Abstract
Distinguishing single-nucleotide variants (SNVs) from errors in whole-genome sequences remains challenging. Here we describe a set of filters, together with a freely accessible software tool, that selectively reduce error rates and thereby facilitate variant detection in data from two short-read sequencing technologies, Complete Genomics and Illumina. By sequencing the nearly identical genomes from monozygotic twins and considering shared SNVs as 'true variants' and discordant SNVs as 'errors', we optimized thresholds for 12 individual filters and assessed which of the 1,048 filter combinations were effective in terms of sensitivity and specificity. Cumulative application of all effective filters reduced the error rate by 290-fold, facilitating the identification of genetic differences between monozygotic twins. We also applied an adapted, less stringent set of filters to reliably identify somatic mutations in a highly rearranged tumor and to identify variants in the NA19240 HapMap genome relative to a reference set of SNVs.
Similar content being viewed by others
Main
The potential applications of whole-genome sequencing in genomic medicine are enormous and range from elucidating disease-causing mutations for monogenic traits to dissecting the molecular genetic basis of complex diseases and discovering somatic alterations in cancer1,2. Many complicated computational analyses are needed, however, to translate raw sequencing data into well-mapped reads, from which a comprehensive list of variants can be derived3,4. The latter process is still difficult, as many detected variants turn out to be genotyping errors. As a result, in many studies, independent mid- to large-scale validation experiments must be done. For instance, >500 somatic SNVs in a lung cancer tumor were validated using mass spectrometry5, whereas other studies resequenced hundreds of SNVs using Sanger sequencing6,7,8. An important drawback of such validation experiments is that they rapidly become as expensive and time-consuming as the whole-genome sequencing experiment itself.
So far, various strategies to improve variant detection in whole-genome sequences have been applied. In family-based studies, relatives enabled the efficient elimination of errors based on Mendelian inheritance patterns, leading to the identification of the culprit gene underlying Miller syndrome9. Another commonly used approach is to apply quality filters that are aimed at selectively removing errors. Every whole-genome sequence reported so far has used filtering to some extent: the most commonly used filters being those that remove sequences with a too-low coverage depth, discard variants with a low-confidence score or eliminate variants located within a cluster of variants3,7,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25. Surprisingly, there is little consensus with respect to which filters should be used and at which threshold they should be applied. As a result, each reported study developed its own heterogeneous set of filters and applied them at various (suboptimal) thresholds. For instance, in the case of the coverage depth filter, thresholds removing sequences with a coverage depth <4×, <10× or <11× were applied6,10,11,16,17,23,24. Additionally, it has not been assessed to which extent filters discard true variants, and how each filter can be optimized in terms of sensitivity and specificity.
In the current study, we therefore optimized a comprehensive set of filters and assessed how each individual filter (or a combination thereof) affected the number of errors, thereby allowing effective filters to be distinguished from those that are less effective. To prove that our filtering strategy was highly efficient, we subsequently applied it using different sequencing technologies to various whole genomes, including those of monozygotic twins, a tumor-normal pair and a HapMap subject.
Results
Development of filters using monozygotic twin genomes
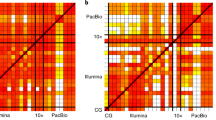
Two peripheral blood leukocyte-derived DNA samples derived from monozygotic twins discordant for schizophrenia were sequenced at high coverage using the short-read sequencing-by-ligation technology from Complete Genomics (CG) and assembled using their Complete Genomics Analysis tools (CGA) (Supplementary Note 1)13. Pairwise comparison of both twin genomes revealed that 95.5% of the reference genome was sequenced in both twins, resulting in 483,725 discordant and 2,846,845 shared SNVs (Table 1). Similar to other reports5,9,13, we discarded positions in which one allele in either of the twins was considered uncertain by CGA. As a result, 8.7% of the reference genome was not considered after removing uncertain calls, leading to 46,376 discordant and 2,725,695 shared SNVs.
As monozygotic twins have near-identical genomes, the majority of discordant SNVs are likely to represent genotyping errors in one of the twins. Inspection of all 46,376 discordant SNVs revealed that a large fraction was located near an indel (19.7%) or cluster of SNVs (25.3%), at positions with low or extremely high coverage (51.9%) or in repetitive DNA regions, such as microsatellites (37.9%). Therefore, in an effort to selectively remove discordant SNVs, we developed two types of filters: (i) filters removing genomic regions of inferior sequence quality (quality filters), and (ii) filters targeting genomic regions with repetitive DNA sequences (repetitive DNA filters). For each individual filter, filter thresholds were determined such that they removed a maximum number of discordant SNVs and a minimum of shared SNVs (Fig. 1 and Supplementary Note 2). Optimized filters were considered effective when the fraction of discordant SNVs removed (Fdiff) was at least twice the fraction of shared SNVs removed (Fshared; Fdiff/Fshared>2). We also calculated the fraction of the genome removed by each filter (Fgenome). Notably, Fgenome does not just represent the fraction of variants removed, but also takes nonvariant sites that are filtered into account (Supplementary Note 2).
