Abstract
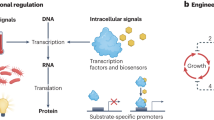
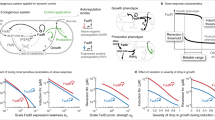
Microbial production of chemicals is now an attractive alternative to chemical synthesis. Current efforts focus mainly on constructing pathways to produce different types of molecules1,2,3. However, there are few strategies for engineering regulatory components to improve product titers and conversion yields of heterologous pathways4. Here we developed a dynamic sensor-regulator system (DSRS) to produce fatty acid–based products in Escherichia coli, and demonstrated its use for biodiesel production. The DSRS uses a transcription factor that senses a key intermediate and dynamically regulates the expression of genes involved in biodiesel production. This DSRS substantially improved the stability of biodiesel-producing strains and increased the titer to 1.5 g/l and the yield threefold to 28% of the theoretical maximum. Given the large number of natural sensors available, this DSRS strategy can be extended to many other biosynthetic pathways to balance metabolism, thereby increasing product titers and conversion yields and stabilizing production hosts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ajikumar, P.K. et al. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science 330, 70–74 (2010).
Atsumi, S., Hanai, T. & Liao, J.C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451, 86–89 (2008).
Steen, E.J. et al. Microbial production of fatty-acid-derived fuels and chemicals from plant biomass. Nature 463, 559–562 (2010).
Zhang, F. & Keasling, J.D. Biosensors and their applications in metabolic engineering. Trends Microbiol. 19, 323–329 (2011).
Keasling, J.D. Manufacturing molecules through metabolic engineering. Science 330, 1355–1358 (2010).
Harcum, S.W. & Bentley, W.E. Heat-shock and stringent responses have overlapping protease activity in Escherichia coli. Implications for heterologous protein yield. Appl. Biochem. Biotechnol. 80, 23–37 (1999).
De Mey, M., Maertens, J., Lequeux, G.J., Soetaert, W.K. & Vandamme, E.J. Construction and model-based analysis of a promoter library for E. coli: an indispensable tool for metabolic engineering. BMC Biotechnol. 7, 34 (2007).
Pfleger, B.F., Pitera, D.J., Smolke, C.D. & Keasling, J.D. Combinatorial engineering of intergenic regions in operons tunes expression of multiple genes. Nat. Biotechnol. 24, 1027–1032 (2006).
Salis, H.M., Mirsky, E.A. & Voigt, C.A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Holtz, W.J. & Keasling, J.D. Engineering static and dynamic control of synthetic pathways. Cell 140, 19–23 (2010).
Farmer, W.R. & Liao, J.C. Improving lycopene production in Escherichia coli by engineering metabolic control. Nat. Biotechnol. 18, 533–537 (2000).
Zhang, F., Rodriguez, S. & Keasling, J.D. Metabolic engineering of microbial pathways for advanced biofuels production. Curr. Opin. Biotechnol. 775–783 (2011).
Fujita, Y., Matsuoka, H. & Hirooka, K. Regulation of FA metabolism in bacteria. Mol. Microbiol. 66, 829–839 (2007).
Cronan, J.E. Jr. In vivo evidence that acyl coenzyme A regulates DNA binding by the Escherichia coli FadR global transcription factor. J. Bacteriol. 179, 1819–1823 (1997).
DiRusso, C.C., Heimert, T.L. & Metzger, A.K. Characterization of FadR, a global transcriptional regulator of FA metabolism in Escherichia coli. Interaction with the fadB promoter is prevented by long chain fatty acyl coenzyme A. J. Biol. Chem. 267, 8685–8691 (1992).
Iram, S.H. & Cronan, J.E. Unexpected functional diversity among FadR FA transcriptional regulatory proteins. J. Biol. Chem. 280, 32148–32156 (2005).
