Abstract
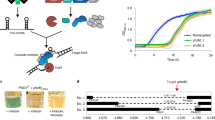
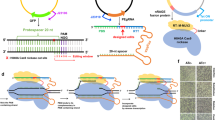
Here we use the clustered, regularly interspaced, short palindromic repeats (CRISPR)–associated Cas9 endonuclease complexed with dual-RNAs to introduce precise mutations in the genomes of Streptococcus pneumoniae and Escherichia coli. The approach relies on dual-RNA:Cas9-directed cleavage at the targeted genomic site to kill unmutated cells and circumvents the need for selectable markers or counter-selection systems. We reprogram dual-RNA:Cas9 specificity by changing the sequence of short CRISPR RNA (crRNA) to make single- and multinucleotide changes carried on editing templates. Simultaneous use of two crRNAs enables multiplex mutagenesis. In S. pneumoniae, nearly 100% of cells that were recovered using our approach contained the desired mutation, and in E. coli, 65% that were recovered contained the mutation, when the approach was used in combination with recombineering. We exhaustively analyze dual-RNA:Cas9 target requirements to define the range of targetable sequences and show strategies for editing sites that do not meet these requirements, suggesting the versatility of this technique for bacterial genome engineering.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Urnov, F.D., Rebar, E.J., Holmes, M.C., Zhang, H.S. & Gregory, P.D. Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 11, 636–646 (2010).
Bogdanove, A.J. & Voytas, D.F. TAL effectors: customizable proteins for DNA targeting. Science 333, 1843–1846 (2011).
Stoddard, B.L. Homing endonuclease structure and function. Q. Rev. Biophys. 38, 49–95 (2005).
Bae, T. & Schneewind, O. Allelic replacement in Staphylococcus aureus with inducible counter-selection. Plasmid 55, 58–63 (2006).
Sung, C.K., Li, H., Claverys, J.P. & Morrison, D.A. An rpsL cassette, janus, for gene replacement through negative selection in Streptococcus pneumoniae. Appl. Environ. Microbiol. 67, 5190–5196 (2001).
Sharan, S.K., Thomason, L.C., Kuznetsov, S.G. & Court, D.L. Recombineering: a homologous recombination-based method of genetic engineering. Nat. Protoc. 4, 206–223 (2009).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Deveau, H., Garneau, J.E. & Moineau, S. CRISPR-Cas system and its role in phage-bacteria interactions. Annu. Rev. Microbiol. 64, 475–493 (2010).
Horvath, P. & Barrangou, R. CRISPR-Cas, the immune system of bacteria and archaea. Science 327, 167–170 (2010).
Terns, M.P. & Terns, R.M. CRISPR-based adaptive immune systems. Curr. Opin. Microbiol. 14, 321–327 (2011).
van der Oost, J., Jore, M.M., Westra, E.R., Lundgren, M. & Brouns, S.J. CRISPR-based adaptive and heritable immunity in prokaryotes. Trends Biochem. Sci. 34, 401–407 (2009).
Brouns, S.J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964 (2008).
Carte, J., Wang, R., Li, H., Terns, R.M. & Terns, M.P. Cas6 is an endoribonuclease that generates guide RNAs for invader defense in prokaryotes. Genes Dev. 22, 3489–3496 (2008).
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011).
Hatoum-Aslan, A., Maniv, I. & Marraffini, L.A. Mature clustered, regularly interspaced, short palindromic repeats RNA (crRNA) length is measured by a ruler mechanism anchored at the precursor processing site. Proc. Natl. Acad. Sci. USA 108, 21218–21222 (2011).
Haurwitz, R.E., Jinek, M., Wiedenheft, B., Zhou, K. & Doudna, J.A. Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 329, 1355–1358 (2010).
Deveau, H. et al. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J. Bacteriol. 190, 1390–1400 (2008).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl. Acad. Sci. USA 109, E2579–E2586 (2012).
Makarova, K.S., Aravind, L., Wolf, Y.I. & Koonin, E.V. Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR-Cas systems. Biol. Direct 6, 38 (2011).
Barrangou, R. RNA-mediated programmable DNA cleavage. Nat. Biotechnol. 30, 836–838 (2012).
Brouns, S.J. Molecular biology. A Swiss army knife of immunity. Science 337, 808–809 (2012).
Carroll, D. A CRISPR approach to gene targeting. Mol. Ther. 20, 1658–1660 (2012).
Bikard, D., Hatoum-Aslan, A., Mucida, D. & Marraffini, L.A. CRISPR interference can prevent natural transformation and virulence acquisition during in vivo bacterial infection. Cell Host Microbe 12, 177–186 (2012).
Sapranauskas, R. et al. The Streptococcus thermophilus CRISPR-Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 39, 9275–9282 (2011).
Semenova, E. et al. Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc. Natl. Acad. Sci. USA 108, 10098–10103 (2011).
Wiedenheft, B. et al. RNA-guided complex from a bacterial immune system enhances target recognition through seed sequence interactions. Proc. Natl. Acad. Sci. USA 108, 10092–10097 (2011).
Zahner, D. & Hakenbeck, R. The Streptococcus pneumoniae beta-galactosidase is a surface protein. J. Bacteriol. 182, 5919–5921 (2000).
Marraffini, L.A., Dedent, A.C. & Schneewind, O. Sortases and the art of anchoring proteins to the envelopes of gram-positive bacteria. Microbiol. Mol. Biol. Rev. 70, 192–221 (2006).