Under the assumption that the number of actual differences between the monozygotic twins is very low, we calculated all discordant and shared SNVs between the twins. We considered discordant SNVs as errors and tested every filter for its capacity to selectively reduce discordances, while keeping as many of the shared variants as possible. Three types of filters were developed: (i) filters removing regions of inferior sequencing quality (quality filters), (ii) filters based on intrinsic genome characteristics (repetitive DNA filters), and (iii) filters selecting variants identified with an independent mapping and SNV calling method (consensus filters). The best individual filters were subsequently combined to remove a maximum number of discordances in the twin genomes. The same rationale was applied on the Yoruban NA19240 genome sequenced by CG and Illumina. The allelic imbalance filter was only applied on Illumina data, whereas the uncertain calls filter was only developed for CG data.
Quality filters and repetitive DNA filters
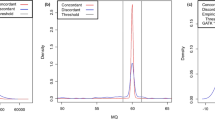
To filter genomic regions of inferior sequencing quality, we selected quality filters based on our observations in the 46,376 discordant SNVs and on a literature survey of whole-genome studies (Supplementary Note 3). In particular, we filtered SNVs according to the coverage depth at a given position, the quality score of an SNV ('variant score'), and the presence of nearby indels or SNV clusters. Standard receiver operating characteristic (ROC) curves and distribution analyses revealed that a coverage depth and variant score threshold of, respectively, <20× and <60 were best to separate discordant SNVs from shared SNVs, although other settings, such as a coverage depth of <10×, can be considered to improve sensitivity at the cost of specificity (Supplementary Note 2). We found that detection of SNVs located near indels or clustered SNVs was most efficiently accomplished by algorithms that removed SNVs located within five base pairs (bp) of an indel or in regions with ∼8 times more SNVs than observed on average across the genome (Supplementary Note 2). Each of these individual filters was found to remove at least twice the fraction of discordant versus shared SNVs at these thresholds (Fdiff/Fshared > 2; Fig. 2). The filters based on the variant score, indels and clustered SNVs discarded very little of the shared twin genome, whereas the coverage depth filter removed a large portion (Fgenome = 4.6%, 4.6%, 4.7% and 28.2%, respectively). The latter clearly illustrates that despite a high average coverage depth (37.9× and 37.3× for the two genomes), a more homogeneous coverage depth distribution is critical to increase the proportion of the genome sequenced at high quality. Overall, the combination of these four quality filters lowered the number of discordant SNVs substantially to 9,537 discordant SNVs, leaving 1,919,487 of the shared SNVs (cumulative Fdiff/Fshared = 3.0 and Fgenome = 28.6%; Table 1).
(a,b) The fraction of SNVs removed by every filter. Shown is the percentage of discordant (Fdiff, light gray) and shared (Fshared, dark gray) variants removed by every filter. (c,d) The ratio Fdiff/Fshared is a measure of the specificity of the individual filters. Filters were considered effective if they removed twice the fraction of discordant versus shared variants (ratio Fdiff/Fshared > 2; dashed line). Filters that failed this criterion are denoted with an asterisk (*). (e,f) The fraction of the reference genome removed by every filter (Fgenome). Data stratified into transcriptome, conserved noncoding and nonconserved noncoding regions are given in Supplementary Note 2. Before filtering, a fraction of the reference genome was not sequenced in both samples (that is, 4.5% for the twins and 4.4% for the NA19240 genomes), representing the lower limit of the Fgenome (dashed line).
Genomic regions with repetitive DNA sequences are prone to error, owing to incorrect mapping to the reference genome. We therefore retrieved all repetitive DNA tracks from the UCSC genome browser26 and designed filters for each of them (Supplementary Note 2). In addition, we optimized a filter for homopolymer stretches. Filters targeting tandem repeats, microsatellites and homopolymer regions were most selective (Fdiff/Fshared > 5) and removed very little of the shared twin genome (Fgenome = 5.8%, 4.5% and 5.2%, respectively). The segmental duplication filter was slightly less effective (Fdiff/Fshared = 2.0; Fgenome = 8.5%), whereas the RepeatMasker and self-chained filters were largely nonspecific and therefore not further considered (Fdiff/Fshared < 2; Fig. 2). When combining the quality and repetitive DNA filters, only 5,387 discordant SNVs and 1,869,370 shared SNVs remained (cumulative Fdiff/Fshared = 2.9 and Fgenome = 35.2%; Table 1). Sanger validation of 252 out of 5,387 discordant SNVs failed to confirm any of these (Supplementary Note 4), thereby revealing that although quality and repetitive DNA filters were very effective, there were still many errors.
