Abstract
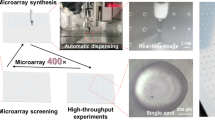
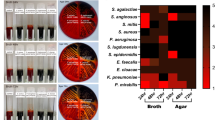
Bacterial attachment and subsequent biofilm formation pose key challenges to the optimal performance of medical devices. In this study, we determined the attachment of selected bacterial species to hundreds of polymeric materials in a high-throughput microarray format. Using this method, we identified a group of structurally related materials comprising ester and cyclic hydrocarbon moieties that substantially reduced the attachment of pathogenic bacteria (Pseudomonas aeruginosa, Staphylococcus aureus and Escherichia coli). Coating silicone with these 'hit' materials achieved up to a 30-fold (96.7%) reduction in the surface area covered by bacteria compared with a commercial silver hydrogel coating in vitro, and the same material coatings were effective at reducing bacterial attachment in vivo in a mouse implant infection model. These polymers represent a class of materials that reduce the attachment of bacteria that could not have been predicted to have this property from the current understanding of bacteria-surface interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
09 May 2014
In the version of this article initially published, the label of the 6th sample across in Figure 5 should have read 4(100%), not B(100%). The error has been corrected in the HTML and PDF versions of the article.
References
Davies, D. Understanding biofilm resistance to antibacterial agents. Nat. Rev. Drug Discov. 2, 114–122 (2003).
Costerton, J.W., Stewart, P.S. & Greenberg, E.P. Bacterial biofilms: a common cause of persistent infections. Science 284, 1318–1322 (1999).
Smith, A.W. Biofilms and antibiotic therapy: is there a role for combating bacterial resistance by the use of novel drug delivery systems? Adv. Drug Deliv. Rev. 57, 1539–1550 (2005).
Danese, P.N. Antibiofilm approaches: prevention of catheter colonization. Chem. Biol. 9, 873–880 (2002).
Krol, J.E. et al. Increased transfer of a multidrug resistance plasmid in Escherichia coli biofilms at the air-liquid interface. Appl. Environ. Microbiol. 77, 5079–5088 (2011).
Darouiche, R.O. et al. A comparison of two antimicrobial-impregnated central venous catheters. N. Engl. J. Med. 340, 1–8 (1999).
Raad, I. et al. Central venous catheters coated with minocycline and rifampin for the prevention of catheter-related colonization and bloodstream infections. A randomized, double-blind trial. Ann. Intern. Med. 127, 267–274 (1997).
Yorganci, K., Krepel, C., Weigelt, J.A. & Edmiston, C.E. Activity of antibacterial impregnated central venous catheters against Klebsiella pneumoniae. Intensive Care Med. 28, 438–442 (2002).
Jaeger, K. et al. Efficacy of a benzalkonium chloride-impregnated central venous catheter to prevent catheter-associated infection in cancer patients. Chemotherapy 47, 50–55 (2001).
Greenfeld, J.I. et al. Decreased bacterial adherence and biofilm formation on chlorhexidine and silver sulfadiazine-impregnated central venous catheters implanted in swine. Crit. Care Med. 23, 894–900 (1995).
Guay, D.R. An update on the role of nitrofurans in the management of urinary tract infections. Drugs 61, 353–364 (2001).
Caillier, L. et al. Synthesis and antimicrobial properties of polymerizable quaternary ammoniums. Eur. J. Med. Chem. 44, 3201–3208 (2009).
Darouiche, R.O., Mansouri, M.D., Gawande, P.V. & Madhyastha, S. Efficacy of combination of chlorhexidine and protamine sulphate against device-associated pathogens. J. Antimicrob. Chemother. 61, 651–657 (2008).
Li, P. et al. A polycationic antimicrobial and biocompatible hydrogel with microbe membrane suctioning ability. Nat. Mater. 10, 149–156 (2011).
Costa, F., Carvalho, I.F., Montelaro, R.C., Gomes, P. & Martins, M.C.L. Covalent immobilization of antimicrobial peptides (AMPs) onto biomaterial surfaces. Acta Biomater. 7, 1431–1440 (2011).
Monds, R.D. & O'Toole, G.A. The developmental model of microbial biofilms: ten years of a paradigm up for review. Trends Microbiol. 17, 73–87 (2009).
Holmes, P.F. et al. Surface-modified nanoparticles as a new, versatile, and mechanically robust nonadhesive coating: suppression of protein adsorption and bacterial adhesion. J. Biomed. Mater. Res. A 91A, 824–833 (2009).
