Abstract
We generated a high-quality reference genome sequence for foxtail millet (Setaria italica). The ∼400-Mb assembly covers ∼80% of the genome and >95% of the gene space. The assembly was anchored to a 992-locus genetic map and was annotated by comparison with >1.3 million expressed sequence tag reads. We produced more than 580 million RNA-Seq reads to facilitate expression analyses. We also sequenced Setaria viridis, the ancestral wild relative of S. italica, and identified regions of differential single-nucleotide polymorphism density, distribution of transposable elements, small RNA content, chromosomal rearrangement and segregation distortion. The genus Setaria includes natural and cultivated species that demonstrate a wide capacity for adaptation. The genetic basis of this adaptation was investigated by comparing five sequenced grass genomes. We also used the diploid Setaria genome to evaluate the ongoing genome assembly of a related polyploid, switchgrass (Panicum virgatum).
Similar content being viewed by others
Main
Grasses of the genus Setaria occur in natural and agricultural ecosystems worldwide, from the tropical forage S. sphacelata (golden timothygrass) to the invasive S. viridis (green foxtail) and S. verticillata (hooked bristlegrass) populations that can be found in the farthest northern reaches of Canada. Five Setaria species, S. faberii (giant foxtail), S. viridis, S. pumila (yellow foxtail), S. geniculata (knotroot foxtail) and S. verticillata, are problematic weeds1. S. viridis is among the most widespread plant species on the planet, and is problematic for crop production due to its repeated evolution of herbicide resistance2.
In Northern China, around 6,000 BC, foxtail millet was domesticated from S. viridis, and together with proso millet (Panicum miliaceum), became a major cereal crop3. Foxtail millet remains a significant crop and dietary staple in this region. It has been suggested, but not proven, that foxtail millet was independently domesticated in Europe a few millennia later4,5,6,7,8. Although the flower structure of S. italica creates a strong bias for inbreeding, the great diversity in S. italica morphology and adaptation suggests that gene flow between S. italica and S. viridis may continue when these two species are grown in sympatry9,10. More recently, directed crosses between S. viridis and S. italica have been used for genetic mapping11 and to introduce herbicide resistance from S. viridis into S. italica breeding germplasm12,13.
Recent interest in the S. italica–S. viridis pair (hereafter referred to as “Setaria”) has centered on their potential as model species for understanding basic biological processes14,15. Some accessions of S. viridis have short cycling times (6 weeks, seed to seed) and tiny statures (20 cm at maturity), and can be efficiently transformed using Agrobacterium-based methods. Setaria is also a diploid, tractable model for polyploid biofuel crops like switchgrass (Panicum virgatum) and Napier grass (Pennisetum purpureum).
Multiple mapping populations, fosmid libraries and mutagenized populations have been developed for the genus Setaria (some for S. italica, others for S. viridis). Here, the Setaria toolkit is expanded with a high-quality, well-annotated reference genome sequence based on Sanger shotgun sequencing, a high-resolution genetic map, an end-sequenced bacterial artificial chromosome (BAC) library, expressed sequence tags (ESTs) from 13 cDNA libraries and 4 RNA-Seq libraries. We compared these sequences with the genomes of S. viridis, switchgrass, sorghum and rice to uncover both shared and unique properties of the independent adaptations of several grasses to life on earth. Detailed comparisons of the diploid S. italica with available polyploid switchgrass genome sequences demonstrated the utility of using a closely related diploid genome sequence to aid assembly of a polyploid plant genome.
Results
Setaria italica phylogeny
Setaria, a member of the grass subfamily Panicoideae, tribe Paniceae is closely related to other Paniceae, including switchgrass, proso millet and pearl millet16,17,18,19 (Fig. 1). Paniceae is closely related to Andropogoneae, which includes maize and sorghum. Most members of Paniceae share a base chromosome number of x = 9, whereas Andropogoneae have x = 10 (ref. 19). Setaria last shared a common ancestor with pearl millet ∼8.3 million years ago (Myr ago), and with Panicum (switchgrass and proso millet) ∼13.1 Myr ago20. This is more recent than its common ancestor with maize and sorghum (∼26 Myr ago), or with rice and Brachypodium (∼52 Myr ago). Setaria is morphologically and developmentally similar to pearl millet21 and can serve as a model for that important dry-land crop. Like maize and sorghum, Setaria uses C4 photosynthesis, but has acquired the C4 pathway independently (Fig. 1)20. Setaria also provides a tractable diploid model for the tetraploids proso millet and switchgrass. A phylogenetic analysis of five nuclear genes (Fig. 1) shows that the proso millet and switchgrass lineages underwent independent polyploidization events, both of which occurred after the divergence of Panicum from Setaria.
Left panel, relationships of the commelinid monocots, showing the order Poales relative to the next closest order with a genome sequence, Arecales (http://www.mobot.org/MOBOT/research/APweb/). Middle panel, relationships among some grass genera (GPWG 2001). Right panel, phylogeny of selected Panicum, Setaria and Pennisetum species. Green, C4 lineage.
Genome sequence
A BAC library (50,688 clones, average insert size of ∼121 kb, ∼12× genomic coverage) was prepared in the vector pIndigoBac536 from total genomic DNA of S. italica inbred Yugu1. BACs were subjected to BAC end sequence (BES) analysis (Supplementary Table 1). With Yugu1 mRNA isolated from a variety of different tissues, at different times in development and exposed to different stress treatments, we generated 13 EST libraries (Supplementary Table 2). A total of 63,286 ESTs were sequenced on the Sanger ABI3730xl platform and another 1,217,156 on the 454 FLX platform. We generated ∼784-million RNA-Seq reads from four developing leaf tissues using the Illumina Genome Analyzer II platform, of which ∼580 million were of appropriate quality to be used for expression analysis (Supplementary Table 2). The distributions of the RNA-Seq reads across the full Setaria genome assembly are available from the Setaria browser at http://www.phytozome.net/.
For full genome analysis, nuclear DNA was prepared from Yugu1 seedlings and subjected to Sanger sequence analysis on ABI3730xl capillary sequencing machines. Separate libraries with several insert sizes (3 kb, 6 kb, 37 kb, 121 kb) were end-sequenced using standard Sanger-based methods. A total of 5,736,559 reads were generated, yielding 4,003,684,795 bp of data with an acceptable PHRED score of 20 (Supplementary Table 3). Assembly of these raw sequencing results, including the BES data, was performed using a modified Arachne v.20071016 software. Assembled sequences were ordered using both the BES data and homology to the sequenced sorghum genome22. These Setaria assemblies were released to the general public in January 2010 as version 1.0.
Nuclear DNA prepared from the tissue of a pool of seedlings derived from the seed of a single plant of S. viridis accession A10 was resequenced using the Illumina Genome Analyzer II platform. The ∼3,500 Mb of generated data provided ∼7× coverage. Apparent orthologs of several candidate domestication genes that were important in other cereal crops (e.g., Q, qsh1, SH4 and tb1)23 were compared between the Yugu1 sequence from domesticated S. italica and the A10 sequence from the undomesticated S. viridis, and no large differences were observed, suggesting either that foxtail millet domestication involved a different set of loci or that the genetic changes (e.g., missense or regulatory mutations) were too subtle to be detected through simple sequence scrutiny.
A genetic map for Setaria
A cross between S. italica inbred B100 and S. viridis accession A10 was performed in 1997, and an F2 generation of this cross was used to generate the first genetic map of Setaria italica/viridis11,24. We used 247 progeny of this cross to construct a recombinant inbred line (RIL) population through eight generations of single-seed descent. This population was mapped with 992 single-nucleotide polymorphism (SNP) markers (Supplementary Note 1) distributed at ∼400-kb intervals across the version 1.0 assembly. The 992 mapped SNPs were distributed over 73 scaffolds, leaving 6.7 Mb of DNA and four scaffolds >50 kb that were not associated with a mapped marker. This yielded a genetic map with nine linkage groups, as expected for the nine chromosome pairs in these two Setaria species. The map covers a total of 1,416 centiMorgans (cM) on chromosomes ranging from 124 cM to 201 cM in size (Supplementary Fig. 1 and Supplementary Table 4).