Consensus mapping and SNV calling filter
In the 1000 Genomes (1KG) Project14,27, a second mapping and SNV calling algorithm to filter SNVs detected by only one of the algorithms was successfully applied as a 'consensus filter'. We used the RTG2.0 technology as an independent mapping and SNV calling method because it is currently the only other method that can handle CG data. Furthermore, it uses substantially different mapping and variant detection algorithms compared to CGA (Supplementary Note 5). Briefly, CGA and RTG2.0 both use a reference assembly approach, whereas CGA performs additional de novo assembly around predicted variant sites. Both methods also use Bayesian variant calling approaches with different probabilities to identify SNVs. Overall, RTG2.0 resulted in a slightly lower number of mapped sequences, but a higher number of discordant and shared SNVs (Supplementary Note 5). The ratio between discordant and shared SNVs was similar between CGA and RTG2.0, indicating that none of the methods substantially outperformed the other. When applying the consensus filter to the unfiltered genome, resulting in SNVs identified by both CGA and RTG2.0, the Fdiff/Fshared value was 6.0 (Fig. 2). Subsequent application of this filter in combination with all other filters showed a substantial overall improvement, resulting in only 846 discordant and 1,704,701 shared SNVs (cumulative Fdiff/Fshared = 2.5 and Fgenome = 32.0%; Table 1).
Adaptive filtering based on error rate reduction
Based on the assumption that discordant SNVs are the result of errors and shared SNVs are correct, we optimized filters to remove as many discordant SNVs and as few shared SNVs as possible. To confirm that shared SNVs are indeed true variants, we used Illumina single-nucleotide polymorphism (SNP) arrays and selected all SNPs heterozygous on the SNP array. There were 227,943 SNPs before and 182,877 SNPs after filtering, of which 99.924% and 99.997%, respectively, were shared SNVs with the same genotype in the twin genomes. Validation of the discordant SNPs revealed that they were all correctly genotyped by CG. Furthermore, another 670 shared SNVs, of which 135 were novel, were also confirmed using Sequenom (Supplementary Table 1). Although we cannot formally exclude the possibility that a few of the remaining shared SNVs represent false positives in both twins, these data convincingly demonstrate that shared SNVs can be considered 'true variants' and discordant SNVs 'errors'. We could therefore also accurately estimate error rates for our filters. In particular, the error rate for 'novel' SNVs decreased from 23.8% before filtering to 0.17% after cumulative filtering. When estimating error rates for 'all' SNVs, error rates decreased from 14.52% to 0.05%, corresponding to a 290-fold improvement. Also, when considering the percentage of the genome that was available for analysis after filtering, per-base detection error rates were 1.79 × 10−4 before filtering and 4.39 × 10−7 after filtering (Table 1).
Despite this substantial reduction in errors, cumulative filtering removes a considerable fraction of the genome (Fgenome = 32%). In some experimental conditions, a subset of filters that removes a smaller fraction of the genome may have to be applied at the expense of a higher number of errors. However, as filters may affect the same SNVs and therefore partly overlap with each other (Supplementary Note 6), such filter combinations cannot be selected based on the performance of individual filters. We therefore calculated error rates and the percentage of the genome removed for each of the 1,048 possible filter combinations (Supplementary Note 7 and Supplementary Table 2), and plotted their effect with respect to true variants (shared SNVs) and errors (discordant SNVs) that remained after filtering (Fig. 3). On average, the filtered fraction of true variants and errors increased gradually with the number of filters applied, with some filter combinations having more pronounced effects on the fraction of errors than others. Notably, when selecting the filter combination, in which the filtered fraction of true variants and errors is optimally balanced (that is, the filter with the best Matthews correlation coefficient (MCC) value), a combination of three individual filters was identified, including the 'near an indel', 'uncertain calls' and 'microsatellite' filters. In particular, this best MCC combination removed only 9% of the genome, while identifying 24,389 discordant and 2,670,244 shared SNVs. The resulting filtering strategy was implemented in the GenomeComb tool, which can be used to combine variant data from multiple genomes, as well as to extensively annotate and filter the combined genomes.
(a) Results of applying filter combinations to monozygotic twin genomes. Each combination is represented by a circle that is sized and colored according to the number of filters combined. The circle's position indicates the fractions of shared SNVs and discordant SNVs that remained after applying the combination. Shared SNVs represent true variants, and discordant SNVs indicate errors. The combination with the best MCC value (that is, the circle closest to the left-upper corner) consisted of three filters: the near-an-indel, uncertain calls and microsatellite filter. (b) Results of filtering NA19240 genomes. Circles drawn as in a. The combination with the best MCC value comprised the consensus mapping and calling filter, near-an-indel filter and variant score filter. For both a and b, the effect of individual filter combinations relative to all other filter combinations at different filter thresholds can be assessed at http://genomecomb.sourceforge.net/publications/filters/filterselection.php.
