Abstract
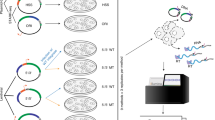
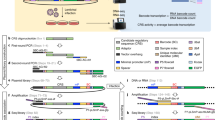
Learning to read and write the transcriptional regulatory code is of central importance to progress in genetic analysis and engineering. Here we describe a massively parallel reporter assay (MPRA) that facilitates the systematic dissection of transcriptional regulatory elements. In MPRA, microarray-synthesized DNA regulatory elements and unique sequence tags are cloned into plasmids to generate a library of reporter constructs. These constructs are transfected into cells and tag expression is assayed by high-throughput sequencing. We apply MPRA to compare >27,000 variants of two inducible enhancers in human cells: a synthetic cAMP-regulated enhancer and the virus-inducible interferon-β enhancer. We first show that the resulting data define accurate maps of functional transcription factor binding sites in both enhancers at single-nucleotide resolution. We then use the data to train quantitative sequence-activity models (QSAMs) of the two enhancers. We show that QSAMs from two cellular states can be combined to design enhancer variants that optimize potentially conflicting objectives, such as maximizing induced activity while minimizing basal activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Lander, E.S. Initial impact of the sequencing of the human genome. Nature 470, 187–197 (2011).
Dorer, D.E. & Nettelbeck, D.M. Targeting cancer by transcriptional control in cancer gene therapy and viral oncolysis. Adv. Drug Deliv. Rev. 61, 554–571 (2009).
Fan, F. & Wood, K.V. Bioluminescent assays for high-throughput screening. Assay Drug Dev. Technol. 5, 127–136 CrossRef (2007).
Loew, R., Heinz, N., Hampf, M., Bujard, H. & Gossen, M. Improved Tet-responsive promoters with minimized background expression. BMC Biotechnol. 10, 81 (2010).
Carey, M., Peterson, C.L. & Smale, S.T. Transcriptional Regulation in Eukaryotes: Concepts, Strategies, and Techniques. Edn. 2 (Cold Spring Harbor Laboratory Press, 2009).
LeProust, E.M. et al. Synthesis of high-quality libraries of long (150mer) oligonucleotides by a novel depurination controlled process. Nucleic Acids Res. 38, 2522–2540 (2010).
Patwardhan, R.P. et al. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis. Nat. Biotechnol. 27, 1173–1175 (2009).
Kinney, J.B., Murugan, A., Callan, C.G. Jr. & Cox, E.C. Using deep sequencing to characterize the biophysical mechanism of a transcriptional regulatory sequence. Proc. Natl. Acad. Sci. USA 107, 9158–9163 (2010).
Panne, D., Maniatis, T. & Harrison, S.C. An atomic model of the interferon-beta enhanceosome. Cell 129, 1111–1123 (2007).
Arnosti, D.N. & Kulkarni, M.M. Transcriptional enhancers: Intelligent enhanceosomes or flexible billboards? J. Cell. Biochem. 94, 890–898 (2005).
Jonsson, J., Norberg, T., Carlsson, L., Gustafsson, C. & Wold, S. Quantitative sequence-activity models (QSAM)–tools for sequence design. Nucleic Acids Res. 21, 733–739 (1993).
Stormo, G.D., Schneider, T.D. & Gold, L. Quantitative analysis of the relationship between nucleotide sequence and functional activity. Nucleic Acids Res. 14, 6661–6679 (1986).
Mayr, B. & Montminy, M. Transcriptional regulation by the phosphorylation-dependent factor CREB. Nat. Rev. Mol. Cell Biol. 2, 599–609 (2001).
Benbrook, D.M. & Jones, N.C. Different binding specificities and transactivation of variant CRE's by CREB complexes. Nucleic Acids Res. 22, 1463–1469 (1994).
Fink, J.S. et al. The CGTCA sequence motif is essential for biological activity of the vasoactive intestinal peptide gene cAMP-regulated enhancer. Proc. Natl. Acad. Sci. USA 85, 6662–6666 (1988).
Kunsch, C., Ruben, S.M. & Rosen, C.A. Selection of optimal kappa B/Rel DNA-binding motifs: interaction of both subunits of NF-kappa B with DNA is required for transcriptional activation. Mol. Cell. Biol. 12, 4412–4421 (1992).
Falvo, J.V., Parekh, B.S., Lin, C.H., Fraenkel, E. & Maniatis, T. Assembly of a functional beta interferon enhanceosome is dependent on ATF-2-c-jun heterodimer orientation. Mol. Cell. Biol. 20, 4814–4825 (2000).