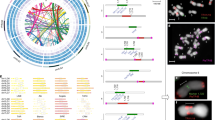
The genetic map indicated several regions of major segregation distortion on seven of the nine Setaria chromosomes. On chromosome II, as many as 95% of the alleles were of the cultivated type (Fig. 2, Supplementary Fig. 2 and Supplementary Table 4). Alleles from the cultivated S. italica parent were mainly on chromosomes II, III, IV, V and IX, whereas alleles from the wild S. viridis parent were overrepresented on chromosome VI. On chromosome VII, alleles from the cultivated parent were overrepresented on the short arm and the proximal region of the long arm, whereas alleles from the wild parent were overrepresented on the long arm. Segregation distortion is common in wide crosses in the angiosperms, especially interspecies crosses25,26, and may reflect reproductive barriers or inadvertent selection during the inbreeding process. Previous studies have shown that segregation distortion does not affect marker order determination27,28, as confirmed in this study by the excellent concordance between the genetic map and the sequence assembly, including in regions where segregation distortion was observed. Large genomic rearrangements (e.g., inversions or translocations) that might produce hybrid fertility issues were not observed in the cross between S. italica and S. viridis, although the comparison of the Yugu1 assembly to the Setaria genetic map did reveal a Yugu1-specific inversion near the centromere on chromosome IX.
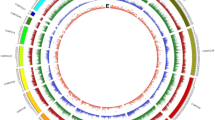
Copy numbers for each track were calculated in 500-kb sliding windows, incrementing every 100 kb. Scale (blue, minimum abundance; red, maximum abundance). Black triangles indicate the estimated position of the centromere on each chromosome. “Other class I TEs” are LINEs, short interspersed nuclear elements (SINEs) and unclassified LTR retrotransposons. “Other class II TEs” are Helitrons, Mutators, hATs, Tc1/Mariners and PIF/Harbingers. Segregation distortion is represented as log10 (A:B ratio). Green indicates no distortion, increasing red intensity indicates significant overrepresentation of S. italica alleles and increasing blue intensity indicates significant overrepresentation of S. viridis alleles. TE, transposable element.
Final genome assembly
The final genomic sequence assembly contains 396.7 Mb of sequence in nine chromosomes and an additional 4.2 Mb in 327 scaffolds (mostly <50 kb in size) that are unanchored by the genetic map, with an estimated genome coverage of ∼80% of the total nuclear DNA (based on a predicted genome size of ∼510 Mb (http://data.kew.org/cvalues/)). The completed assembly was compared against 42 randomly selected, fully sequenced BAC clones derived from the S. italica BAC library and against nine randomly selected, fully sequenced fosmid clones containing DNA from S. viridis accession A10. Of the 51 regions investigated, all were represented and all genes were colinear with the whole genome shotgun (WGS) sequencing assembly. The finished Yugu1 BAC clones were 98.7% identical with the Yugu1 WGS sequence. In addition to a small portion of the unaligned bps (0.29%) that were caused by gaps in the genome assembly, five of the 42 BACs exhibited a major discrepancy when compared to the assembly, all of which are repetitive DNA collapses of 5–10 kb in comparison to the clone sequence. The majority of the remaining differences may be due to instances of retained heterozygosity in the Yugu1 accession or to differences between the Yugu1 seed stocks used as DNA sources for the BAC library and WGS sequencing.
Setaria genome annotation and analysis
Transposable element and gene (as represented by exons) distributions across two chromosomes are shown in Figure 2. At least 40% of the Yugu1 genome is composed of transposable elements, a number that is consistent with the low abundances found in small grass genomes like rice (∼40%) and sorghum (∼62%) and much lower than the >80% seen in genomes like maize and wheat22,29,30. As usually observed in plant genomes, the long terminal repeat (LTR) retrotransposons are the most abundant class, comprising >25% of the total nuclear genome. Exons for protein-encoding genes are predicted to comprise ∼46 Mb, or ∼9%, of the genome. Centromere locations were inferred from regions that were the lowest in the ratio of cM to Mb and centered on the regions with the highest ratio of transposable elements to exons (Fig. 2 and Supplementary Fig. 2, black arrows).
In common with the maize genome30 each class of transposable elements in Setaria exhibits a different bias in accumulation across the chromosomes. LTR retrotransposons, especially the gypsy superfamily that comprises most of the repetitive DNA in the genome, are enriched in the pericentromeric heterochromatin (Fig. 2). Other element classes, such as long interspersed nuclear elements (LINEs) and most DNA elements, are rare in the pericentromeric regions. A notable exception is the CACTA family of DNA elements that, as in maize30, is found at fairly even distribution across all genomic regions. Using the standard LTR retrotransposon dating approach31 and degree of element homology for other transposable element classes, insertion dates were approximated for each element family (Supplementary Fig. 3). The data indicate a very recent burst of LTR retrotransposon amplification, peaking within the last few hundred thousand years, whereas the LINE and CACTA elements exhibit a broader range of activity over the last 6 million years. Helitron elements seem to have undergone two major bursts of activity, one ∼1.8 Myr ago and a more dramatic event ∼4 Myr ago (Supplementary Fig. 3), both of which date long before the concurrent Helitron bursts ∼0.2 Myr ago observed in maize, rice and sorghum32.
Small RNAs (smRNAs) were analyzed by comparing ∼10.3 million smRNA reads from inflorescence and leaf (http://smallrna.udel.edu/project_data.php) of Setaria inbred Yugu1 with known microRNAs (miRNAs) (Supplementary Table 5) and with the annotated transposable element component of the Setaria genome (Supplementary Fig. 4). Forty-eight families of miRNA transcripts were found that were 100% identical in sequence to mature miRNAs from 14 investigated species, including several monocots, dicots and the moss Physcomitrella patens. Twenty-four nucleotide smRNAs with strong homology to transposable elements were particularly abundant. Of the smRNA sequences associated with the 48 conserved miRNA families, >93% were found at >95% identity in the Setaria genome assembly, indicating good recovery of this class of gene in the current genome sequence (data not shown).
The ratio of physical to genetic map distance is typically a few hundred kb/cM, but varies by three orders of magnitude across the genome, from less than 50 kb/cM in some distal chromosome regions to several tens of Mb/cM in the centromeric regions (Supplementary Fig. 5). Large regions of suppressed recombination were identified on nearly all chromosomes. The only exception was chromosome VIII, where the region of low recombination was less extensive compared to the other chromosomes. One possible explanation might be the presence of a substantial gap in the assembly in that region. In all cases, the lowest cM to Mb ratios were found in the same regions on a chromosome that had the greatest gypsy retrotransposon abundance (Fig. 2 and Supplementary Figs. 2 and 5). The predicted centromere locations indicated that chromosome VII is acrocentric whereas the other chromosomes are metacentric or submetacentric, which is the typical and likely ancestral structure of all cereal chromosomes33,34,35.
Full genome annotation yielded a predicted 24,000 to 29,000 protein-encoding genes (Online Methods and Supplementary Notes 2–4). Genes (median exon and intron sizes of 163 bp and 135 bp, respectively) and peptides (median length 329 amino acids) and number of exons per gene (average 4.5) were consistent with other grasses, as well as Arabidopsis, reflecting the high degree of gene structure conservation across angiosperms (Supplementary Table 6). The annotated Setaria genome includes 10,059 single-intron genes, compared to a predicted 11,112 and 6,217 in rice and sorghum, respectively.
We compared Setaria, sorghum and rice genomes, and found that they exhibit extensive synteny. Interestingly, although this investigation was only at the highest scale of chromosome structure, we observed that the sorghum genome exhibited fewer chromosomal rearrangements relative to rice than did the Setaria genome (Supplementary Note 5).