True genetic differences in monozygotic twins
As somatic mutations have been reported at a rate of 4.6 × 10−10 bp per generation28,29, it is expected that very few SNVs are found in one monozygotic twin and not the other. A previous whole-genome sequencing experiment of monozygotic twins discordant for multiple sclerosis did not, however, identify true genetic differences30. In an effort to identify such differences in our twins discordant for schizophrenia, all 846 discordant SNVs identified after cumulative filtering were validated using Sanger sequencing. Of the 814 SNVs that could be sequenced, 561 were false positive in one of the twins and 251 were false negative. However, after Sanger sequencing in both directions, two SNVs were confirmed as actual differences (Supplementary Note 4). On the other hand, because 24,389 discordant SNVs were detected after applying the best MCC filters, validation of all discordant SNVs is not realistic, thereby illustrating that the most stringent combination of filters is needed under certain experimental conditions.
The first SNV located on chromosome 4 (position 189,411,955) is heterozygous in the healthy twin. This variant is located in a genomic evolutionary rate profiling (GERP) constraint element of 135 bp with the closest gene >100 kb away31. Intriguingly, the second SNV turned out to be a mosaic variant with a higher frequency of the variant allele in the schizophrenic twin. To confirm this observation, we amplified by PCR the genomic region of both twins and cloned region using the TOPO TA cloning kit. Sequencing of the resulting bacterial clones revealed that 17 out of 60 clones (28%) and 2 out of 54 clones (4%) were variant in the schizophrenic and healthy twin, respectively (Supplementary Note 4). In particular, this variant was located on chromosome X (position 146,050,487) in a nonintact LINE-1 sequence (L1MA4)32, with the closest known gene being a microRNA (miRNA) cluster located ∼28-kb downstream of the variant.
Somatic mutations in ovarian tumor genomes
As many whole-genome sequencing applications involve tumor genomes, we assessed whether our filters were applicable to the genome of a primary serous ovarian tumor and its matched normal DNA (Supplementary Note 1). Serous ovarian tumors are known to have highly rearranged genomes and chromosomal instability33,34. Illumina SNP arrays confirmed that ∼190 chromosomal aberrations and loss-of-heterozygosity regions were present in this tumor (Supplementary Note 8). Additionally, because tumors collected during surgery are often characterized by infiltrating normal cells, we established that our tumor consists of ∼50% normal cells. The structural aberrations and infiltration of nonmalignant cells may seriously complicate the detection of somatic variants in tumor genomes. It is therefore expected that this unfiltered tumor genome will be highly enriched in genotyping errors.
We first performed tumor-normal comparisons and assessed whether step-wise application of the filters to both genomes affected the number of discordant variants (Supplementary Note 8). Overall, 58.7% of all discordances were removed after cumulative filtering compared to only 19.3% of the shared variants. When the same tumor sample was resequenced (after construction of a new sequencing library from the same DNA pool), allowing us to independently validate filters in a true (tumor) replicate, individual filters had similar effects on the number of discordant and shared variants between both replicates (Supplementary Note 9 and Supplementary Table 3). Overall, this indicates that our filtering strategy was also effective in tumor genomes. Nevertheless, substantially more differences were found in the tumor-normal comparison compared to the twins (24,523 versus 846). Highly amplified and loss-of-heterozygosity regions were enriched in these differences, as indicated by the fact that 15.9% of the differences were found on chromosomes 13 and 17, which both show loss of heterozygosity throughout the whole chromosome, but represent only 6.4% of the genome.
To identify putative driver mutations, we restricted our analysis to homozygous-to-heterozygous changes in the coding region. After cumulative filtering, 21 missense mutations were identified, 19 of which were confirmed using Sequenom (Supplementary Table 1). In particular, we found mutations in TP53 and MLH1, which are both recognized cancer genes35,36. On the other hand, when applying the MCC filter combination, we identified 117 somatic variants, 50 of which were confirmed by Sequenom or Sanger sequencing (Table 2). Thus, by cumulative filtering we detected only 38% of the true-positive mutations with high confidence (validation rate of 90%), whereas application of a less stringent set of filters identified all true-positive mutations albeit with lower confidence (validation rate of 43%; Supplementary Note 8). Analysis of an additional tumor-normal pair characterized by fewer chromosomal aberrations (∼120) and a larger tumor percentage (80%), revealed a larger fraction of true-positives after stringent filtering (22 out of 34; 65%) and a higher validation rate after MCC filtering (82%; Supplementary Note 10 and Supplementary Table 4). This demonstrates that our filters were effective in both tumor genomes, but that the extent to which they improved the sensitivity and specificity varied depending on tumor content and the degree of chromosomal instability.
Finally, to assess which of these mutations represent driver mutations, we sequenced a second serous ovarian tumor with high chromosome instability and genotyped an independent series of 120 serous ovarian tumors for all 50 somatic mutations. We confirmed that out of the 50 somatically altered genes in the first serous ovarian tumor, MLH1 was mutated on a different position in the second tumor. Genotyping of 120 additional tumors for these 50 mutations revealed that mutations P238T in PPP1R3A and I71M in SUPT5H were present in a second, whole-genome sequenced serous ovarian tumor (Supplementary Note 10). This strongly suggests that at least these two genes represent driver genes of serous ovarian cancer. Bioinformatics analyses further predicted that 26 out of 50 somatic mutations, including those affecting TP53, MLH1, PPP1R3A and SUPT5H, have a putatively detrimental effect on protein function (Supplementary Note 11 and Supplementary Table 5). In addition, network and over-representation analyses of Gene Ontology terms on all mutated genes showed that ovarian cancer signaling pathways were over-represented (Supplementary Note 11).