Schneider, T.D. & Stormo, G.D. Excess information at bacteriophage T7 genomic promoters detected by a random cloning technique. Nucleic Acids Res. 17, 659–674 (1989).
Bishop, C.M. Pattern Recognition and Machine Learning (Springer, 2006).
De Mey, M., Maertens, J., Lequeux, G.J., Soetaert, W.K. & Vandamme, E.J. Construction and model-based analysis of a promoter library for E. coli: an indispensable tool for metabolic engineering. BMC Biotechnol. 7, 34 (2007).
Quan, J. et al. Parallel on-chip gene synthesis and application to optimization of protein expression. Nat. Biotechnol. 29, 449–452 (2011).
Matzas, M. et al. High-fidelity gene synthesis by retrieval of sequence-verified DNA identified using high-throughput pyrosequencing. Nat. Biotechnol. 28, 1291–1294 (2010).
Bernstein, B.E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Edelman, G.M., Meech, R., Owens, G.C. & Jones, F.S. Synthetic promoter elements obtained by nucleotide sequence variation and selection for activity. Proc. Natl. Acad. Sci. USA 97, 3038–3043 (2000).
Schlabach, M.R., Hu, J.K., Li, M. & Elledge, S.J. Synthetic design of strong promoters. Proc. Natl. Acad. Sci. USA 107, 2538–2543 (2010).
Holland, J.H. Adaptation in Natural and Artificial Systems: AN Introductory Analysis with Applications to Biology, Control, and Artificial Intelligence Edn. 1 (MIT Press, 1992).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J.R. Stat. Soc. B 57, 289–300 (1995).
Treves, A. & Panzeri, S. The upward bias in measures of information derived from limited samples. Neural Comput. 7, 399–407 (1995).
Acknowledgements
The authors would like to thank E.M. LeProust and S. Chen of Agilent for oligonucleotide library synthesis, R.P. Deering for assistance with Sendai virus infections and the staff of the Broad Institute and the Bauer Core facilities for assistance with data generation. This project was supported by funds from the Broad Institute, the Harvard Stem Cell Institute (T.S.M.), National Human Genome Research Institute grant R01HG004037 (M.K.), the Simons Center for Quantitative Biology at Cold Spring Harbor Laboratory (J.B.K.), National Science Foundation (NSF) grant PHY-0957573 (C.G.C., T.T.) and NSF grant PHY-1022140 (A. Mur.).
Author information
Authors and Affiliations
Contributions
A. Mel., X.Z., P.R., A.G. and T.S.M. developed MPRA and performed the molecular biology experiments. L.W. cultured the cells, and performed the plasmid transfections and luciferase assays. A.Mur., T.T., S.F., C.G.C., J.B.K., M.K., E.S.L. and T.S.M. analyzed the data. T.S.M. wrote the main text with substantial input from all authors. C.G.C. and J.B.K. wrote the Supplementary Notes with substantial input from A. Mur. and T.S.M.
Corresponding author
Ethics declarations
Competing interests
A patent application describing ideas presented in this article has been filed by the Broad Institute.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 5,6, Supplementary Notes and Supplementary Figs. 1–10 (PDF 5994 kb)
Supplementary Table 1
CRE variants (XLSX 5145 kb)
Supplementary Table 2
IFNB variants (XLSX 3389 kb)
Supplementary Table 3
CRE mutagenesis/models (XLSX 39 kb)
Supplementary Table 4
IFNB mutagenesis/models (XLSX 36 kb)
Rights and permissions
About this article
Cite this article
Melnikov, A., Murugan, A., Zhang, X. et al. Systematic dissection and optimization of inducible enhancers in human cells using a massively parallel reporter assay. Nat Biotechnol 30, 271–277 (2012). https://doi.org/10.1038/nbt.2137
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2137
This article is cited by
-
Perfect and imperfect views of ultraconserved sequences
Nature Reviews Genetics (2022)
-
Genomic frontiers in congenital heart disease
Nature Reviews Cardiology (2022)
-
Massively parallel identification of functionally consequential noncoding genetic variants in undiagnosed rare disease patients
Scientific Reports (2022)
-
Functional dissection of inherited non-coding variation influencing multiple myeloma risk
Nature Communications (2022)
-
Quantitative-enhancer-FACS-seq (QeFS) reveals epistatic interactions among motifs within transcriptional enhancers in developing Drosophila tissue
Genome Biology (2021)