Comparison of Setaria with switchgrass
Using two different methods, including comparison of homologous gene sets and alignment of switchgrass ESTs to the Setaria genome sequence, we showed that the Setaria and switchgrass lineages diverged 3–7 Myr ago (Supplementary Note 6).
To investigate the relative structure and evolution of switchgrass and Setaria chromosomes, a switchgrass genetic map36 was aligned with the Setaria and sorghum genome sequences, allowing the identification of several translocations and inversions that differentiate these genomes (Supplementary Note 7). To compare the structure of the switchgrass genome to Setaria at the level of microcolinearity, we shotgun sequenced and fully assembled eight switchgrass fosmids (each containing at least two genes) using Sanger sequencing methods and Phred/Phrap programs. Using standard approaches to annotate the fosmid genes37, we identified 35 protein-encoding genes, all of which had orthologs in both sorghum and Setaria, and were present in the same order in both species. There were, however, minor differences in gene content in the homologous regions due to insertions in Setaria (the insertion of a single novel gene in one Setaria region) and sorghum (four unique, nonadjacent sorghum genes were found in the gene-dense region surrounding a galacturonosyltransferase 10 gene) (data not shown). This small sample set indicates that both sorghum and Setaria are good genome models for switchgrass.
The polyploid switchgrass genome is a challenge for WGS assembly, so we used BES data from switchgrass38 as the starting point for a comparison with the Setaria and sorghum genome assemblies. We aligned 351,152 switchgrass BES reads with the annotated genes from the most recent assemblies of sorghum and Setaria, using a cross-species exploration parameter set, to find switchgrass BACs that had an identified gene at both ends. The terminal genes of these BACs were then compared to the current genome assemblies for both sorghum22 and Setaria. Alignments were screened for quality (>75% base pair identity, E-value < 1.0e-20 and <500-kb separation of pairs), and duplicate alignments were removed before accumulating statistics. The switchgrass BES aligned ∼24% more frequently with Setaria than with sorghum (2,798 versus 2,254 respective colinearities, of which 928 were shared by both species) (Table 1). When colinearity was observed for both sorghum and Setaria, the colinear genes were found to be, on average, about 15% farther apart in sorghum compared with Setaria. Inversion of one of the terminal genes in the BES pair was found to be more common in Setaria than sorghum (Table 1), but this type of rearrangement requires preservation of regional gene content, which is higher in Setaria than in sorghum. Although further analysis will be needed to determine the precise level of colinearity between switchgrass and other plant genomes, these BES comparisons document superior microcolinearity with Setaria compared with sorghum.
Assessing switchgrass genome assembly
To separate the genomic content of the tetraploid reference switchgrass genotype (Alamo AP-13) into the correct subgenomes, one must take advantage of the divergence between the subgenomes without fracturing the contigs into chimeric allelic variants within the subgenomes caused by the heterozygosity of the sequenced variety. We used the Setaria sequence to evaluate the accuracy of ongoing switchgrass genomic sequence assembly.
By aligning the switchgrass WGS assembly against the Setaria reference, we found that it was possible to determine for each genic region whether the subgenomes were properly split, collapsed into a single reference, or fractured into three or four allelic variant contigs (Fig. 3). For the gene shown in Figure 3, contig 1 represents the 3′ portion of the gene, merging the subgenomes into a single contig, whereas contig 2 and contig 3 are split into subgenome components, as are contigs 4, 5 and 6. At 132 kb, all four alleles are present because the variation is sufficient to separate subgenomes into unique haplotypes. Additionally, contig 4 and contig 5 show a subgenome-specific insertion relative to Setaria. Future studies for each gene region, combined with data for individual contigs, will permit increasing stringency of assembly for collapsed regions in order to recover the subgenomes.
The upper line scales show positions in the Setaria assembly for a region encoding a ubiquitin ligase. The transcript for this gene, annotated in Setaria, is represented by exons (tan boxes) and introns (thin lines) on the 'transcript' row. The multicolored bar below the transcript shows the Setaria assembly, with the degree of homology to switchgrass indicated by the height of the color peaks within the bar. The multicolored bars below are the switchgrass contigs, which could now be assembled because of their microcolinearity with Setaria. Note that the four switchgrass haplotypes have become anywhere from one (3′ end) to four (tannish-green middle region) assemblies for this gene. Contigs 4 and 5 have a small subgenome-specific insertion (see white space in Setaria assembly).
Genetic basis of adaptation
The evolution of C4 photosynthesis occurred independently in the Setaria and sorghum lineages20,39 (Fig. 1). In C4 photosynthesis, carbon is shuttled as a C4 acid from the mesophyll to the bundle sheath cells to create a CO2 pump that greatly reduces photorespiratory losses associated with hot, arid environments40 (Supplementary Fig. 6). A comparison of amino acid identity in the primary carbon shuttle enzymes PEPC, PPDK and MDH among the sequenced grass genomes reveals the expected gradient of amino acid conservation (Supplementary Fig. 6). The highest conservation is observed between maize and sorghum orthologs and identity is higher within the panicoids than between the panicoids and the pooid grasses rice and Brachypodium. This is consistent with recent recruitment of these enzymes into the C4 pathway from ancestral C3 isoforms. From analysis of the Setaria genomes, it seems that the C4 isoform of malic enzyme was recruited from a different C3 paralog in Setaria than in maize and sorghum, because sequence identity is higher between the Andropogoneae and Pooideae isoforms than the S. italica isoform (Supplementary Fig. 6 and Supplementary Table 7). A more detailed analysis of sequence variation among these malic enzyme isoforms could reveal novel signatures of C4 evolution, including targets of positive selection such as those described for PEPC41.
To investigate the molecular basis of drought tolerance in grasses, we performed clustering of the protein sequences from four grass genomes and used them to identify six drought-associated gene clusters that have significantly more gene members in drought-tolerant species (Setaria and sorghum), as compared with drought-susceptible species (maize and rice) (Table 2).
Genes that control flowering time have major effects upon adaptation to local environments, making them important targets for breeding and biotechnological modification. Most temperate grasses such as Brachypodium, wheat and barley use the cold temperatures of winter (vernalization) to induce flowering in spring, but Setaria does not1. Instead, it appears to rely on photoperiod, with a conserved CONSTANS pathway and an additional EARLY HEADING DATE1 (Ehd1)–GRAIN HEADING DATE7 (Ghd7) pathway. Vernalization, photoperiod and autonomous pathways in both grasses and Arabidopsis are integrated by FLOWERING LOCUS T (FT), a mobile signal whose RNA or protein is thought to move between the leaves (site of signal reception) and the apical meristem (site of floral induction)42. FT belongs to the PEBP gene family, which is much expanded in grasses compared to Arabidopsis, with 19 copies in rice43 and 22 in Setaria (this study). There are two subgroups of genes in the grass clade most similar to FT, one of which contains the confirmed rice floral promoting genes HEADING DATE3a and RFT1 (ref. 44). This group also contains three Setaria homologs (Supplementary Fig. 7).
Discussion
The Setaria genome analysis yielded ∼24,000 protein-encoding genes by a manually validated sample sequence approach and ∼29,000 by a largely automated analysis of the WGS sequence assembly. The sample sequence approach used a conservative set of criteria for gene calling, thereby minimizing the common error where plant gene number prediction is 10–50% too high, because of transposable elements miscalled as genes in annotations that are done without extensive manual curation45,46. Hence, the most likely gene number for Setaria is 24,000–29,000, in line with gene complements of other diploid grasses like rice and sorghum.
We carried out a preliminary analysis of the switchgrass WGS assembly onto the Setaria diploid framework, and identified numerous cases where paralogous assemblies could be separated from homoeologous assemblies. This demonstrated the utility of a high-quality reference genome for assembly of an outbred polyploid species. We also investigated microcolinearity by comparing short sequenced segments and switchgrass BES data. The superior microcolinearity exhibited between switchgrass and Setaria supports the future power of Setaria to identify both common and novel genes and/or pathways in important Paniceae crops.