Adaptive filtering of an Illumina-sequenced HapMap genome
To validate our filtering pipeline on other sequencing platforms, we adapted our filters for genomes sequenced with Illumina technology. In particular, we used the Yoruban NA19240 HapMap genome that was sequenced at high coverage in the 1KG Project14. Except for the uncertain calls filter, which could not be adapted because Illumina does not report uncertain calls, all filters were successfully optimized (Fig. 1 and Supplementary Note 12). Furthermore, as other studies have applied an allelic imbalance filter3,14, this filter was also optimized for Illumina data. Because NA19240 has also been sequenced by CG (Supplementary Note 1), we calculated the number of discordant and shared SNVs between CG and Illumina data to assess the effect of every individual filter. For this comparison, we used both the unfiltered and the cumulatively filtered CG genome as a reference (Supplementary Note 12 and Supplementary Table 6). Similar to the twin genomes, the most effective filters were those that selectively removed discordant SNVs. Ilumina filters with the best performance were the 'variant score', 'consensus' and 'clustered SNV' filters (Fdiff/Fshared > 5 versus the unfiltered CG genome). Notably, out of all five repetitive DNA filters, only the homopolymer filter was effective on Illumina data (Fig. 2). Overall, when both genomes were cumulatively filtered, only 643 SNVs were discordant and 2,449,744 SNVs were shared between CG and Illumina genomes, corresponding to 0.13% and 63.7% of the original discordant and shared SNVs.
Cross-platform validation and error rate reduction
Finally, we also assessed how Illumina and CG filters improve error rates in the NA19240 genome. The unique availability of CG, Illumina and SOLiD data for NA19240 enabled us to construct a 'golden' variant set, consisting of all unfiltered SNVs shared by at least two sequencing technologies, as well as all SNVs identified by Mendelian inheritance studies for NA19240 in the 1KG Project14 (Supplementary Note 13). In total, this 'golden' set contained 4,092,560 SNVs. Under the assumption that all SNVs detected by more than one sequencing platform are 'true', this golden variant set represents the most complete and accurate list of SNVs available for any genome to date. Notably, we detected more false-positive errors in the unfiltered Illumina data (15.27% versus 4.17% with CG) and more false-negative errors in the CG data (4.97% versus 0.28% with Illumina). Nevertheless, after application of all filters, false-positive and false-negative rates decreased substantially to near-concordance on both platforms, with false-positive and false-negative rates of ∼1% and 0.1%, respectively (Table 3). Overall, these data indicate that our filtering pipeline, independently of the sequencing technology used, substantially reduces error rates in whole genome sequences.
Discussion
In the present study, we optimized a filtering pipeline on monozygotic twins sequenced with CG and adapted this pipeline for genomes sequenced with Illumina using publically available data. Although similar filters have been applied previously, to our knowledge no other single study has reported on the systematic optimization of filter thresholds and assessment of whether their application indeed improves the quality of a genome. Intriguingly, the most effective CG filters were those that removed SNVs located in simple tandem repeat regions or near an indel, whereas the most effective Illumina filters were the consensus and variant score filters. These observations suggest that CG, owing to its specific read structure, is less effective in correctly identifying SNVs in repetitive DNA sequences or indels, but that its in-house developed mapping and SNV calling algorithms are otherwise carefully tuned. On the other hand, indels or repeat regions represent a less prominent source of errors for Illumina data, whereas standard mapping and SNV calling algorithms are more prone to identifying false-positive SNVs.
An important finding of our study is that cumulative application of all filters decreased the error rate by 290-fold in the twins. This error rate was based on the number of discordant and shared SNVs in the twins. Therefore, it did not account for false-negative SNVs (that is, variants missed by CG but identified by other technologies) and provides an estimate of the reproducibility for CG sequencing. On the other hand, in the NA19240 genomes sequenced by CG and Illumina, error rates decreased by 8.11-fold and 15.35-fold, respectively, after filtering. As these error rates were based on discordant and shared SNVs relative to a golden variant set obtained by three independent technologies, they reflect the cross-platform reproducibility of whole-genome sequencing. As such, these error rates also account for platform-specific false-negative SNVs, but not for false negatives missed by all three sequencing technologies.
Up to 29% or 16% of the genome is removed by CG or Illumina filters, respectively. Depending on the experimental conditions, specific combinations of filters, which remove a smaller proportion of the genome at the cost of a higher number of errors, might be necessary. The filter combination with the best MCC value is interesting in this respect, as it considers the cost of removing a 'true variant' or 'error' equal. In CG and Illumina sequences, this best MCC combination removed only 4.7% and 4.9%, respectively, of the genome. Alternatively, thresholds of individual filters, such as the coverage filter, can also be adapted to remove a smaller proportion of the genome. Another attractive approach is to consider our filters as a prioritization method, whereby SNVs are not removed, but ranked based on error rates obtained for the combination of filters affecting the SNVs, as discussed in further detail in Supplementary Note 14.
