Abstract
Genes underlying mutant phenotypes can be isolated by combining marker discovery, genetic mapping and resequencing, but a more straightforward strategy for mapping mutations would be the direct comparison of mutant and wild-type genomes. Applying such an approach, however, is hampered by the need for reference sequences and by mutational loads that confound the unambiguous identification of causal mutations. Here we introduce NIKS (needle in the k-stack), a reference-free algorithm based on comparing k-mers in whole-genome sequencing data for precise discovery of homozygous mutations. We applied NIKS to eight mutants induced in nonreference rice cultivars and to two mutants of the nonmodel species Arabis alpina. In both species, comparing pooled F2 individuals selected for mutant phenotypes revealed small sets of mutations including the causal changes. Moreover, comparing M3 seedlings of two allelic mutants unambiguously identified the causal gene. Thus, for any species amenable to mutagenesis, NIKS enables forward genetics without requiring segregating populations, genetic maps and reference sequences.
Similar content being viewed by others
Main
Forward genetic screens have been of fundamental importance in elucidating biological mechanisms in model species1. Their success, however, has relied on the feasibility of mutant gene isolation. Identification of causal mutations typically begins with genetic mapping, followed by candidate gene sequencing and complementation studies using transformation. Advances in DNA sequencing technologies have tremendously accelerated genetic mapping by combining bulk segregant analysis, that is, pooling recombinant genomes, with whole-genome sequencing, usually referred to as mapping by sequencing2,3. This approach is now becoming standard for mutation mapping and identification in many model species3,4,5,6,7,8,9,10,11,12 and has even been applied to decipher quantitative traits with complex genetic architectures13,14. Recently, mutagen-induced changes have been used as novel markers, allowing mapping of mutations using isogenic mapping populations10,15. Nevertheless, all mapping-by-sequencing methods rely on resequencing, a method for whole-genome reconstruction based on aligning sequences to a reference sequence. Therefore, this requirement restricts the application of the technique to species for which such a reference genome sequence is available.
Many reference-sequence assembly projects are currently in progress, including ones for most of the major crop species and breeding animals. However, even with an existing reference sequence, extending mapping-by-sequencing methods beyond the sequenced reference accessions has proved technically challenging. Mutant alleles of genes that are not present in the reference sequence cannot be identified within resequencing data alone. In particular, fast-evolving genes, such as those involved in disease resistance, might not always be represented in the reference sequence16,17.
Alternative solutions for mapping-by-sequencing in species without reference sequences have been proposed, such as mapping-by-sequencing based on reference sequences of related species or expressed sequence tag collections11,18. However, all of these methods greatly rely on low sequence divergence and high levels of synteny between the mutant genome and alignment target. Recently, methods for direct genome comparison of multiple samples without a reference sequence were introduced, but none has proven to be accurate and precise enough for the identification of mutations19,20,21.
NIKS is a method for reference-free genome comparison based solely on the frequencies of short subsequences within whole-genome sequencing data. It is geared toward identifying mutagen-induced, small-scale, homozygous differences between two highly related genomes, independent of their inbred or outbred background, and provides a route to identification of mutations without requiring any prior information about reference sequences or genetic maps.
Results
Principles and performance of NIKS
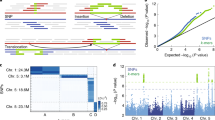
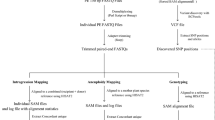
NIKS relies on the analysis of k-mers, which are defined as sub-sequences of length k of a sequencing read. NIKS starts by assessing the frequency of each k-mer within the sequencing data of each sample using the k-mer–counting software Jellyfish22. K-mers that overlap with sequencing errors will be of low frequency, as these errors are not present in all reads from the corresponding region, and it is therefore possible to separate them from reads that are error free (Fig. 1). Amplification artifacts in sequencing can increase the frequency of error-based k-mers and have the potential to introduce error-based k-mers into the set of error-free k-mers. Thus, filtering for identical reads before running NIKS can reduce the impact of sequencing errors. Differences between genomes give rise to multiple, sample-specific and overlapping k-mers. NIKS identifies and then merges all sample-specific k-mers into longer sequences (or seeds).