We investigated the properties of the Setaria genome, including its level of diversity, the transmission of that diversity, and its redistribution by recombinational and mutational processes. Transposable elements were very abundant, recently active and distributed nonrandomly. Hence, because of their ability to break chromosomes, to acquire and amplify genes or gene fragments, and to serve as sites of ectopic recombination47, transposable elements are likely candidates for participation in macro and micro rearrangements. Recombination was distributed nonrandomly, with cold spots in the regions flanking centromeres. Although the SNP analyses indicated a high level of diversity in Setaria, the cross we analyzed was observed to exhibit strong segregation distortion for some chromosomal regions. Hence, diversity will not be evenly transmitted and could be lost by a standard breeding approach. The molecular markers developed in this project will allow identification of diverse and underrepresented chromosomal haplotypes so that they can be retained and recombined for future allele mining.
One major biological question is how different species become unique organisms. To understand the origins of adaptation, we think it is particularly useful to investigate multiple species, especially when they have independently evolved an ability to prosper under similar environmental conditions. With the sequencing of the Setaria genome, evolutionary geneticists now have an annual, temperate, C4, drought- and cold-tolerant grass that they can comprehensively compare to other plants that have or have not evolved these adaptations. In the brief examples of adaptation analysis in this study, particular traits were targeted that are relevant for biotechnical improvement, namely drought tolerance, photosynthetic efficiency and flowering control.
With a completed genome sequence, the door is now open for further development of Setaria as a model plant. This model can be applied to understanding such phenomena as cell wall composition, growth rate, plant architecture and input demand that are pertinent to the development of biofuel crops. In addition to its use as a panicoid model for switchgrass, pearl millet, maize and Miscanthus, Setaria has the model characteristics that will encourage its development as a study system for any biological process, with pertinence to the entire plant kingdom and beyond.
Methods
Phylogenetic modeling.
Phylogenetic analysis of Panicum species used sequences of alcohol dehydrogenase1 (adh1), knotted1 (kn1), poly-A binding protein1 (pabp1), cellulase1 (PvCel1) and cellulase2 (PvCel2). Loci were amplified by PCR using exon-anchored primers, cloned and sequenced on both strands. Genome-specific sequences were easily identified by pairwise comparisons and by phylogenetic analysis; within P. virgatum, A and B genome sequences differed by about 7% in the introns. The number of sequence types recovered from each plant corresponded with genome size and reported ploidal level. Separate analyses were run for each locus under Bayesian inference (MrBayes 3.1.2)49 and parsimony (MP) (using PAUP* 4.0). For combined data sets, maximum likelihood analyses were also conducted using GARLI v0.95 (ref. 50).
Individual gene trees were not strongly incongruent, so the gene sequences were concatenated and analyzed as a single supermatrix. The tree presented in Figure 1 is pruned from a larger analysis that included 25 species of Panicum (data not shown).
Nuclear DNA isolation from Setaria.
A single, highly inbred Yugu1 plant was used for S. italica DNA preparation for the WGS project. Approximately 50 g of frozen tissue from whole seedlings were ground to a fine powder with a mortar and pestle and immediately suspended in a sucrose extraction buffer (SEB) following option Y as described51. The suspension was filtered through two layers of cheesecloth and miracloth to remove particulates, and 10% Triton X-100 (v/v) in SEB solution was then added to the filtered suspension at a 1:20 volume ratio. Nuclei were isolated from the suspension using a series of centrifugations at 650g for 15 min. Nuclei were then resuspended in 5 ml of buffer AP1 from the DNeasy plant maxi-kit (Qiagen, Valencia, CA), and manufacturer's instructions were followed to isolate plastid-free DNA.
Library production.
The switchgrass (variety Alamo) and S. viridis (accession A10) fosmid libraries were prepared from nuclear DNA preparations in the vector pCC2FOS (Epicentre Technologies, Madison, WI), as previously described37. The S. italica and P. virgatum BAC libraries were prepared by the Clemson University Genomics Institute in vector pIndigoBac536 (Supplementary Table 1). EST libraries were generated from a variety of tissues, treatments and developmental stages (see below) using the vector pDNRLib (Supplementary Table 2).
Tissues and treatments for EST analysis.
Yugu1 seed were germinated in sterile vermiculite and water under normal greenhouse conditions. To reduce growth of microorganisms, we amended the water with Captan (1 tbs/gal) and erythromycin (384 mg/gal). After germination, seedlings were grown for 3–5 weeks in sterile vermiculite with a nutrient solution (water amended with 20:10:20 fertilizer as per manufacturer's recommendation) before starting the treatment regime. After treatment, tissues were harvested and immediately placed in liquid nitrogen. RNA was then extracted using the Omega Biotek Plant Maxi Kit (Norcross, GA). The RNA was DNase treated and quality was assessed on a Bioanalyzer 2100 from Agilent Technologies (Santa Clara, CA). RNAs from a total of 13 different tissues and/or treatments were isolated for the generation of ESTs. These treatments included 8 whole seedling treatments as follows: (i) 20% 6000 polyethylene glycol w/v in nutrient solution for 24 h, (ii) cold treatment at 4° for 2.5 h, (iii) heat shock treatment at 45° for 1 h, (iv) salicylic acid (SA) treatment where [1 mM] SA is thoroughly misted on the plants that were harvested 24 h later, (v) salt stress treatment where the NaCl concentration of the nutrient solution is increased to [250 mM], (vi) full light for 48 h, (vii) no light for 48 h and (viii) no treatment. Additionally, three floral spike treatments (spikes were harvested either within 1 week of anthesis, on the day of initial anthesis, or 3 d after initial anthesis), 1 leaf treatment (insect feeding was simulated using an 18-gauge needle to puncture leaves in many locations, leaves were harvested 24 h later), and 1 root treatment (roots harvested 5 weeks after germination) were performed.
WGS sequencing.
All WGS sequencing reads for Yugu1 were collected with standard Sanger sequencing protocols on ABI 3730xl capillary sequencing machines at the Department of Energy Joint Genome Institute in Walnut Creek, California, and the HudsonAlpha Institute, Huntsville, Alabama. Four different-sized libraries were used as templates for the subclone sequencing process and both ends were sequenced: 2,512,497 reads from the 2.89 kb sized library, 2,747,135 reads from the 6.4 kb sized library, 371,711 reads from the 36.7 kb fosmid library, and 105,216 reads from the 121 kb BAC library were produced.
Genome assembly and chromosome anchoring.
The WGS sequence reads were assembled using a modified version of Arachne v.20071016 with parameters maxcliq1 = 100, correct1_passes = 0 and BINGE_AND_PURGE = True52. This produced 597 scaffold sequences (548 that were >1 kb), with N50 of 12.3 Mb, 56 scaffolds > 100 kb, and total scaffold size of 402.4 Mb of which only 1.2% was represented by gaps. Each scaffold was screened against bacterial proteins, organelle sequences and GenBank nr and removed if found to be a contaminant. For further quality control, additional scaffolds were removed as likely false if they consisted of >95% 24 mers that occurred four other times in the scaffolds >50 kb or if the scaffold contained only unanchored RNA sequences.
For the map integration and chromosome-scale pseudomolecule construction, markers from the genetic map were placed using the highest scoring location in the genome with BLAT and blastn. Thirteen breaks were made in scaffolds based on marker discontiguity and 54 joins were made to form the pseudomolecules. Each map join is denoted with 10,000 Ns. These pseudomolecules were then rearranged to match the existing linkage group numbers for the S. italica × S. viridis genetic map. The first nine scaffolds of the release correspond to the chromosome-scale pseudomolecules. The chromosome-scale pseudomolecules contain 396.7 Mb out of 400.9 Mb (99.0%) of the assembled sequence. The final assembly contains 336 scaffolds that cover 400.9 Mb of the genome with a contig N50 of 126.3 kb and a scaffold N50 of 47.3 Mb.
Screening and final assembly release.
