genome Program Grants R01HG005115, R01HG006321 and R01HG006283 and graduate research fellowship DGE-0718124 from the National Science Foundation (to A.A.).
Author information
Authors and Affiliations
Contributions
A.H.L., I.M.D., A.A., J.S. and J.H.G. designed the research. H.B., A.A., I.C.N., J.M.C., J.M.S., R.D. and K.D. performed the research. A.H.L., I.M.D., B.C.R., H.B., I.C.N., J.M.C. and K.W.L. analyzed the data. A.H.L., J.H.G. and J.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
I.M.D. has a commercial interest with Illumina Inc. The University of Washington has filed a provisional patent on technologies described herein.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Notes 1–5 and Supplementary Tables 1–6 (PDF 4418 kb)
Supplementary Software
Software code. (ZIP 22513 kb)
Rights and permissions
About this article
Cite this article
Laszlo, A., Derrington, I., Ross, B. et al. Decoding long nanopore sequencing reads of natural DNA. Nat Biotechnol 32, 829–833 (2014). https://doi.org/10.1038/nbt.2950
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2950
This article is cited by
-
Peptide sequencing based on host–guest interaction-assisted nanopore sensing
Nature Methods (2024)
-
MoS2 nanopore identifies single amino acids with sub-1 Dalton resolution
Nature Communications (2023)
-
High-resolution discrimination of homologous and isomeric proteinogenic amino acids in nanopore sensors with ultrashort single-walled carbon nanotubes
Nature Communications (2023)
-
Detection of phosphorylation post-translational modifications along single peptides with nanopores
Nature Biotechnology (2023)
-
De novo design of transmembrane nanopores
Science China Chemistry (2022)