Whole-genome sequencing data of two related genomes is analyzed for the frequency of all k-mers. K-mer frequency histograms provide the power to distinguish between native k-mers (area highlighted in gray) and k-mers overlapping with sequencing errors. Comparing the two sets of k-mers of two highly related genomes discloses sample-specific, overlapping k-mers that result from subtle differences between the genomes. These sample-specific k-mers are then merged to a seed if they can be paired with a homologous, but not identical, seed in the other sample. Read pairs that share at least one k-mer with a seed pair will be used for local assemblies. This results in contigs that are centered on the mutated sites.
To identify only those differences that were introduced during mutagenesis, NIKS considers only those seeds that feature a homologous, but not identical, seed in the second sample (seed pairing; Supplementary Fig. 1a). In fact, the two seeds of such a seed pair represent the wild-type and the mutant allele, respectively, and are distinguished only by the mutagen-induced mutations themselves. Mutations that are spaced by less than k – 1 bp and small indels will be combined in one elongated seed (Supplementary Fig. 1b,c). Larger indels might not result in one contiguous seed for the wild-type allele, but can be too complex to be assembled into one seed. Thus, NIKS additionally screens for other types of seed pairing. Seeds representing deletions can match up to two seeds in the other sample. These two seeds represent the breakpoints of the deletion in the wild-type sample. If both breakpoints of the deletion yield a seed, NIKS will identify a two-sided junction, and if only one breakpoint generates a seed it will produce a one-sided junction (Supplementary Fig. 1d,e).
Like homozygous mutations, heterozygous mutations and mutations in repetitive regions will introduce novel k-mers into the mutant sample. However, unlike for homozygous mutations, both the wild-type and the novel mutant allele will be present in the mutant sample. The presence of the wild-type allele in the mutant sample will then prevent the generation of a seed pair. In detail, NIKS records all k-mers that support the wild-type allele in the mutant sample and vice versa (mirror counts). Only mutations that feature less than a total of k k-mers from the respective other allele will be considered as homozygous.
In a last step, NIKS generates local de novo assemblies to extend the sequences associated with the mutated site23. Usually, this results in contigs of multiple hundreds of base pairs in length, including the mutated site. These contigs allow for functional annotation to assess the putative effect of mutations either by de novo gene predictions or alignments against known gene annotations.
To evaluate the performance of NIKS, we simulated 160 whole-genome sequencing experiments by first introducing ∼2,000 random, homozygous, single-base mutations into the mouse and maize genome reference sequences and then sequencing at 17, 25, 35 and 50-fold genome coverage24,25 (Supplementary Notes). The reference genomes are comparable in size—the mouse genome is 2.6 Gb and maize is 2.0 Gb—but differ drastically in their repeat content (Supplementary Fig. 2). To distinguish between mutations in unique regions and those in repeats, we classified each position into one of three classes according to the number of overlapping, repetitive k-mers (Supplementary Notes).
We compared the simulated sequencing data of each mutant to the sequencing data of the reference strain using NIKS (Supplementary Fig. 3). Within unique regions NIKS' sensitivity was >90%, and slightly increased with higher coverage levels. Notably, it was almost the same in the mouse and the maize experiments, indicating that sensitivity in unique regions, which typically represent large parts of the complement of genes, is not influenced by the overall repeat content of a genome. As expected, NIKS did not identify any of the mutations within repetitive regions, though it did identify some of the mutations in regions where repetitive and unique reads are present. From 25× genome coverage and up, the percentage of correctly identified mutations among all predictions (positive predictive value) was >98% across all experiments and backgrounds.
In addition to mutation identification, marker development in nonmodel organisms would profit from reference-independent methods. Applying NIKS to a whole-genome sequencing data set from two natural varieties of Arabidopsis thaliana, we were able to identify nearly 300,000 single-nucleotide polymorphisms (Supplementary Table 1). However, because of the high density of polymorphisms between natural accessions, dedicated tools, such as Cortex21, may be more suitable for this task (Supplementary Notes).
Testing NIKS on seven previously analyzed rice mutants
Ethyl methanesulfonate (EMS) mutations have been identified by resequencing isogenic bulked, or pooled, recombinants of seven rice mutants10. To reduce the high load of mutations introduced by EMS, researchers10 analyzed DNA pools of backcrossed recombinants rather than single-mutant genomes. The mutations were induced in the elite cultivar background of Hitomebore, which has no assembled reference sequence. By creating a pseudo-reference sequence and resequencing bulk segregant populations at coverage levels of 12–17×, the authors identified 3–11 putative candidate EMS mutations for six of the seven mutants.