Notably, adaptive filtering based on the selectivity and specificity of various filter combinations can be critically important for the discovery of genetic variants. Cumulative filtering was essential, for instance, to discover two genetic differences in our monozygotic twins. The first SNV, owing to its location in a gene desert, did not provide any obvious causality for the discordant schizophrenia phenotype of the twins. The second SNV is located in a nonintact LINE-1 element (L1) and was characterized by a substantially higher contribution of the variant allele in the schizophrenic twin. This can be explained by a post-zygotic mutation before the twinning event and an unequal distribution of mutated cells during the twinning event. Alternatively, as nonintact retroviral sequences may retain their ability to be retrotransposed by active transposons nearby during embryogenesis32, retrotransposon activity could have introduced the observed mosaicism. Moreover, as L1-activity has been implicated in neurological diseases37 and retroviral RNAs have been linked to schizophrenia38, this SNV may confirm a potentially relevant disease mechanism contributing to schizophrenia.
Finally, we also demonstrated that the filters improved somatic variant detection in serous ovarian tumors. We confirmed that several mutated genes were also affected in other serous ovarian tumors, thereby indicating that they are causally contributing to tumorigenesis. Notably, ovarian tumors were characterized by a considerable fraction of infiltrating nonmalignant cells and a highly rearranged chromosomal architecture. We found that, depending on the extent to which these characteristics were present, the number of false-negative variants after stringent filtering were higher and the validation rate upon less stringent filtering was lower, thereby suggesting that highly rearranged tumors with many nonmalignant cells, will require additional methods, such as the development of an SNP caller aware of the allelic fraction, to improve variant detection. In conclusion, we demonstrate that adaptive filtering leads to fewer errors, allowing variants of interest, which are not identified without applying the filters, to be reliably detected.
Methods
Sample selection and sequence generation.
Seven samples were selected for whole-genome sequencing by Complete Genomics (CG): two blood samples from a female, Caucasian monozygotic twin pair discordant for schizophrenia (further referred to as Twin 1 and Twin 2), two tumor and matched normal samples taken from two Caucasian ovarian cancer patients (referred to as Tumor 1 or 2 and Normal 1 or 2) and a tumor from an additional ovarian tumor sample (referred to as Tumor 3). The DNA sample from Tumor 1 was sequenced twice, that is, the same DNA sample was sent to CG in two separate batches and independent sequencing libraries were prepared to generate a replicate whole-genome sequence of Tumor 1. The monozygotic twin sisters were 42 years old at the time of blood sampling. The diseased twin was diagnosed with schizophrenia of the paranoid type (DSM IV TR diagnosis 295.30). The tumor-normal pair (Tumor 1 and Normal 1) was collected from a 75-year old patient diagnosed with serous grade 3 ovarian cancer. The second tumor-normal pair (Tumor 2 and Normal 2) was collected from a 61-year old patient diagnosed with grade 3 clear-cell ovarian carcinoma. Sample Tumor 3 was selected because it had a similar copy-number profile as Tumor 1. The sample was also collected during surgery from a patient diagnosed with serous grade 3 ovarian cancer at the age of 56. All three tumors were primary chemonaive tumors. Informed consent was obtained for all individuals. DNA from all samples was extracted using the Qiagen DNAEasy kit.
Paired-end sequencing was performed with the CG service provider using a proprietary sequencing-by-ligation technology. CG also performed primary data analysis, including image analysis, base calling, alignment and variant calling. Reads were mapped to the NCBI Build 36.1 reference genome using a fast algorithm and initial mappings were expanded by local de novo assembly on all regions of the genome that contain SNVs relative to the reference genome. The mapping algorithm included in the CG Analysis (CGA) toolset was used. More detailed explanation of the CG technology and analysis tool set, as well as sequencing statistics obtained for the seven genomes, can be found in Supplementary Note 1.
Sequencing of the HapMap individual NA19240.