Henry, M.F. & Cronan, J.E. Jr. A new mechanism of transcriptional regulation: release of an activator triggered by small molecule binding. Cell 70, 671–679 (1992).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1–I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
van Aalten, D.M., DiRusso, C.C. & Knudsen, J. The structural basis of acyl coenzyme A-dependent regulation of the transcription factor FadR. EMBO J. 20, 2041–2050 (2001).
Lanzer, M. & Bujard, H. Promoters largely determine the efficiency of repressor action. Proc. Natl. Acad. Sci. USA 85, 8973–8977 (1988).
Spencer, A.K., Greenspan, A.D. & Cronan, J.E. Jr. Thioesterases I and II of Escherichia coli. Hydrolysis of native acyl-acyl carrier protein thioesters. J. Biol. Chem. 253, 5922–5926 (1978).
Corchero, J.L. & Villaverde, A. Plasmid maintenance in Escherichia coli recombinant cultures is dramatically, steadily, and specifically influenced by features of the encoded proteins. Biotechnol. Bioeng. 58, 625–632 (1998).
Friehs, K. Plasmid copy number and plasmid stability. Adv. Biochem. Eng. Biotechnol. 86, 47–82 (2004).
Wilkinson, S.P. & Grove, A. Ligand-responsive transcriptional regulation by members of the MarR family of winged helix proteins. Curr. Issues Mol. Biol. 8, 51–62 (2006).
Carothers, J.M., Goler, J.A., Juminaga, D. & Keasling, J.D. Model-driven engineering of RNA devices to quantitatively program gene expression. Science 334, 1716–1719 (2012).
Lee, T.S. et al. BglBrick vectors and datasheets: A synthetic biology platform for gene expression. J. Biol. Eng. 5, 12 (2011).
Engler, C., Kandzia, R. & Marillonnet, S. A one pot, one step, precision cloning method with high throughput capability. PLoS ONE 3, e3647 (2008).
Hajimorad, M., Gray, P.R. & Keasling, J.D. A framework and model system to investigate linear system behavior in Escherichia coli. J. Biol. Eng. 5, 3 (2011).
Ramsey, S., Orrell, D. & Bolouri, H. Dizzy: stochastic simulation of large-scale genetic regulatory networks. J. Bioinform. Comput. Biol. 3, 415–436 (2005).
Acknowledgements
The authors would like to thank W. Holtz, E. Steen and N. Hillson for discussion and critical reading of the manuscript. This work was supported in part by the Synthetic Biology Engineering Research Center, which is funded by National Science Foundation award no. 0540879, and by the Joint BioEnergy Institute, which is funded by the US Department of Energy, Office of Science, Office of Biological and Environmental Research, through contract DE-AC02-05CH11231. F.Z. is supported by the Postdoctoral Fellowships Program of the Natural Sciences and Engineering Research Council of Canada.
Author information
Authors and Affiliations
Contributions
F.Z. and J.D.K. conceived the project and designed the experiments. F.Z. performed the experiments. J.M.C. designed and performed the DSRS modeling. F.Z., J.M.C. and J.D.K. analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
J.D.K. has a financial interest in Amyris, LS9 and Lygos.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–5 and Supplementary Figures 1–9 (PDF 860 kb)
Rights and permissions
About this article
Cite this article
Zhang, F., Carothers, J. & Keasling, J. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol 30, 354–359 (2012). https://doi.org/10.1038/nbt.2149
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2149
This article is cited by
-
State-of-art engineering approaches for ameliorated production of microbial lipid
Systems Microbiology and Biomanufacturing (2024)
-
A hybrid in silico/in-cell controller for microbial bioprocesses with process-model mismatch
Scientific Reports (2023)
-
Synthetic biology promotes the capture of CO2 to produce fatty acid derivatives in microbial cell factories
Bioresources and Bioprocessing (2022)
-
Development and characterization of a glycine biosensor system for fine-tuned metabolic regulation in Escherichia coli
Microbial Cell Factories (2022)
-
Spatial–temporal regulation of fatty alcohol biosynthesis in yeast
Biotechnology for Biofuels and Bioproducts (2022)