We reanalyzed all seven samples with NIKS (Supplementary Notes and Supplementary Fig. 4). Because NIKS performs a two-sample comparison, each of the seven samples was compared to six other samples separately. After removing mutations identified in only one comparison and keeping only canonical EMS mutations, we retained 7–21 mutations per sample (Table 1 and Supplementary Tables 2 and 3). To compare both analyses, we carried out a functional characterization of the mutations using the reference annotations of rice26. For four mutants, NIKS revealed the same candidate genes reported previously10. Among those was the experimentally validated mutation in the sample Hit1917-pl1. For two of the remaining mutants, our analysis revealed a distinct set of causal candidate genes. No candidate mutation had been previously reported for the last mutant. Our analysis revealed a mutation in an intron that is putatively retained in an alternatively spliced isoform, which is expressed in various developmental stages (Supplementary Notes and Supplementary Fig. 5). In summary, NIKS identified candidate mutations for all seven rice mutants, appearing to be at least as accurate in our hands as the previously published method10, and using the reference sequence only for functional analysis.
Mutation identification without reference or genetic map
To perform NIKS on a undescribed mutant of a nonmodel species, we selected A. alpina mutants from a recent EMS mutagenesis screen27. A. alpina is a perennial Brassicaceae species with an estimated genome size of 375 Mb for which no reference sequence, annotation or genetic map is available. One of the selected mutants floral defective 1 (fde1) displayed floral homeotic defects (Fig. 2a). A second mutant perpetual flowering 1-1 (pep1-1) was previously shown, through a homology-based candidate gene approach, to carry a splice-site lesion in the PEP1 gene that is responsible for the phenotype27 (Fig. 2b). Comparing the two genomes using NIKS, we aimed to identify the unknown lesion in fde1 and simultaneously confirm the pep1-1 mutation.
(a) fde1 mutant flowers are deformed compared to flowers in wild-type plants. (b) pep1-1 mutant plants flower without vernalization compared to wild type, which has an obligate requirement for vernalization to flower. (c) Alignment of a NIKS contig of pep1-1 against the A. thaliana reference sequence reveals the splice-site mutation in the A. alpina ortholog of FLOWERING LOCUS C (FLC) previously reported to be responsible for the pep1-1 phenotype. The mutant base is shown in red, the exonic regions in A. thaliana are shown in yellow, the inferred exon of A. alpina in blue and all noncoding nucleotides in gray. (d) Alignment of a NIKS contig of fde1 against the A. thaliana reference sequence revealing a putative amino acid change likely to alter an aspartic acid to an asparagine within the 4th coding exon of the A. alpina ortholog of AP2. The frame used to translate the nucleotide sequence to an amino acid sequence was inferred from the annotation of A. thaliana. Causal mutations are shown in red. Exonic sequences associated with the mutation are shown in blue and exons of A. thaliana in yellow. Only relevant parts of the alignments are shown.
To reduce the high load of mutations, we compared pooled DNA of backcrossed recombinants rather than single-mutant genomes, as described for the rice mutants10 (Supplementary Notes). After backcrossing to the nonmutagenized progenitor, F1 individuals were self-pollinated and DNA of 97 and 86 F2 plants of pep1-1 and fde1, respectively, with the mutant phenotype were pooled for whole-genome sequencing (Supplementary Table 4). Applying NIKS to whole-genome sequencing data at a genome fold coverage of 67 (pep1-1) and 158 (fde1), we found 29 seed pairs that represent homozygous mutagen-induced changes (Supplementary Notes, Supplementary Fig. 6a,b, Supplementary Fig. 7 and Supplementary Table 5). To demonstrate that such high levels of coverage were not necessary to identify these mutations, we repetitively bootstrapped the data at different coverage levels and still retained, on average, >20 at 17× coverage (Supplementary Notes and Supplementary Table 6). All 29 changes were C:G ↔ T:A mutations, as expected for EMS mutagenesis28. Based on the known bias of EMS mutations, we defined all T and A alleles as mutant alleles, thereby assigning 16 mutations to fde1 and 13 to pep1-1, respectively (Table 2). We have resequenced all 16 fde1 mutations by PCR amplification of adjacent regions followed by Sanger sequencing and verified the presence of all predicted changes (Supplementary Table 7). Finally, local assemblies of all read pairs, that harbored k-mers, shared with the seed pairs reconstructed contigs up to 922 bp in length, surrounding all 29 mutations in both genomes. These were used to unambiguously identify the fde1 mutation.