Individual NA19240 is the daughter of a Yoruba family in Ibadan, Nigeria. The Yoruban trio (mother NA19238, father NA19239 and daughter NA19240) has been included in the HapMap project39 and was sequenced in pilot 2 of the 1000 Genomes Project (1KG Project)14. In particular, individual NA19240 was sequenced using the Genome Analyzer II (Illumina) and SOLiD technology (Life Technologies). Furthermore, this NA19240 Yoruban genome was also sequenced with CG sequencing technology in a large sequencing effort, in which whole-genome sequences of 69 HapMap genomes were sequenced at high coverage and were made publicly available. NA19240 CG data were downloaded as processed variant files and coverage depth data from the Complete Genomics FTP site (ftp://ftp2.completegenomics.com/YRI_trio/ASM_Build37_2.0.0/NA19240/), whereas NA19240 Illumina data were downloaded as preprocessed, BWA40 mapped reads from the 1KG Data Portal (ftp://ftp.1000genomes.ebi.ac.uk:21//vol1/ftp/data/NA19240/alignment/). High coverage data for all chromosomes were downloaded as separate, preprocessed bam files (NA19240.chrom*.ILLUMINA.bwa.YRI.high_coverage.20100311.bam). For more details we refer readers to the 1KG Project website (http://www.1000genomes.org/) or to the 1KG publication14. SNV calling on the 1KG NA19240 data was performed using the GenomeAnalysisToolKit (GATK version v1.0.4418) Unified Genotyper41 developed at the Broad Institute, by using the same settings as suggested for the pilot 2 data in the 1KG Project (http://www.broadinstitute.org/gsa/wiki/index.php/Unified_genotyper): stand_call_conf 30 (which sets the minimum phred-scaled Qscore; 30 is the standard threshold for high-pass calling), stand_emit_conf 10 (each variant with at least Q10 confidence to be nonreference is shown) and –all-bases (output for each position in the genome is generated, allowing one to determine the coverage depth at each position for further filtering purposes). GATK produces a variant file in VCF format. Detailed information on availability and processing details of the NA19240 sequencing data can be found in Supplementary Note 1.
Evaluation and optimization of SNV filters.
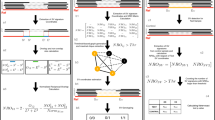
To determine the effect of each filter, we used several metrics according to the definitions below:
Fshared = the proportion of shared SNVs removed by applying the filter, defined as  , where Nshared,UF represents the total number of shared SNVs in the unfiltered genome and Nshared,F represents the number of shared SNVs after applying the filter.
, where Nshared,UF represents the total number of shared SNVs in the unfiltered genome and Nshared,F represents the number of shared SNVs after applying the filter.
Fdiff = the proportion of discordant SNVs removed by applying the filter, defined as  , where Ndiff,UF represents the total number of discordant SNVs in the unfiltered genome and Ndiff,F the number of discordant SNVs after applying the filter.
, where Ndiff,UF represents the total number of discordant SNVs in the unfiltered genome and Ndiff,F the number of discordant SNVs after applying the filter.
Fdiff/Fshared = the ratio of the proportion of discordant SNVs removed versus the proportion of shared SNVs removed; this ratio reflects how many errors versus true SNVs are removed.
Fgenome = percentage of the genome removed after applying the filters.
All twelve filters were developed, optimized and tested individually. These filters included the uncertain calls, variant score, coverage depth, clustered SNVs, near-an-indel, simple repeat, microsatellites, segmental duplications, self-chained regions, Repeat Masker, homopolymer run, near a homopolymer run, consensus mapping and SNV calling filters. Detailed definitions for these filters can be found in Supplementary Note 2. Consensus mapping and SNV calling was performed by algorithms developed by Real Time Genomics (SlimNGS technology, RTG2.0) and were applied to the original sequencing data received from CG and Illumina. Full details are given in Supplementary Note 5.
For all filters handling continuous values, standard ROC curves and MCC values were used to determine optimal Fdiff/Fshared cutoff values (Supplementary Note 2). In particular, we calculated ROC curves for each of the filters according to the following definition: the 'true variant rate' or sensitivity = Fshared and the 'error rate' or specificity = Fdiff . The MCC value for each point in an ROC curve is defined as follows:

Distribution curves for discordant and shared SNVs were used to identify relevant ranges for each filter. The distribution curves give an intuitive interpretation of the optimal threshold, whereas ROC curves provide a quantitative interpretation of how effective these filters are. For each individual filter, we also calculated the fraction of the genome removed (Fgenome). We also combined each of the various individual filters into filter combinations; Fdiff/Fshared and Fgenome values were calculated for each filter combination and are described in Supplementary Note 7. All algorithms and scripts were implemented in the Tcl, C and Perl programming languages.
Concordance analysis using Illumina Omni1-Human arrays.
Genome-wide SNP genotyping of >1 million SNPs was performed using Illumina Human-Omni1 SNP arrays on an Illumina iSCAN (Illumina) for Twin 1, Twin 2, Tumor 1 and Normal 1. Genotype calls on the SNP array were compared with the SNVs obtained from the whole genomes. Positions that were discordant with whole-genome sequence data in all four samples were removed from the analysis because they most likely represent systematic errors on the SNP array. Concordance rates were calculated for all four samples and are shown in Supplementary Note 4.
Validation of shared and discordant SNVs in the monozygotic twins.