The remaining scaffolds were classified in various bins depending on sequence content. Contamination was identified using megablast against GenBank nr and blastp against a set of known microbial proteins. No scaffolds were identified as due to contamination. Additional scaffolds were classified as unanchored rDNA (53), chloroplast (9), mitochondrial (1) and repetitive (108) DNA. Another 50 scaffolds were removed that were <1 kb in sequence length. The resulting final statistics are shown in Supplementary Table 3c.
Large insert clone sequencing.
A total of 105,216 BAC end sequence reads were attempted using Sanger technology on the entire S. italica BAC library. This project generated 93,989 reads (89.3%) with acceptable quality, thereby generating ∼4 Mb of data and 44,284 cases of high-quality reads at both ends of individual BACs.
BAC clones were selected randomly from the S. italica library, were sequenced by the Sanger method to full-length and finished to a high-degree of accuracy. After manual inspection of the assembled sequences, finishing was performed both by resequencing plasmid subclones and by walking on plasmid subclones or the BAC clone using custom primers. All finishing reactions were performed using dGTP BigDye Terminator Chemistry (Applied Biosystems). Hard-to-sequence gaps or small repeats were completed using small insert shatter libraries generated using the Roche/454 sequencing technology or transposon libraries generated using Sanger technology. Finished clones contain no gaps and are estimated to contain less than one error per 10,000 bp.
Fosmids were analyzed by shotgun Sanger sequencing to 8–12× redundancy on the ABI3730xl, as previously described, followed by standard assembly and annotation approaches37.
RNA-Seq analysis.
S. italica (Yugu1) seedlings were grown under an 80:20 mix of metal halide/capsylite halogen lamps at light intensity of 550 μmol/m2/sec, 12 h light, 31 °C L/22 °C D and 50% relative humidity. Tissue was collected from leaf 4 at 10 days after planting and 3 h into the light period from four 1-cm segments: 0–1cm above the leaf-four ligule; 1–0 cm below the leaf-three ligule; 2–3 cm above the leaf-three ligule; and 2–1 cm below the leaf-four tip. Tissue was pooled from 25 seedlings, and three biological replicates were harvested. RNA was extracted using Trizol reagent and libraries constructed using Illumina reagents according to the manufacturer's protocols. Approximately 580 million paired-end reads that passed filtering were used to validate gene model predictions. RNA-Seq–based annotation reconstruction was performed using Cufflinks53 with the alignment data generated by TopHat54 using default settings. Cufflinks-produced gene features were generated by Cufflinks output and compared to reference annotation.
Sequence validation.
Finished Setaria BAC sequences were compared against the assembly using BLAT55 and then a fine alignment was made with a localized Needleman-Wunsch analysis (as implemented at JGI in BioPerl::deAlign; Y.M. Chan, Stanford Human Genome Center, personal communication) to identify discrepancies between the clones and assembly. The alignment showed 5,079,945 out of 5,148,103 bps matching from the clones. This lack of a perfect match suggests that the Yugu1 employed for BAC library construction (which was derived from seed of a separate set of plants from those highly inbred Yugu1 seed used for the WGS) may have contained some residual heterozygosity. These clones were also plotted against the assembly using Dotter and the resulting plots examined for major rearrangements and discrepancies (Supplementary Figs. 8 and 9).
Gene prediction in the full genome sequence.
Gene predictions were performed through the standard JGI plant gene annotation pipeline56. Genes were identified from BLAT alignments of PASA (Program to Assemble Spliced Alignments) EST assemblies and EXONERATE (a splice-site-informed software package for peptide, EST and genome alignment) alignments of homologous peptides from the Arabidopsis, Brachypodium, rice and sorghum proteomes with 2 kb of wiggle room. The genomic sequence, predicted peptides and EST open reading frame for each candidate gene were fed into GenomeScan, FGENESH+ and FGENESH_EST for gene prediction. A best gene prediction per locus was chosen based on EST assembly and homologous peptide alignment support. The selected gene predictions were then fed into the PASA pipeline, where EST assemblies were obtained for gene model improvement, including the addition of untranslated regions. PASA-improved gene model transcripts were subjected to filtering based on how well the transcript coding sequence was supported by ESTs and/or peptide homology, and not overlapped with repeats for more than 20% of the candidate gene length. The filtered gene model peptides were assigned PFAM and PANTHER domains, and gene models with 30% or more of their length assigned with transposon domains were filtered out (Supplementary Note 2).
Transposable element annotation.
The strategy for transposable element discovery consisted of three steps: (1) finding intact transposable elements, (2) building a transposable element database by constructing exemplars57, and (3) scanning the genome to find all (intact and truncated) elements with RepeatMasker (http://www.repeatmasker.org/RepeatModeler.html). Three types of methods were used to discover intact transposable elements in the genome. The first were structural search tools to find LTR retrotransposons (LTR_FINDER58 and LTR_STRUC59) and Helitrons60. The second were transposable element protein-based scans used to find protein-encoding transposable elements such as LINEs, Tc1/Mariner, hAT, Mutator, PIF/Harbinger and CACTA elements. The genome was first scanned by blastp (E-value = 10−5) using known transposable element protein domains as query; then matched regions as well as their flanking sequences were extracted and grouped according to domains. Subsequently, for each group, the domain sequences of group members were aligned and a neighbor-joining tree was built. For clades in the tree, corresponding DNA sequences were aligned and the alignment was inspected manually to find intact elements. The third approach was de novo repeat discovery using the programs RepeatModeler (www.repeatmasker.org/RepeatModeler.html) and MITE-Hunter61 to find noncoding transposable elements such as SINEs and MITEs. All of the program outputs were manually inspected to eliminate artifacts.
Dating times of transposable element insertion and amplification.
The insertion dates for LTR retrotransposons were estimated by the degree of divergence of their two LTRs, as previously described31. The dates of amplification for other types of transposable elements were estimated by comparing the degree of homology of the most closely related copies32. This latter approach, due to the frequent loss of some transposable element copies (primarily by segregation) will tend to miss many transposition events, thus providing an underrepresentation of transposition and an overrepresentation of more ancient (that is, fixed) transposons.
Gene annotation on fully sequenced BACs and fosmids.
BACs and fosmids were first subjected to AUGUSTUS62 using maize parameters to predict gene models, then known transposable elements and/or repeats were identified by comparing predicted gene models to the S. italica transposable element database built in this research and an in-house transposable element protein database. The filtered models were then compared to the GenBank nonredundant protein database, the TAIR10 A. thaliana proteins (http://www.arabidopsis.org/) and IRGSP/RAP build 5 O. sativa proteins (http://rapdb.dna.affrc.go.jp/). Comparison results were layered in Apollo and manual inspection was done to eliminate unreliable gene models following the criteria described previously45. The retained gene models were compared to the current S. italica gene set to evaluate the gene coverage in the current assembly. All of the BACs and fosmids were compared to the current assembly to investigate whether they were recovered by the assembly. Criteria for recovery were as previously described63.
SNP Analysis.
SNP markers for genetic mapping were identified from sequence data generated from seven lanes of 2 × 76 paired-end Illumina reads from a library of 48 pooled A10 × italica RILs. Bases were aligned to the reference Yugu1 genome using Burrows Wheeler Transform64, trimming ends of base quality Q < 15. Only reads with mapping quality q ≥ 29 and with both members of the pair mapping to the genome were used for subsequent SNP analysis. Such reads covered the genome with an average depth of about 27×. A total of 317,973,400 positions in the genome had coverage between 12× and 48× of Q30 or better. Such positions were defined as eligible sites, that is, sites with sufficient information to assess whether or not the pooled RILs show variation at the site. Of the eligible sites, 3,149,093 (∼1%) were polymorphic. Hence, to find SNP markers for genome mapping, targeted SNPs were required to have an alignment depth between 12 and 48 reads, base and mapping quality scores of at least 30, and 60 bp of monomorphic sequence on either side of the SNP site. Using these criteria, a set of 290,317 SNPs was extracted from the original 3,149,093 and 992 of these were used for generating a genetic map.