FDE1 is the ortholog of A. thaliana AP2
We annotated the effect of the 29 mutations using two independent methods. First, all 29 contigs were aligned against a collection of publicly available full-length reference sequences29 (NCBI Genomic Reference Sequences). Surprisingly, we could not identify any reliable hits for seven of these contigs (e-value, 1e-05), whereas all others provided reliable hits against the reference sequence of A. thaliana, as well as against other genomes30. For uniformity, we considered only hits against A. thaliana (Supplementary Notes). Based on these alignments, we inferred the structure of putative genes of A. alpina and assigned a putative effect to each of these mutations (Table 2). Six of the pep1-1 mutations overlapped with the annotation of genes. Among those, we identified the known causal mutation, aligned to a splice donor site of the PEP1 ortholog FLC27,31 (Fig. 2c). Six of the fde1 mutations aligned against genes. One of them altered the putative amino acid sequence of the A. alpina homolog APETALA 2 (AaAP2) (Fig. 2d). The A. thaliana ap2 mutant alleles cause a flower deformation very similar to that observed in fde1 (ref. 32) (Fig. 2a).
In a second, independent attempt to annotate the effect of mutations, we performed de novo gene predictions using the 29 contigs harboring the wild-type allele followed by the annotation of the effect of the respective mutation (Table 2 and Supplementary Notes). Notably, 19 out of 22 predicted effects were identical to the homology-based annotation, including both causal changes. Three of them differed, although one showed homology to a transposable element, which was excluded in the homology-based annotation. In addition, for one of the seven contigs without a significant homology, a gene model was predicted.
To confirm that the mutation in AaAP2 caused the flower phenotype of the fde1 mutant, we further characterized three additional A. alpina mutants, designated fde1-2, fde1-3 and fde1-4, displaying the same flower deformation as fde1 (Supplementary Fig. 8a–c). The assembled contig sequence surrounding the AaAP2 mutation was used to design primers to amplify AP2 cDNA from the additional mutants. fde1-2 contained a mutation that causes an amino acid change; fde1-3, a mutation that causes an early stop codon; and fde1-4, a splice site mutation, which was close to the original fde1 mutation (Supplementary Notes). The presence of mutations in AaAP2 in four of the isolated mutants corroborated that we had identified the causal mutation (Supplementary Fig. 8d).
Thus, NIKS correctly identified both causal mutations among a small set of candidate mutations, which could be functionally annotated through public databases and independently annotated de novo (Table 2).
NIKS identifies footprints of a 169-kb deletion
In contrast to chemically induced mutations, those induced by fast neutron radiation (FNR) can be more complex, such as deletions that span multiple kilobases33. To determine NIKS' power to identify complex changes, we applied it to the genome of the rice mutant hebiba, which was isolated from an FNR-mutagenized population of Oryza sativa ssp. japonica cv. Nihonmasari and displayed perturbed seedling photomorphogenesis34. Sixty-two hebiba mutants, identified from a segregating population, and a pool of 100 wild-type genomes were sequenced up to 80.7- and 99.5-fold coverage, respectively, and compared against each other (Supplementary Notes, Supplementary Fig. 9 and Supplementary Tables 4 and 5). In addition to 92 small-scale changes, NIKS identified footprints for three large-scale changes, represented by three contigs assembled in the mutant sample (Supplementary Notes and Supplementary Table 8). Two of these recapitulated single-base mutations in repetitive regions. To annotate a potential large-scale disruption identified by the remaining contig, we aligned the contig against the reference sequence using BLAST, which revealed a 169-kb deletion that was subsequently confirmed by PCR (Supplementary Notes and Supplementary Fig. 10). Simultaneously with our work, it has been shown that the hebiba phenotype is, in fact, induced by this deletion35.
Owing to the complexity and length of the deleted sequence, NIKS could not assemble the complete 169-kb sequence of the wild-type allele, and a reference sequence was required for the identification of the deletion. However, NIKS did correctly assemble the newly generated junction at the deletion site into a 587-bp contig.
Mutation identification by analyzing two independent alleles
Backcrossing to wild-type progenitors has been successfully used to reduce the high number of noncausal EMS mutations10. However, this requires at least two more generations and is not practical for species with long generation times. Recently, we proposed that the most straightforward approach for the identification of mutations would be direct sequencing of two or more allelic mutants36. Although each mutant genome would feature many mutations, only genes with lesions in all mutants need to be considered as candidates (Supplementary Fig. 11). The expected random overlap of disrupted genes between two independently mutagenized genomes is extremely small and effectively disappears when more than two alleles are considered (Supplementary Notes and Supplementary Fig. 12).