Motamedi, M.R., Szigety, S.K. & Rosenberg, S.M. Double-strand-break repair recombination in Escherichia coli: physical evidence for a DNA replication mechanism in vivo. Genes Dev. 13, 2889–2903 (1999).
Hosaka, T. et al. The novel mutation K87E in ribosomal protein S12 enhances protein synthesis activity during the late growth phase in Escherichia coli. Mol. Genet. Genomics 271, 317–324 (2004).
Costantino, N. & Court, D.L. Enhanced levels of lambda Red-mediated recombinants in mismatch repair mutants. Proc. Natl. Acad. Sci. USA 100, 15748–15753 (2003).
Edgar, R. & Qimron, U. The Escherichia coli CRISPR system protects from lambda lysogenization, lysogens, and prophage induction. J. Bacteriol. 192, 6291–6294 (2010).
Marraffini, L.A. & Sontheimer, E.J. Self versus non-self discrimination during CRISPR RNA-directed immunity. Nature 463, 568–571 (2010).
Fischer, S. et al. An archaeal immune system can detect multiple Protospacer Adjacent Motifs (PAMs) to target invader DNA. J. Biol. Chem. 287, 33351–33363 (2012).
Gudbergsdottir, S. et al. Dynamic properties of the Sulfolobus CRISPR-Cas and CRISPR/Cmr systems when challenged with vector-borne viral and plasmid genes and protospacers. Mol. Microbiol. 79, 35–49 (2011).
Wang, H.H. et al. Genome-scale promoter engineering by coselection MAGE. Nat. Methods 9, 591–593 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR-Cas systems. Science doi:10.1126/science.1231143 (3 January 2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science doi:10.1126/science.1232033 (3 January 2013).
Cho, S.W., Kim, S., Kim, J.M. & Kim, J.-S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. advance online publication, doi:10.1038/nbt.2507 (29 January 2013).
Hwang, W.Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. advance online publication, doi:10.1038/nbt.2501 (29 January 2013).
Hoskins, J. et al. Genome of the bacterium Streptococcus pneumoniae strain R6. J. Bacteriol. 183, 5709–5717 (2001).
Havarstein, L.S., Coomaraswamy, G. & Morrison, D.A. An unmodified heptadecapeptide pheromone induces competence for genetic transformation in Streptococcus pneumoniae. Proc. Natl. Acad. Sci. USA 92, 11140–11144 (1995).
Horinouchi, S. & Weisblum, B. Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol resistance. J. Bacteriol. 150, 815–825 (1982).
Horton, R.M. In vitro recombination and mutagenesis of DNA: SOEing Together tailor-made genes. Methods Mol. Biol. 15, 251–261 (1993).
Podbielski, A., Spellerberg, B., Woischnik, M., Pohl, B. & Lutticken, R. Novel series of plasmid vectors for gene inactivation and expression analysis in group A streptococci (GAS). Gene 177, 137–147 (1996).
Husmann, L.K., Scott, J.R., Lindahl, G. & Stenberg, L. Expression of the Arp protein, a member of the M protein family, is not sufficient to inhibit phagocytosis of Streptococcus pyogenes. Infect. Immun. 63, 345–348 (1995).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Acknowledgements
We thank V. Fischetti and C. Euler for plasmid pLZ12spec, D. Court for the HME63 strain, J. Kern for plasmid pKD46, the Rockefeller University Genomic Resource Center for technical support and J. Roberts for the MG1655 strain. D.B. is supported by a Harvey L. Karp Discovery Award and the Bettencourt Schuller Foundation. D.C. is supported by the Medical Scientist Training Program. F.Z. is supported by a US National Institutes of Health (NIH) Director's Pioneer Award (DP1MH100706), Transformative R01, the Keck, McKnight, Gates, Damon Runyon, Searle Scholars, Klingenstein, and Simons Foundations, Bob Metcalfe, Mike Boylan and Jane Pauley. L.A.M. is supported by the Searle Scholars Program, the Rita Allen Scholars Program, a Irma T. Hirschl Award and a NIH Director's New Innovator Award (1DP2AI104556-01).
Author information
Authors and Affiliations
Contributions
W.J., D.B. and L.A.M. designed the experiments; W.J., D.B. and D.C. performed experiments; W.J., D.B., F.Z. and L.A.M. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Discussion, Supplementary Figures 1–11 and Supplementary Tables 1–3 (PDF 2790 kb)
Supplementary Data
Supplementary Data (XLSX 36 kb)
Rights and permissions
About this article
Cite this article
Jiang, W., Bikard, D., Cox, D. et al. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol 31, 233–239 (2013). https://doi.org/10.1038/nbt.2508
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2508
This article is cited by
-
Stable expression of HIV-1 MPER extended epitope on the surface of the recombinant probiotic bacteria Escherichia Coli Nissle 1917 using CRISPR/Cas9
Microbial Cell Factories (2024)
-
Massively parallel profiling of RNA-targeting CRISPR-Cas13d
Nature Communications (2024)
-
BacPE: a versatile prime-editing platform in bacteria by inhibiting DNA exonucleases
Nature Communications (2024)
-
Bacterial genome engineering using CRISPR-associated transposases
Nature Protocols (2024)
-
Use of Cas9 Targeting and Red Recombination for Designer Phage Engineering
Journal of Microbiology (2024)