To assess the local variation in divergence across the genome, eligible sites were combined into adjacent 10-kb bins spanning no more than 20 kb in real genomic space. Examples of such distributions are shown in Supplementary Figure 10.
Mapping populations and construction of a genetic map.
The mapping population was derived by eight generations of single-seed descent from the original S. italica acc. B100 × S. viridis acc A10. For F2 population development, see reference 11. From the identified set of 290,317 SNPs flanked by 60 bp of monomorphic sequence, SNPs that were spaced at ∼100 kb intervals along the draft Yugu1 assembly were selected using an in-house perl script and analyzed using the Illumina Array Design Tool. The final set of 1,536 SNPs that made up the four 384-SNP custom Golden Gate panels (OPAs) was selected to (1) have an Array Design Score ≥ 0.91 and (2) evenly cover as many scaffolds ≥50 kb in the draft assembly as possible. Each of the four 384-SNP OPAs was used to genotype the parents and 188 RIL progeny (F8 generation) from the B100 × A10 cross. Genotyping was done using the Illumina VeraCode technology detected by the Illumina BeadXpress Reader System and analyzed using the Illumina GenomeStudio Data Analysis Software and an in-house perl script. Linkage analysis was carried out using JoinMap 4 (ref. 65). Linkage groups were established at a LOD (logarithm (base 10) of odds) score ≥ 6. Marker orders were determined using the maximum likelihood algorithm and verified manually. The genetic map was drawn using MapChart66.
Interspecies genome comparisons.
The annotated transcripts in the foxtail millet genome assembly (35,158 loci in nine chromosomes, JGI v2.1; this manuscript), sorghum assembly (34,008 loci in ten chromosomes, JGI v1.0)22 and rice assembly (34,781 representative gene loci in 12 chromosomes, IRGSP build 5)67,68 were downloaded. Only the first transcript for each gene was retrieved. A blastp search was conducted with the sorghum and rice gene sets against the foxtail millet gene set. The top two hits with an E-value < 1e-5 were recorded. S. italica genes that were identified by five or more rice and/or sorghum genes were excluded from the set of homologous gene pairs.
The homologous gene pairs were used to draw pairwise comparative dot plots (Supplementary Fig. 11) using R69, and to detect syntenic blocks using the software MCscan70. The colinear segments for all possible pairs of chromosomes were detected using an empirical scoring scheme, min {−log10E, 50} for each gene pair and a −1 gap penalty for each 10-kb distance between any two consecutive gene pairs. Syntenic blocks with a score >300 and E-value <1e-10 were retrieved.
Use of the Setaria physical map in assisting the assembly of the switchgrass WGS sequence.
Peptides obtained from the phytozome website were aligned using tblastn (-f 500 –m 8 –e 1.0E-2) to the current set of switchgrass contigs. The subset of contigs containing peptide alignments were extracted and aligned to the Setaria genome using exploratory cross species parameters (-p blastn -r 1 –q -1 –G 1 –E 2 –W 9 –F “m D” –U –m 8). Best placements were selected based on the total number of aligned bases, and a specific gene on the front end of scaffold_1 in the Setaria genome was randomly selected for example purposes.
Gene family analyses.
For analysis of drought-induced genes, the protein sequences from S. italica, S. bicolor (http://www.phytozome.net/), Z. mays (http://www.phytozome.net/) and O. sativa (http://www.phytozome.net/) were clustered using an all-against-all blastp search of these protein sequences followed by clustering analysis using TRIBE-MCL with an inflation value of 1.2 (ref. 71) and those clusters with five or more members were selected for further analysis. Upregulated genes in drought stress were obtained using blastn, with an e-value cutoff of 1 × 10−10 using published EST data72,73. Possible significance of differences in gene copy numbers was determined using a cumulative Poisson distribution and adjusted with Benjamini and Hochberg correction48.
Phylogenetic analysis of flowering-time genes.
Peptide sequences for genes involved in flowering-time pathways in Arabidopsis, rice and Brachypodium were identified from the literature44 and orthologous grass sequences obtained from GenBank and Phytozome (http://www.phytozome.net/). Additional sequences from Setaria were found using tblastn routines against a database of genomic sequence data created from the draft 8× Setaria assembly, in order to identify proteins that may not have been annotated in the Phytozome database. Sequences were aligned in an online server version of MAFFT version 6 (ref. 74), using the L-INS-I 'slow and accurate' option for alignment of sequences with conserved domains and large gaps. Aligned FASTA files were then imported into Geneious75 version 5.4.4 and maximum likelihood trees were computed in PhyML76, using evolutionary models calculated in ProtTest77. Analysis of the support of clades produced in maximum likelihood trees was carried out on 100 bootstrapped data sets. Patterns of gene duplication and deletion were estimated from the phylogenies, and likely Setaria homologs of flowering-time genes were identified through phylogenetic position, by reciprocal BLAST analysis, and, where possible, by a pattern of synteny in the Phytozome and the Plant Genome Duplication Database70,78. For analysis of variation between foxtail and green millet, we used individual genes of interest obtained from the Setaria genome as target sequences for assembly of the green foxtail genomic reads in Geneious version 5.4.4. These reads were then assembled into contigs and compared with the Setaria sequence in MacClade version 4.08 (ref. 79). Both coding and genomic sequences were used as targets, so that the recovered green foxtail sequences could be annotated and coding regions easily compared.
Small RNA analysis.
Trimmed S. italica small RNA sequences (smRNA) were downloaded from http://smallrna.udel.edu/project_data.php on December 28, 2011. These sequences were searched against all transposable elements annotated in this study using BLAT with the following parameters: -minScore = 18, -tileSize = 7.
To identify putative miRNA families in Setaria, we downloaded known mature miRNA sequences from miRBase (Release 18)80. miRBase data were filtered to include only Viridiplantae species. The smRNA data set used above was then searched against these miRBase sequences. Sequences were required to perfectly match along the entire length of the shortest sequence in the comparison. miRNA family coverage across the assembly was assessed by searching all smRNA sequences associated with each family against current scaffolds using BLAT as above, except that only full-length, perfect matches to smRNAs were counted.
Accession codes.
Data from the whole genome shotgun sequencing project have been deposited at DDBJ/EMBL/GenBank under the accession AGNK00000000. The version described in this paper is the first version, AGNK01000000. EST and RNA-Seq data have been deposited at NCBI in the EST database (JK546897–JK608602) and in the short read archive (SRX116346–SRX116357). BES data for foxtail millet and switchgrass are in GSS (GS026189–GS122438, HR309496–HR503629 and JM786703–JM972700) at the National Center for Biotechnology Information (NCBI).
References
Dekker, J. The foxtail (Setaria) species-group. Weed Sci. 51, 641–656 (2003).
Scott, B.A., Vangessel, M.J. & White-Hansen, S. Herbicide-resistant weeds in the United States and their impact on extension. Weed Technol. 23, 599–603 (2009).
Bettinger, R.L., Barton, L. & Morgan, C. The origins of food production in north China: A different kind of agricultural revolution. Evol. Anthropol. 19, 9–21 (2010).
Harlan, J.R. Crops and Man. (American Society of Agronomy, Madison, Wisconsin, 1975).
Jusuf, M. & Pernes, J. Genetic variability of foxtail millet (Seteria italica P. Beauv.). Theor. Appl. Genet. 71, 385–391 (1985).
Le Thierry d'Ennequin, M., Panaud, O., Toupance, B. & Sarr, A. Assessment of genetic relationships between Setaria italica and its wild relative S. viridis using AFLP markers. Theor. Appl. Genet. 100, 1061–1066 (2000).
Hunt, H. et al. Millets across Eurasia: chronology and context of early records of the genera Panicum and Setaria from archaeological sites in the Old World. Veg. Hist. Archaeobot. 17, 5–18 (2008).
Hirano, R. et al. Genetic structure of landraces in foxtail millet (Setaria italica (L.) P. Beauv.) revealed with transposon display and interpretation to crop evolution of foxtail millet. Genome 54, 498–506 (2011).