As a proof-of-concept demonstration, we compared the genomes of pep1-1 and pep1-2, which were isolated in the same EMS screen27 and flower without vernalization. Both mutants share lesions in the same gene and form a complementation group; F1 generation plants resulting from a cross between pep1-1 and pep1-2 also flower without vernalization. The mutation in pep1-2 was confirmed by targeted sequencing of PEP1, revealing an early stop codon in the second exon (Supplementary Notes and Fig. 3).
(a) pep1-1 and pep1-2 flower without vernalization. pep1-2 does not complement the early flowering phenotype of pep1-1. F1 (pep1-1/2) plants, resulted from the cross of pep1-1 with pep1-2, flower in long-day conditions suggesting that they carry independent lesions within the same gene. (b) Flowering time analysis measured as number of leaves at flowering. pep1-1, pep1-2 and their F1 hybrid flowered whereas wild-type plants never flower in long days. Error bars, mean ± s.d, n = 12. (c) Alignment of a NIKS contig of pep1-2 against the A. thaliana reference sequence identifies a premature stop codon introduced in the second exon of PEP1. The frame used to annotate the nucleotide sequence was inferred through alignment to A. thaliana. The causal mutation is shown in red. Exonic sequence associated with the mutation is shown in blue and the homologous exon of A. thaliana in yellow.
We generated 51- and 105-fold whole-genome coverage sequencing data from pools of 35 M3plants, which were derived from seeds after self-pollination of the original mutant plants, of pep1-1 and pep1-2, respectively. NIKS identified 779 seed pairs, each of them revealing one genomic difference between pep1-1 and pep1-2 (Supplementary Table 9 and Supplementary Fig. 6c,b). Of these 779 changes, only five lacked the canonical EMS change. All others could be assigned to either of the mutants and were annotated by aligning the mutation-associated contigs against the reference sequence of A. thaliana. Of the mutant contigs of pep1-1 and pep1-2, 165 and 94, respectively, aligned to genes. However, only three of these genes were common for both mutants (Table 3). Functional annotation revealed only one gene, for which the mutations in both genomes affected the integrity of the gene. This gene was PEP1, the A. alpina ortholog of FLC and causal for the phenotype of pep1-1 and pep1-2 (ref. 27).
Discussion
We introduced strategies for identification of mutagen-induced homozygous changes in unique regions in both of two highly related genomes, such as mutant and wild-type genomes, without requiring a reference genome. For this purpose, we present a whole-genome comparison method, NIKS, which accurately predicts subtle differences based on whole-genome sequencing data alone.
However, mutants may contain hundreds of mutagen-induced changes that hamper the direct readout of causal mutations. One way to reduce the number of mutations is backcrossing to the wild-type, followed by sequencing of pooled genomes of F2 individuals10,15. With this strategy we identified an unknown mutation in A. alpina. To investigate whether this would have been possible with conventional means, we performed short-read alignment against the reference sequence of A. thaliana and de novo assembly followed by whole-genome comparisons, but with neither of these attempts was it possible to unambiguously identify all mutagen-induced changes (Supplementary Notes).
To classify genomic differences as mutant or wild-type allele, we used mutagen-specific biases for the type of mutation they introduce. Of all changes identified in our A. alpina data, 98.8% were canonical EMS changes. However, even if the mutagen does not introduce a bias in the mutational spectrum, analyzing multiple mutants or the wild-type genome unambiguously identifies mutant and wild-type alleles. In the current implementation, NIKS supports the comparison of two samples at a time, which conceptually could be expanded to an unlimited number of samples, similar to the approaches implemented in Cortex, a sophisticated usage of colored de Bruijn graphs21.
Cortex and similar tools facilitate simultaneous whole-genome assembly and variation identification19,20,21. Whole-genome assembly structures can leverage the identification of complex differences between the samples, which is a nontrivial task when considering genome structures such as repeats and low-complexity regions. Such tools can additionally integrate a reference sequence into the assembly structure and thereby further compensate for difficulties while assembling long deletions de novo. Even without a reference sequence, NIKS identified the footprints of one large-scale deletion within a rice mutant, but it assembled only the mutant allele correctly. The reconstruction of the respective wild-type allele still required a reference sequence.
