Abstract
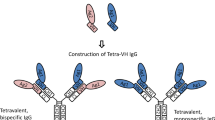
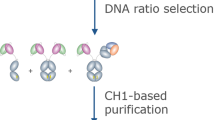
Robust generation of IgG bispecific antibodies has been a long-standing challenge. Existing methods require extensive engineering of each individual antibody, discovery of common light chains, or complex and laborious biochemical processing. Here we combine computational and rational design approaches with experimental structural validation to generate antibody heavy and light chains with orthogonal Fab interfaces. Parental monoclonal antibodies incorporating these interfaces, when simultaneously co-expressed, assemble into bispecific IgG with improved heavy chain–light chain pairing. Bispecific IgGs generated with this approach exhibit pharmacokinetic and other desirable properties of native IgG, but bind target antigens monovalently. As such, these bispecific reagents may be useful in many biotechnological applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chames, P. & Baty, D. Bispecific antibodies for cancer therapy: the light at the end of the tunnel? mAbs 1, 539–547 (2009).
Demarest, S.J. & Glaser, S.M. Antibody therapeutics, antibody engineering, and the merits of protein stability. Curr. Opin. Drug Discov. Devel. 11, 675–687 (2008).
Lum, L.G. & Thakur, A. Targeting T cells with bispecific antibodies for cancer therapy. BioDrugs 25, 365–379 (2011).
Jin, H. et al. MetMAb, the one-armed 5D5 anti-c-Met antibody, inhibits orthotopic pancreatic tumor growth and improves survival. Cancer Res. 68, 4360–4368 (2008).
Strop, P. et al. Generating bispecific human IgG1 and IgG2 antibodies from any antibody pair. J. Mol. Biol. 420, 204–219 (2012).
Labrijn, A.F. et al. Efficient generation of stable bispecific IgG1 by controlled Fab-arm exchange. Proc. Natl. Acad. Sci. USA 110, 5145–5150 (2013).
Spiess, C. et al. Bispecific antibodies with natural architecture produced by co-culture of bacteria expressing two distinct half-antibodies. Nat. Biotechnol. 31, 753–758 (2013).
Bostrom, J. et al. Variants of the antibody herceptin that interact with HER2 and VEGF at the antigen binding site. Science 323, 1610–1614 (2009).
Ridgway, J.B., Presta, L.G. & Carter, P. 'Knobs-into-holes' engineering of antibody CH3 domains for heavy chain heterodimerization. Protein Eng. 9, 617–621 (1996).
Klein, C. et al. Progress in overcoming the chain association issue in bispecific heterodimeric IgG antibodies. mAbs 4, 653–663 (2012).
Merchant, A.M. et al. An efficient route to human bispecific IgG. Nat. Biotechnol. 16, 677–681 (1998).
Schaefer, W. et al. Immunoglobulin domain crossover as a generic approach for the production of bispecific IgG antibodies. Proc. Natl. Acad. Sci. USA 108, 11187–11192 (2011).
Leaver-Fay, A., Jacak, R., Stranges, P.B. & Kuhlman, B. A generic program for multistate protein design. PLoS ONE 6, e20937 (2011).
Havranek, J.J. & Harbury, P.B. Automated design of specificity in molecular recognition. Nat. Struct. Biol. 10, 45–52 (2003).
Ashworth, J. et al. Computational redesign of endonuclease DNA binding and cleavage specificity. Nature 441, 656–659 (2006).
Grigoryan, G., Reinke, A.W. & Keating, A.E. Design of protein-interaction specificity gives selective bZIP-binding peptides. Nature 458, 859–864 (2009).
Tan, P.H., Sandmaier, B.M. & Stayton, P.S. Contributions of a highly conserved VH/VL hydrogen bonding interaction to scFv folding stability and refolding efficiency. Biophys. J. 75, 1473–1482 (1998).
Igawa, T. et al. VH/VL interface engineering to promote selective expression and inhibit conformational isomerization of thrombopoietin receptor agonist single-chain diabody. Protein Eng. Des. Sel. 23, 667–677 (2010).
De Groot, A.S. & Martin, W. Reducing risk, improving outcomes: bioengineering less immunogenic protein therapeutics. Clin. Immunol. 131, 189–201 (2009).
Nahta, R., Hung, M.C. & Esteva, F.J. The HER-2-targeting antibodies trastuzumab and pertuzumab synergistically inhibit the survival of breast cancer cells. Cancer Res. 64, 2343–2346 (2004).
Schmiedel, J., Blaukat, A., Li, S., Knochel, T. & Ferguson, K.M. Matuzumab binding to EGFR prevents the conformational rearrangement required for dimerization. Cancer Cell 13, 365–373 (2008).
Ye, X. et al. An anti-Axl monoclonal antibody attenuates xenograft tumor growth and enhances the effect of multiple anticancer therapies. Oncogene 29, 5254–5264 (2010).
Michaelson, J.S. et al. Anti-tumor activity of stability-engineered IgG-like bispecific antibodies targeting TRAIL-R2 and LTbetaR. mAbs 1, 128–141 (2009).
Gunasekaran, K. et al. Enhancing antibody Fc heterodimer formation through electrostatic steering effects: applications to bispecific molecules and monovalent IgG. J. Biol. Chem. 285, 19637–19646 (2010).
Garber, E. & Demarest, S.J. A broad range of Fab stabilities within a host of therapeutic IgGs. Biochem. Biophys. Res. Commun. 355, 751–757 (2007).