Wang, C. et al. Population genetics of foxtail millet and its wild ancestor. BMC Genet. 11, 90–102 (2010).
Darmency, H. Incestuous relations of foxtail millet (Setaria italica) with its parents and cousins, in Crop Ferality and Volunteerism (ed. Gressel, J.) 81–96 (CRC Press, 2005).
Wang, Z.M., Devos, K.M., Liu, C.J., Wang, R.Q. & Gale, M.D. Construction of RFLP-based maps of foxtail millet, Setaria italica (L.) P. Beauv. Theor. Appl. Genet. 96, 31–36 (1998).
Naciri, Y., Darmency, H., Belliard, J., Dessaint, F. & Pernès, J. Breeding strategy in foxtail millet, Setaria italica (L.P. Beauv.), following interspecific hybridization. Euphytica 60, 97–103 (1992).
Darmency, H. & Pernes, J. Use of wild Setaria viridis (L.) Beauv. to improve triazine resistance in cultivated S. italica (L.) by hybridization. Weed Res. 25, 175–179 (1985).
Doust, A.N., Kellogg, E.A., Devos, K.M. & Bennetzen, J.L. Foxtail millet: A sequence-driven grass model system. Plant Physiol. 149, 137–141 (2009).
Brutnell, T.P. et al. Setaria viridis: A model for C-4 photosynthesis. Plant Cell 22, 2537–2544 (2010).
Clayton, W.D. & Renvoize, S.A. Genera Graminum: Grasses of the World. Kew Bulletin Additional Series XIII (Royal Botanic Gardens, Kew, 1986).
Kellogg, E.A., Aliscioni, S.S., Morrone, O., Pensiero, J. & Zuloaga, F. A phylogeny of Setaria (Poaceae, Panicoideae, Paniceae) and related genera, based on the chloroplast gene ndhF. Int. J. Plant Sci. 170, 117–131 (2009).
Aliscioni, S.S., Giussani, L.M., Zuloaga, F.O. & Kellogg, E.A. A molecular phylogeny of Panicum (Poaceae: Paniceae): tests of monophyly and phylogenetic placement within the Panicoideae. Am. J. Bot. 90, 796–821 (2003).
Giussani, L.M., Cota-Sánchez, J.H., Zuloaga, F.O. & Kellogg, E.A. A molecular phylogeny of the grass subfamily Panicoideae (Poaceae) shows multiple origins of C4 photosynthesis. Am. J. Bot. 88, 1993–2012 (2001).
Vicentini, A., Barber, J.C., Giussani, L.M., Aliscioni, S.S. & Kellogg, E.A. Multiple coincident origins of C4 photosynthesis in the Mid-to Late Miocene. Glob. Change Biol. 14, 2963–2977 (2008).
Doust, A.N. & Kellogg, E.A. Inflorescence diversification in the panicoid “bristle grass” clade (Paniceae, Poaceae): evidence from molecular phylogenies and developmental morphology. Am. J. Bot. 89, 1203–1222 (2002).
Paterson, A.H. et al. The Sorghum bicolor genome and the diversification of grasses. Nature 457, 551–556 (2009).
Sang, T. Genes and mutations underlying domestication transitions in grasses. Plant Physiol. 149, 63–70 (2009).
Devos, K.M., Wang, Z.M., Beales, J., Sasaki, T. & Gale, M.D. Comparative genetic maps of foxtail millet (Setaria italica) and rice (Oryza sativa). Theor. Appl. Genet. 96, 63–68 (1998).
Zamir, D. & Tadmor, Y. Unequal segregation of nuclear genes in plants. Bot. Gaz. 147, 355–358 (1986).
Jenczewski, E. et al. Insight on segregation distortions in two intraspecific crosses between annual species of Medicago (Leguminosae). Theor. Appl. Genet. 94, 682–691 (1997).
Lorieux, M., Perrier, X., Goffinet, B., Lanaud, C. & González León, D. Maximum-likelihood models for mapping genetic markers showing segregation distortion. 2. F2 populations. Theor. Appl. Genet. 90, 81–89 (1995).
Lu, H., Romero-Severson, J. & Bernardo, R. Chromosomal regions associated with segregation distortion in maize. Theor. Appl. Genet. 105, 622–628 (2002).
Devos, K.M., Ma, J., Pontaroli, A.C., Pratt, L.H. & Bennetzen, J.L. Analysis and mapping of randomly chosen BAC clones from hexaploid bread wheat. Proc. Natl. Acad. Sci. USA 102, 19243–19248 (2005).
Schnable, P.S. et al. The B73 maize genome: complexity, diversity and dynamics. Science 326, 1112–1115 (2009).
SanMiguel, P., Gaut, B.S., Tikhonov, A., Nakajima, Y. & Bennetzen, J.L. The paleontology of intergene retrotransposons of maize: dating the strata. Nat. Genet. 20, 43–45 (1998).
Yang, L. & Bennetzen, J.L. Distribution, diversity, evolution and survival of Helitrons in the maize genome. Proc. Natl. Acad. Sci. USA 106, 19922–19927 (2009).
Kim, J.-S. et al. Chromosome identification and nomenclature of Sorghum bicolor. Genetics 169, 1169–1173 (2005).
Kimber, G. The addition of the chromosomes of Aegilops umbellulata to Triticum aestivum (var. “Chinese Spring”). Genet. Res. 9, 111–114 (1967).
Devos, K.M. & Gale, M.D. Genome relationships: The grass model in current research. Plant Cell 12, 637–646 (2000).
Okada, M. et al. Complete switchgrass genetic maps reveal subgenome colinearity, preferential pairing and multilocus interactions. Genetics 185, 745–760 (2010).
Pontaroli, A.C. et al. Gene content and distribution in the nuclear genome of Fragaria vesca. Plant Genome 2, 93–101 (2009).
Sharma, M.K. et al. A genome-wide survey of switchgrass genome structure and organization. PLoS ONE 7, e33892 (2012).
Christin, P.A., Salamin, N., Kellogg, E.A., Vicentini, A. & Besnard, G. Integrating phylogeny into studies of C4 variation in the grasses. Plant Physiol. 149, 82–87 (2009).
Sage, R.F. The evolution of C4 photosynthesis. New Phytol. 161, 341–370 (2004).
Christin, P.A., Salamin, N., Savolainen, V., Duvall, M.R. & Besnard, G. C4 photosynthesis evolved in grasses via parallel adaptive genetic changes. Curr. Biol. 17, 1241–1247 (2007).
Corbesier, L. et al. FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis. Science 316, 1030–1033 (2007).
Chardon, F. & Damerval, C. Phylogenomic analysis of the PEBP gene family in cereals. J. Mol. Evol. 61, 579–590 (2005).
Higgins, J.A., Bailey, P.C. & Laurie, D.A. Comparative genomics of flowering time pathways using Brachypodium distachyon as a model for the temperate grasses. PLoS ONE 5, e10065–e10090 (2010).
Liu, R. et al. GeneTrek analysis of the maize genome. Proc. Natl. Acad. Sci. USA 104, 11844–11849 (2007).
Bennetzen, J.L., Coleman, C., Ma, J., Liu, R. & Ramakrishna, W. Consistent over-estimation of gene number in complex plant genomes. Curr. Opin. Plant Biol. 7, 732–736 (2004).
Bennetzen, J.L. Transposable elements, gene creation and genome rearrangement in flowering plants. Curr. Opin. Genet. Dev. 15, 621–627 (2005).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. B Stat. Methodology 57, 289–300 (1995).
Ronquist, F. & Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Zwickl, D.J. Genetic Algorithm Approaches for the Phylogenetic Analysis of Large Biological Sequence Datasets under the Maximum Likelihood Criterion. PhD thesis, Univ. Texas, Austin (2006).
Peterson, D.G., Tomkins, J.P., Frisch, D.A., Wing, R.A. & Paterson, A.H. Construction of plant bacterial artificial chromosome (BAC) libraries: an illustrated guide. J. Agric. Genomics 5, 1–100 (2000).