If one wants to pinpoint the causal mutation among a set of candidate mutations, it is helpful to annotate the effect of mutations on genes, either by imputing public gene annotation10 or de novo gene annotation. De novo gene predictions turned out to be very similar to homology-based gene predictions, indicating that annotation of the effect of mutations can be done independently of prior knowledge.
Another solution to reduce the number of candidate mutations emerges by direct sequencing of genomes carrying two or more independent alleles, followed by a subsequent search for genes that harbor unique mutations in the same gene in multiple genomes. In a proof-of-principle experiment, we screened the M3 genomes of two allelic A. alpina mutants for common genes carrying mutations, and we were able to unambiguously identify the causal gene. This analysis required prior identification of different alleles, which might not be straightforward for all species and phenotypes, but on the other hand, bypasses the generation of mapping populations. To our knowledge, this describes the first successful report of mutation identification using whole-genome sequencing of mutant genomes of a complementation group without relying on any kind of recombination.
Our method opens up many possibilities of forward genetics supported by whole-genome sequencing for any species that is amenable to mutagen screens. Thus, the approach has the potential to ease the access to all those species without reference sequences whose complex genomes defy current assembly methods. However, the largest impact might come from the use of NIKS to clone genes from nonmodel species that exhibit important traits36.
Given that sequencing costs continue to decline, we anticipate that the genomes of all mutants of a forward genetic screen can be sequenced. By knowing all genomic and phenotypic differences, researchers will be able to group mutants into putative in silico allelic groups. With this information at hand, a small number of targeted complementation tests will serve as a first line of mutation validation for all those genes that feature multiple alleles in the screen. Depending on the size of the screen and on the number of genes that contribute to the phenotype of interest, this can be a powerful way to identify multiple mutants simultaneously. For example, simulations of forward genetic screens show that the analysis of 100 mutant genomes would be enough to identify over 27 allelic groups, assuming a screen size of 40,000 individuals and a phenotype that includes 75 genes (Supplementary Fig. 13). NIKS, thus, has the potential to reduce the workload of mutant identification to nothing more than whole-genome sequencing and comparison.
Methods
NIKS implementation.
NIKS is a Java and bash-based pipeline and implements all analysis steps performed in this study, including k-mer counting and selection, seed generation and pairing, and mutation-centric contig assembly. NIKS can be downloaded from https://sourceforge.net/projects/niks/.
k-mer counting and selection.
NIKS starts by assessing the frequency of all k-mers within whole-genome sequencing data of each sample using the efficient k-mer–counting software Jellyfish22. For k > 31 NIKS implements simple counting and sorting algorithms. Here we used k = 31 to make k as large as possible, while still being able to utilize the advantages of Jellyfish. In general, k needs to be smaller than read size, but large enough to allow for unique assignments to the genome. K-mers that overlap with sequencing errors will be of no use, which implies a k value significantly smaller than read length. Thus, the optimal choice of k depends on read quality and genome complexity, and cannot be generalized. In practical applications, however, it might be advantageous to run NIKS with multiple different values for k.
Defining two-thirds of the first local minimum in a histogram of k-mer frequencies as an intrinsic cut-off, NIKS can distinguish between native and sequencing error–based k-mers (Fig. 1). Insufficient coverage will hamper identification of this local minimum and make NIKS consider true all k-mers that occur at least twice. This allows reliable querying of samples for the presence or absence of k-mers. NIKS then extracts all k-mers present in one sample, but not in the other (sample-specific k-mers).
Seed generation and pairing.
Single point mutations give rise to k sample-specific, overlapping k-mers. Joining k-mers that overlap by k – 1 bases merges sample-specific k-mers to seeds. For this, NIKS selects all k-mers of a sample that share at least one m-mer with the sample-specific k-mers of the other sample, where m is smaller than k (here, k = 31 and m = 25). Thereby, NIKS can preselect seeds that will yield valid seed pairs while allowing for disruptions at each end of a seed (Supplementary Fig. 1). This set of k-mers is then screened for k-mers that do not overlap with other sample-specific k-mers on one side, in order to identify putative start or end points of seeds (end k-mers). Starting with each end k-mer, an exhaustive walk, by combining k-mers that overlap by k – 1 bases, is conducted without adding the same k-mer twice and stopping if another end k-mer is reached. To avoid repetitive regions and to reduce the computational load, seeds with the summed coverage of the combined k-mers larger than 10,000 bases are discarded.