Cheng, G. et al. Zwitterionic carboxybetaine polymer surfaces and their resistance to long-term biofilm formation. Biomaterials 30, 5234–5240 (2009).
Cheng, G., Zhang, Z., Chen, S.F., Bryers, J.D. & Jiang, S.Y. Inhibition of bacterial adhesion and biofilm formation on zwitterionic surfaces. Biomaterials 28, 4192–4199 (2007).
Hook, A.L. et al. High-throughput methods applied in biomaterial development and discovery. Biomaterials 31, 187–198 (2010).
Pernagallo, S., Wu, M., Gallagher, M.P. & Bradley, M. Colonising new frontiers-microarrays reveal biofilm modulating polymers. J. Mater. Chem. 21, 96–101 (2011).
Anderson, D.G., Levenberg, S. & Langer, R. Nanoliter-scale synthesis of arrayed biomaterials and application to human embryonic stem cells. Nat. Biotechnol. 22, 863–866 (2004).
Mei, Y. et al. Mapping the interactions among biomaterials, adsorbed proteins, and human embryonic stem cells. Adv. Mater. 21, 2781–2786 (2009).
Mei, Y. et al. Combinatorial development of biomaterials for clonal growth of human pluripotent stem cells. Nat. Mater. 9, 768–778 (2010).
Yang, J. et al. Polymer surface functionalities that control human embryoid body cell adhesion revealed by high-throughput surface characterization of combinatorial material microarrays. Biomaterials 31, 8827–8838 (2010).
Berger, H., Hacker, J., Juarez, A., Hughes, C. & Goebel, W. Cloning of the chromosomal determinants encoding hemolysin production and mannose-resistant hemagglutination in Escherichia coli. J. Bacteriol. 152, 1241–1247 (1982).
Hidron, A.I. et al. Antimicrobial-resistant pathogens associated with healthcare-associated infections: annual summary of data reported to the national healthcare safety network at the Centers for Disease Control and Prevention, 2006–2007. Infect. Control Hosp. Epidemiol. 29, 996–1011 (2008).
Schumm, K. & Lam, T.B. Types of urethral catheters for management of short-term voiding problems in hospitalised adults. Cochrane DB. Syst. Rev. 16, CD004013 (2008).
Katsikogianni, M. & Missirlis, Y.F. Concise review of mechanisms of bacterial adhesion to biomaterials and of techniques used in estimating bacteria-material interactions. Eur. Cell. Mater. 8, 37–57 (2004).
O'Toole, G., Kaplan, H.B. & Kolter, R. Biofilm formation as microbial development. Annu. Rev. Microbiol. 54, 49–79 (2000).
Flemming, H.C. & Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 8, 623–633 (2010).
Urquhart, A.J. et al. High-throughput surface characterisation of a combinatorial material library. Adv. Mater. (Deerfield Beach Fla.) 19, 2486–2491 (2007).
Davies, M.C. et al. High-throughput surface characterization: a review of a new tool for screening prospective biomedical material arrays. J. Drug Target. 18, 741–751 (2010).
Hook, A.L. et al. Polymers with hydro-responsive topography identified using high-throughput AFM of an acrylate microarray. Soft Matter 7, 7194–7197 (2011).
Taylor, M. et al. Partial least squares regression as a powerful tool for investigating large combinatorial polymer libraries. Surf. Interface Anal. 41, 127–135 (2009).
Urquhart, A.J. et al. TOF-SIMS analysis of a 576 micropatterned copolymer array to reveal surface moieties that control wettability. Anal. Chem. 80, 135–142 (2008).
Dalton, H.M. & March, P.E. Molecular genetics of bacterial adhesion and biofouling. in Handbook of Bacterial Adhesion: Principles, Methods, and Applications (eds. An, Y.H. & Friedman, R.J.) 43–51 (Humana Press, 2000).
Ong, Y.L., Razatos, A., Georgiou, G. & Sharma, M.M. Adhesion forces between E. coli bacteria and biomaterial surfaces. Langmuir 15, 2719–2725 (1999).
Conrad, J.C.C.J.C. et al. Flagella and pili-mediated near-surface single-cell motility mechanisms in P. aeruginosa. Biophys. J. 100, 1608–1616 (2011).
Williams, P., Winzer, K., Chan, W.C. & Camara, M. Look who's talking: communication and quorum sensing in the bacterial world. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 362, 1119–1134 (2007).