Scheer, J.M. et al. Reorienting the Fab domains of trastuzumab results in potent HER2 activators. PLoS ONE 7, e51817 (2012).
Li, B. et al. Bispecific antibody to ErbB2 overcomes trastuzumab resistance through comprehensive blockade of ErbB2 heterodimerization. Cancer Res. 73, 6471–6483 (2013).
Feige, M.J. et al. An unfolded CH1 domain controls the assembly and secretion of IgG antibodies. Mol. Cell 34, 569–579 (2009).
Pejchal, R. et al. A potent and broad neutralizing antibody recognizes and penetrates the HIV glycan shield. Science 334, 1097–1103 (2011).
Lewis, S.M. & Kuhlman, B.A. Anchored design of protein-protein interfaces. PLoS ONE 6, e20872 (2011).
McLellan, J.S. et al. Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9. Nature 480, 336–343 (2011).
Gray, J.J. et al. Protein-protein docking with simultaneous optimization of rigid-body displacement and side-chain conformations. J. Mol. Biol. 331, 281–299 (2003).
Fleishman, S.J. et al. RosettaScripts: a scripting language interface to the Rosetta macromolecular modeling suite. PLoS ONE 6, e20161 (2011).
Cooper, S. et al. Predicting protein structures with a multiplayer online game. Nature 466, 756–760 (2010).
Leaver-Fay, A. et al. Scientific benchmarks for guiding macromolecular energy function improvement. Methods Enzymol. 523, 109–143 (2013).
Franklin, M.C. et al. Insights into ErbB signaling from the structure of the ErbB2-pertuzumab complex. Cancer Cell 5, 317–328 (2004).
Casimiro, D.R., Wright, P.E. & Dyson, H.J. PCR-based gene synthesis and protein NMR spectroscopy. Structure 5, 1407–1412 (1997).
Dong, J. et al. Stable IgG-like bispecific antibodies directed toward the type I insulin-like growth factor receptor demonstrate enhanced ligand blockade and anti-tumor activity. J. Biol. Chem. 286, 4703–4717 (2011).
Doern, A. et al. Characterization of inhibitory anti-insulin-like growth factor receptor antibodies with different epitope specificity and ligand-blocking properties: implications for mechanism of action in vivo. J. Biol. Chem. 284, 10254–10267 (2009).
Powell, H.R. The Rossmann Fourier autoindexing algorithm in MOSFLM. Acta Crystallogr. D Biol. Crystallogr. 55, 1690–1695 (1999).
Evans, P.R. An introduction to data reduction: space-group determination, scaling and intensity statistics. Acta Crystallogr. D Biol. Crystallogr. 67, 282–292 (2011).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
McRee, D.E. XtalView/Xfit–A versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 125, 156–165 (1999).
Acknowledgements
This work was supported by the Lilly Research Laboratories and the Lilly Research Award Program (LRAP). We thank J. Hannah and B. Gutierrez for their help with transient transfection, R. Yuan and D. He for assistance with protein purification, M. Batt and J. Fitchett for their training and up keep of the mass spectrometry facility at Lilly. B. Stranges provided suggestions for the computational docking protocol. Use of the Advanced Photon Source, an Office of Science User Facility operated for the US Department of Energy (DOE) Office of Science by Argonne National Laboratory, was supported by the US DOE under Contract No. DE-AC02-06CH11357. Use of the Lilly Research Laboratories Collaborative Access Team (LRL-CAT) beamline at Sector 31 of the Advanced Photon Source was provided by Eli Lilly Company, which operates the facility. We thank S. Wasserman, S. Sojitra, J. Koss for data collection and operation of the beamline.
Author information
Authors and Affiliations
Contributions
All authors contributed to the concepts of the study. S.M.L. and B.K. performed the computational design, with advice from A.L.-F. B.K., A.K.C., S.M.T., S.M.L., S.J.D. and X.W. contributed to the rational designs. X.W., A.S., H.L.R., E.M.C., E.M.S., G.G., C.H., F.H., C.H.-E. and S.J.D. performed the experimental work. A.P. and S.A. performed the crystallization and structure determinations. B.K. and S.J.D. conceived the project. Writing of the paper was done in close collaboration by B.K., S.A., S.M.L. and S.J.D.
Corresponding authors
Ethics declarations
Competing interests
X.W., A.P., A.S., F.H., E.M.S., C.H., A.K.C., S.T., S.A. and S.J.D. are employees of Eli Lilly
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1–5 and Supplementary Protocols 1–3 (PDF 10445 kb)
Rights and permissions
About this article
Cite this article
Lewis, S., Wu, X., Pustilnik, A. et al. Generation of bispecific IgG antibodies by structure-based design of an orthogonal Fab interface. Nat Biotechnol 32, 191–198 (2014). https://doi.org/10.1038/nbt.2797
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2797
This article is cited by
-
CD20/TNFR1 dual-targeting antibody enhances lysosome rupture-mediated cell death in B cell lymphoma
Cancer Immunology, Immunotherapy (2023)
-
Bispecific T cell engagers: an emerging therapy for management of hematologic malignancies
Journal of Hematology & Oncology (2021)
-
Production of IgG1-based bispecific antibody without extra cysteine residue via intein-mediated protein trans-splicing
Scientific Reports (2021)
-
A new approach to produce IgG4-like bispecific antibodies
Scientific Reports (2021)
-
Cancer immune therapy with PD-1-dependent CD137 co-stimulation provides localized tumour killing without systemic toxicity
Nature Communications (2021)