In the case of point mutations, optimal seeds are 2*k – 1 bp long and centered on the mutated sites, but in rare cases some seeds do not extend completely, resulting in seeds that are shorter than expected. Multiple closely linked mutations introduce (k – 1 + s) sample-specific k-mers, where s refers to the length of the mutated region. Long indels also give rise to sample-specific overlapping k-mers. In particular, at the breakpoint of deletions, novel sequence is introduced. If this sequence is unique in comparison to the contrasting genome it will generate unique k-mers, which will give rise to sample-specific seeds. NIKS identifies these seeds by their partial homology to the breakpoint in the other sample (Supplementary Fig. 1c–e). Seeds featuring sequence similarity to multiple other seeds' ends in the other sample are excluded.
Wrongly scored phenotypes or undetected sequencing errors can lead to the presence of k-mers, which represent the nonmutagenized allele and can mask real mutations. By default, NIKS discards seeds with support from more than k k-mers for the second allele in the other sample. This discards seeds with at least two contradicting reads covering the mutation. As seed pairing requires consistent support from both samples, it introduces high levels of specificity into NIKS.
Mutation-centric assemblies.
To extend all valid seeds, NIKS extracts all read pairs sharing at least one of the k-mers with a seed. Each such set of read pairs is assembled with Velvet, applying standard parameters, except for the sample specific insert length and automated estimation of the coverage cutoff23. Assembled contigs that are longer than the respective seed, but perfectly match the seed, replace the seed.
Accession code.
Short read data have been deposited in the EBI short read archive under accession number ERP002265.
Accession codes
References
Page, D.R. & Grossniklaus, U. The art and design of genetic screens: Arabidopsis thaliana. Nat. Rev. Genet. 3, 124–136 (2002).
Michelmore, R.W., Paran, I. & Kesseli, R.V. Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc. Natl. Acad. Sci. USA 88, 9828–9832 (1991).
Schneeberger, K. et al. SHOREmap: simultaneous mapping and mutation identification by deep sequencing. Nat. Methods 6, 550–551 (2009).
Blumenstiel, J.P. et al. Identification of EMS-induced mutations in Drosophila melanogaster by whole-genome sequencing. Genetics 182, 25–32 (2009).
Cuperus, J.T. et al. Identification of MIR390a precursor processing-defective mutants in Arabidopsis by direct genome sequencing. Proc. Natl. Acad. Sci. USA 107, 466–471 (2010).
Birkeland, S.R. et al. Discovery of mutations in Saccharomyces cerevisiae by pooled linkage analysis and whole-genome sequencing. Genetics 186, 1127–1137 (2010).
Zuryn, S., Le Gras, S., Jamet, K. & Jarriault, S. A strategy for direct mapping and identification of mutations by whole genome sequencing. Genetics 186, 427–430 (2010).
Laitinen, R.A.E., Schneeberger, K., Jelly, N.S., Ossowski, S. & Weigel, D. Identification of a spontaneous frame shift mutation in a nonreference Arabidopsis accession using whole genome sequencing. Plant Physiol. 153, 652–654 (2010).
Austin, R.S. et al. Next-generation mapping of Arabidopsis genes. Plant J. 67, 715–725 (2011).
Abe, A. et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 30, 174–178 (2012).
Galvão, V.C. et al. Synteny-based mapping-by-sequencing enabled by targeted enrichment. Plant J. 71, 517–526 (2012).
Leshchiner, I. et al. Mutation mapping and identification by whole genome sequencing. Genome Res. 22, 1541–1548 (2012).
Ehrenreich, I.M. et al. Dissection of genetically complex traits with extremely large pools of yeast segregants. Nature 464, 1039–1042 (2010).
Turner, T.L., Stewart, A.D., Fields, A.T., Rice, W.R. & Tarone, A.M. Population-based resequencing of experimentally evolved populations reveals the genetic basis of body size variation in Drosophila melanogaster. PLoS Genet. 7, e1001336 (2011).
Hartwig, B., James, G.V., Konrad, K., Schneeberger, K. & Turck, F. Fast Isogenic mapping-by-sequencing of ethyl methanesulfonate-induced mutant bulks. Plant Physiol. 160, 591–600 (2012).
Cai, D. et al. Positional cloning of a gene for nematode resistance in sugar beet. Science 275, 832–834 (1997).
Song, J. et al. Gene RB cloned from Solanum bulbocastanum confers broad spectrum resistance to potato late blight. Proc. Natl. Acad. Sci. USA 100, 9128–9133 (2003).