Ferrieres, L. & Clarke, D.J. The RcsC sensor kinase is required for normal biofilm formation in Escherichia coli K-12 and controls the expression of a regulon in response to growth on a solid surface. Mol. Microbiol. 50, 1665–1682 (2003).
Gilbert, K.B., Kim, T.H., Gupta, R., Greenberg, E.P. & Schuster, M. Global position analysis of the Pseudomonas aeruginosa quorum-sensing transcription factor LasR. Mol. Microbiol. 73, 1072–1085 (2009).
Harder, P., Grunze, M., Dahint, R., Whitesides, G.M. & Laibinis, P.E. Molecular conformation in oligo(ethylene glycol)-terminated self-assembled monolayers on gold and silver surfaces determines their ability to resist protein adsorption. J. Phys. Chem. B. 102, 426–436 (1998).
Taylor, M., Urquhart, A.J., Zelzer, M., Davies, M.C. & Alexander, M.R. Picoliter water contact angle measurement on polymers. Langmuir 23, 6875–6878 (2007).
Messina, M. Gene Regulation in Pseudomonas Aeruginosa: from Environmental Signals to Responses via Global Post-Transcriptional Control and Intracellular Messaging. PhD thesis, Univ. Nottingham (2010).
Qazi, S.N.A., Rees, C.E.D., Mellits, K.H. & Hill, P.J. Development of GFP vectors for expression in Listeria monocytogenes and other low G+C gram positive bacteria. Microb. Ecol. 41, 301–309 (2001).
Brooks, T. & Keevil, C.W. A simple artificial urine for the growth of urinary pathogens. Lett. Appl. Microbiol. 24, 203–206 (1997).
Diggle, S.P. et al. The galactophilic lectin, LecA, contributes to biofilm development in Pseudomonas aeruginosa. Environ. Microbiol. 8, 1095–1104 (2006).
Johansson, E.M.V. et al. Inhibition and dispersion of Pseudomonas aeruginosa biofilms by glycopeptide dendrimers targeting the fucose-specific lectin LecB. Chem. Biol. 15, 1249–1257 (2008).
Kuklin, N.A. et al. Real-time monitoring of bacterial infection in vivo: Development of bioluminescent staphylococcal foreign-body and deep-thigh-wound mouse infection models. Antimicrob. Agents Chemother. 47, 2740–2748 (2003).
Acknowledgements
Funding from the Wellcome Trust (grant no. 085245 and support from N. Shepherd) and the Medical Research Council UK (for the in vivo work; grant no. G0802525) is gratefully acknowledged. M. Alexander gratefully acknowledges the Royal Society for the provision of his Wolfson Research Merit Award. Assistance with ToF-SIMS measurements from D. Scurr is kindly acknowledged. Assistance with the preparation of polymer for in vivo studies by E. Eaves, N. Nguyen and J. Li is kindly acknowledged.
Author information
Authors and Affiliations
Contributions
M.R.A., M.C.D., P.W., R.L. and D.G.A. conceived the high-throughput strategy. Y.M., A.L.H. and J.Y. prepared the microarrays. J.Y. coated catheters with hit polymers. J.Y. and A.L.H. performed the HT-SC. C.-Y.C. performed the biological assays with support from S.A. D.J.I. prepared polymer for in vivo assays. J.L. and A.C. performed the in vivo assays. A.L.H., C.-Y.C., J.Y., M.R.A., M.C.D., P.W., R.B., D.G.A. and R.L. contributed to the analysis of data and the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Discussion, Supplementary Table 1 and Supplementary Figures 1–17 (PDF 1455 kb)
Rights and permissions
About this article
Cite this article
Hook, A., Chang, CY., Yang, J. et al. Combinatorial discovery of polymers resistant to bacterial attachment. Nat Biotechnol 30, 868–875 (2012). https://doi.org/10.1038/nbt.2316
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2316
This article is cited by
-
Fusion and classification algorithm of octacalcium phosphate production based on XRD and FTIR data
Scientific Reports (2024)
-
An injectable and biodegradable zwitterionic gel for extending the longevity and performance of insulin infusion catheters
Nature Biomedical Engineering (2023)
-
Bacterial capsular polysaccharides with antibiofilm activity share common biophysical and electrokinetic properties
Nature Communications (2023)
-
Uniform amphiphilic cellulose nanocrystal films
Polymer Journal (2022)
-
Evaluation of the relative potential for contact and doffing transmission of SARS-CoV-2 by a range of personal protective equipment materials
Scientific Reports (2022)