Jaffe, D.B. et al. Whole-genome sequence assembly for mammalian genomes: Arachne 2. Genome Res. 13, 91–96 (2003).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Kent, W.J. BLAT–the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Goodstein, D.M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186 (2012).
Baucom, R.S. et al. Exceptional diversity, non-random distribution, and rapid evolution of retroelements in the B73 maize genome. PLoS Genet. 5, e1000732 (2009).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–268 (2007).
McCarthy, E.M. & McDonald, J.F. LTR_STRUC: a novel search and identification program for LTR retrotransposons. Bioinformatics 19, 362–367 (2003).
Yang, L. & Bennetzen, J.L. Structure-based discovery and description of plant and animal Helitrons. Proc. Natl. Acad. Sci. USA 106, 12832–12837 (2009).
Han, Y. & Wessler, S.R. MITE-Hunter: a program for discovering miniature inverted-repeat transposable elements from genomic sequences. Nucleic Acids Res. 38, e199 (2010).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644 (2008).
Al-Dous, E.K. et al. De novo genome sequencing and comparative genomics of date palm (Phoenix dactylifera). Nat. Biotechnol. 29, 521–527 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Stam, P. Construction of integrated genetic linkage maps by means of a new computer package: JoinMap. Plant J. 3, 739–744 (1993).
Voorrips, R.E. MapChart: software for the graphical presentation of linkage maps and QTLs. J. Hered. 93, 77–78 (2002).
Itoh, T. et al. Curated genome annotation of Oryza sativa ssp. japonica and comparative genome analysis with Arabidopsis thaliana. Genome Res. 17, 175–183 (2007).
Tanaka, T. et al. The Rice Annotation Project Database (RAP-DB): 2008 update. Nucleic Acids Res. 36, D1028–D1033 (2008).
R Development Core Team. R: A Language and Environment for Statistical Computing. (R Foundation for Statistical Computing, Vienna, 2009).
Tang, H. et al. Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res. 18, 1944–1954 (2008).
Enright, A.J., Van Dongen, S. & Ouzounis, C.A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Zhang, J. et al. Construction and application of EST library from Setaria italica in response to dehydration stress. Genomics 90, 121–131 (2007).
Lata, C., Sahu, P.P. & Prasad, M. Comparative transcriptome analysis of differentially expressed genes in foxtail millet (Setaria italica L.) during dehydration stress. Biochem. Biophys. Res. Commun. 393, 720–727 (2010).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Drummond, A.J. et al. Geneious v5.4 (http://www.geneious.com/; 2011).
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Abascal, F., Zardoya, R. & Posada, D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21, 2104–2105 (2005).
Tang, H. et al. Synteny and collinearity in plant genomes. Science 320, 486–488 (2008).
Maddison, D.R. & Maddison, W.P. MacClade 4. Analysis of Phylogeny and Character Evolution (Sinauer Associates, 2000).
Kozomara, A. & Griffiths-Jones, S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 39, D152–D157 (2011).
Acknowledgements
The studies conducted by the US Department of Energy Joint Genome Institute, the BioEnergy Science Center and the Joint Bioenergy Institute are supported by the Office of Science of the US Department of Energy under contract numbers DE-AC02-05CH11231, DE-AC05-00OR22725 and DE-AC02-05CH11231, respectively. This research was also supported by grants from the Department of Energy (DE-FG02-08ER64636), National Institute of Food and Agriculture Plant Feedstock Genomics for Bioenergy Program (no. 2008-35504-04851), National Science Foundation (IOS-0701736 and DBI-0607123), the Oklahoma Center for the Advancement of Science and Technology (PSB08-007, PS11-035B), and the University of Missouri Research Board. We thank C. Saski at Clemson University for the construction of the Setaria italica and switchgrass BAC libraries, M. Udvardi at the Noble Foundation for providing switchgrass genomic DNA and B. Meyers at the University of Delaware for small RNA annotation advice.
Author information
Authors and Affiliations
Contributions
J.L.B. designed the project, interpreted experimental results and took primary responsibility for writing the manuscript. J.S. was scientific director for reference S. italica sequencing, assembly and map integration. H.W. analyzed all of the transposable elements in the Setaria genome. R.P. prepared all of the RNAs for the EST analysis and participated in switchgrass fosmid library construction, and in the screening, sequencing and genome comparison for switchgrass and Setaria fosmids. J.H. led in all switchgrass fosmid activities. A.C.P. prepared the Setaria fosmid library and sequenced and/or annotated some of these fosmids. M.E. prepared nuclear DNA from S. italica and P. virgatum. L.F. analyzed SNP frequency and distribution. J.N.V. designed, performed, interpreted and described all bioinformatic analyses of small RNAs in this project. J.G. led BES data generation, BAC quality control analysis and project management at HudsonAlpha. J.J. led on switchgrass resource analysis. K.B. coordinated sequence data generation for Setaria genomes, RILs and transcripts at JGI. E.L. directed sequencing and assembly of Setaria ESTs. U.H. identified SNPs for mapping, helped in SNP diversity and/or distribution analysis, and analyzed switchgrass polymorphism for characterization of homoeologous divergence. S.D. assembled libraries and managed the library production activities at JGI. X. Wang developed the genotyping arrays, coordinated the genotyping and constructed the Setaria genetic map. X. Wu conducted the whole-genome comparative analyses between Setaria, sorghum and rice. T.M. led on annotation of the Setaria genome, analysis of switchgrass ESTs compared to Setaria, and analyzed synteny between switchgrass and Setaria. J.T. generated comparative sequence data for Panicum species. M.M.-H. analyzed comparative genomic data to identify possible homologs of flowering-time genes from Arabidopsis in Setaria and other grasses. L.W. and P.L. purified RNA, generated RNA-Seq data and interpreted RNA-Seq results. C.-Y.Y. and X.Y. performed comparative analysis of drought-associated genes. M.S., R.S. and P.C.R. designed and characterized the switchgrass BAC libraries. O.P. generated the Setaria RIL population used for mapping. E.A.K. led the phylogenetic work on switchgrass and phylogenetic placement of switchgrass. T.P.B. planned and managed RNA-Seq analysis. A.N.D. conducted comparative genomic and phylogenetic analysis on flowering time and domestication genes and curated seed collections. G.A.T. performed comparative analysis of drought-associated genes. D.R. oversaw all sequence generation and analysis. K.M.D. led the genetic mapping work and the whole-genome comparative analyses, and assisted with the interpretation of these data sets. All authors contributed to the design, implementation, interpretation and manuscript generation on this project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Notes 1–7, Supplementary Tables 1–3, 5–9 and Supplementary Figures 1–12 (PDF 8356 kb)
Supplementary Table 4
Marker names, location of the markers on the genetic map (in cM), position of the markers on the genome assembly (in bp), genotypic scores in the 188 B100 X A10 RILs analyzed and segregation distortion statistics (XLS 2758 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Noncommercial-Share Alike license (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This license does not permit commercial exploitation, and derivative works must be licensed under the same or similar license.
About this article
Cite this article
Bennetzen, J., Schmutz, J., Wang, H. et al. Reference genome sequence of the model plant Setaria. Nat Biotechnol 30, 555–561 (2012). https://doi.org/10.1038/nbt.2196
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2196
This article is cited by
-
Genome-wide identification of AAAP gene family and expression analysis in response to saline-alkali stress in foxtail millet (Setaria italica L.)
Scientific Reports (2024)
-
MDSi: Multi-omics Database for Setaria italica
BMC Plant Biology (2023)
-
Variation of growth and transcriptome responses to arbuscular mycorrhizal symbiosis in different foxtail millet lines
Botanical Studies (2023)
-
Pangenomic analysis identifies structural variation associated with heat tolerance in pearl millet
Nature Genetics (2023)
-
An E2-E3 pair contributes to seed size control in grain crops
Nature Communications (2023)