Wurtzel, O., Dori-Bachash, M., Pietrokovski, S., Jurkevitch, E. & Sorek, R. Mutation detection with next-generation resequencing through a mediator genome. PLoS ONE 5, e15628 (2010).
Peterlongo, P., Schnel, N., Pisanti, N., Sagot, M.F. & Lacroix,, V. Identifying SNPs without a reference genome by comparing raw reads. in String Processing and Information Retrieval (eds. Chavez, E. & Lonardi, S.) 147–158 (Springer, 2010).
Ratan, A., Zhang, Y., Hayes, V.M., Schuster, S.C. & Miller, W. Calling SNPs without a reference sequence. BMC Bioinformatics 11, 130 (2010).
Iqbal, Z., Caccamo, M., Turner, I., Flicek, P. & McVean, G. De novo assembly and genotyping of variants using colored de Bruijn graphs. Nat. Genet. 44, 226–232 (2012).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Zerbino, D.R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Church, D.M. et al. Lineage-specific biology revealed by a finished genome assembly of the mouse. PLoS Biol. 7, e1000112 (2009).
Schnable, P.S. et al. The B73 maize genome: complexity, diversity, and dynamics. Science 326, 1112–1115 (2009).
International Rice Genome Sequencing Project. The map-based sequence of the rice genome. Nature 436, 793–800 (2005).
Wang, R. et al. PEP1 regulates perennial flowering in Arabis alpina. Nature 459, 423–427 (2009).
Greene, E.A. et al. Spectrum of chemically induced mutations from a large-scale reverse-genetic screen in Arabidopsis. Genetics 164, 731–740 (2003).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Wang, R. et al. Aa TFL1 confers an age-dependent response to vernalization in perennial Arabis alpina. Plant Cell 23, 1307–1321 (2011).
Bowman, J.L., Smyth, D.R. & Meyerowitz, E.M. Genes directing flower development in Arabidopsis. Plant Cell 1, 37–52 (1989).
Li, X. et al. A fast neutron deletion mutagenesis-based reverse genetics system for plants. Plant J. 27, 235–242 (2001).
Riemann, M. et al. Impaired induction of the jasmonate pathway in the rice mutant hebiba. Plant Physiol. 133, 1820–1830 (2003).
Riemann, M. et al. Identification of rice ALLENE OXIDE CYCLASE mutants and the function of jasmonate for defence against Magnaporthe oryzae. Plant J. (accepted) http://dx.doi.org/10.1111/tpj.12115 (24 January 2013).
Schneeberger, K. & Weigel, D. Fast-forward genetics enabled by new sequencing technologies. Trends Plant Sci. 16, 282–288 (2011).
Acknowledgements
The authors would like to thank D. Weigel for helpful discussions and suggestions on the manuscript, G. Marcais for support in using the Jellyfish software, P. Nick for providing hebiba mutant seeds, M. Riemann for support during the hebiba mutant analysis and the Max Planck Genome Center Cologne for their help on mutant sequencing. This work has been supported by grant DFG SPP1530 (“Flowering time control: from natural variation to crop improvement”) (M.C.A. and K.S.).
Author information
Authors and Affiliations
Contributions
K.S. and G.C. designed the study. K.J.V.N. designed and implemented NIKS. M.C.A., C.G., B.H., U.P. and F.T. provided biological material and performed experiments. K.J.V.N., G.V.J. and K.S. performed data analysis. K.S. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Notes, Supplementary Figures 1–13 and Supplementary Tables 1–9 (PDF 610 kb)
Rights and permissions
About this article
Cite this article
Nordström, K., Albani, M., James, G. et al. Mutation identification by direct comparison of whole-genome sequencing data from mutant and wild-type individuals using k-mers. Nat Biotechnol 31, 325–330 (2013). https://doi.org/10.1038/nbt.2515
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2515
This article is cited by
-
Broadening the horizon of crop research: a decade of advancements in plant molecular genetics to divulge phenotype governing genes
Planta (2022)
-
Growth adaptability and foreign gene stability of TaLEA transgenic Populus simonii × nigra
Annals of Forest Science (2021)
-
Role of BrSDG8 on bolting in Chinese cabbage (Brassica rapa)
Theoretical and Applied Genetics (2020)
-
Principles and approaches of association mapping in plant breeding
Tropical Plant Biology (2020)
-
Comparative Subsequence Sets Analysis (CoSSA) is a robust approach to identify haplotype specific SNPs; mapping and pedigree analysis of a potato wart disease resistance gene Sen3
Plant Methods (2019)