While developing our filters, we assumed that shared variants between the twin genomes are true variants. To confirm this, we used Illumina SNP arrays, as described above. To confirm that discordant variants identified during the filtering process were true-positive of false-positive variants, we used Sanger sequencing. Sanger sequencing was done using 10 ng of genomic DNA with 10 pmol of each primer in a standard PCR reaction, followed by ExoSAPit treatment (Amersham Biosciences) and subsequently sequenced using the Big Dye terminator cycle sequencing kit v3.1 according to the manufacturer's instructions (Applied Biosystems). Sequencing reactions were run on a 3730XL DNA Analyzer and the resulting trace files were analyzed using NovoSNPv3.0 (ref. 42). Two large validation experiments were performed with Sanger sequencing: the validation of 324 discordant SNVs randomly selected after the initial filtering steps and the validation of 846 discordant SNVs remaining after all cumulative filters were applied. Detailed data for these experiments are listed in Supplementary Table 1.
Validation of a mosaic variant between the monozygotic twins.
We confirmed that the variant on chromosome X was a mosaic variant between the twins. The PCR product spanning this SNV was cloned for each of the twins separately using the TOPO TA cloning kit as described by the manufacturer (Invitrogen). The resulting bacterial colonies were randomly picked, heat denatured and the insert PCR was amplified using the plasmid-derived forward and reverse primers. The resulting PCR products were subsequently Sanger sequenced.
Copy number analysis.
The ASCAT algorithm43 was used to detect copy number aberrations and loss-of-heterozygosity regions in the tumor genome using Illumina Human-Omni1 SNP arrays.
Validation of somatic missense mutations in the tumor-normal genomes.
Standard Sequenom MassARRAY genotyping experiments were done to validate somatic missense variants in the tumor and matched normal genomes, according to the manufacturer's conditions. Automated genotyping calls were generated using the MassARRAY RTTM software (Sequenom) and were validated by manual review of the raw mass spectra. The following approach was used: variants that failed to be successfully genotyped in the first round of validation were subsequently redesigned for a second attempt using a new set of Sequenom primers (e.g., by designing new extension primers that annealed on the other DNA strand as the extension primer from the first validation round). When variants also failed to be successfully genotyped in this second round, they were considered as 'failed genotyping using Sequenom'. As Tumor 1 was characterized by a high number of infiltrating normal cells, it was possible that Sequenom was not able to positively validate a number of true-positive variants. We therefore also carried out Sanger sequencing on the variants from Tumor 1 that failed to be confirmed with Sequenom in the first validation round. All data from these validation experiments in the tumors (that is, confirmed somatic variants, false-positive and false-negative variants) are accessible in Supplementary Table 1.
Functional annotation of SNVs.
Several computational tools and databases were used to predict the functional effect of coding and noncoding SNVs, including known or predicted protein coding genes, miRNAs and their target sites26, TRANSFAC transcription factor binding sites44, OregANNO annotated regulatory sites45, Vista Enhancer sites46, conservation as indicated by the presence of GERP constraint elements17, phastcons conserved elements47 and repeat elements. The effect of coding mutations was assessed using SIFT48, PolyPhen49 and CanPredict50. Ingenuity Pathway Analysis (IPA) was used to analyze which pathways were affected by the somatic mutations (Supplementary Note 11).
The GenomeComb software tool.
We have made our filtering pipeline freely available to the community under the name GenomeComb. The software tool can be downloaded from http://genomecomb.sourceforge.net/. Filter databases are available for reference genomes NCBI36/hg18 and NCBI37/hg19, and standard filtering protocols are provided on the tool's website. GenomeComb has been developed for Complete Genomics– and GATK-generated variants, but can easily be applied to other platforms through the use of the standard Variant Call Format (VCF). By using the query language provided, users can customize filters and annotations specifically for their research questions. GenomeComb can also be used as a prioritization tool, to rank SNVs based on their probability of being an error. The annotated, tab-separated variant files are the standard output of GenomeComb. A graphical tool, allowing researchers to flexibly select a number of filters and assess how the selected filter combination affects the number of shared and discordant variants relative to the other filter combinations, is also available. Together with the number of shared and discordant SNVs (that is, true SNVs and errors), the 'total' and 'per novel' variant error rates, as well as the percentage of the genome removed for each of the 1,048 filter combinations, are provided.
Accession of sequencing data.
Genome sequence data, all unfiltered variants identified in the twins and tumor-normal pairs and all the annotated variant files have been deposited at the European Genotype Phenotype Archive (http://www.ebi.ac.uk/ega/) under restricted access, with accession numbers EGAS00001000158 (tumor-normal genomes) and EGAS00001000152 (monozygotic twin genomes). The NA19240 sequence data are freely available at ftp://ftp2.completegenomics.com/YRI_trio/ASM_Build37_2.0.0/NA19240/ and ftp://ftp.1000genomes.ebi.ac.uk:21//vol1/ftp/data/NA19240/alignment/), whereas the annotated variant files can be at downloaded at http://genomecomb.sourceforge.net/.
Accession codes
References
Ashley, E.A. et al. Clinical assessment incorporating a personal genome. Lancet 375, 1525–1535 (2010).
Cirulli, E.T. & Goldstein, D.B. Uncovering the roles of rare variants in common disease through whole-genome sequencing. Nat. Rev. Genet. 11, 415–425 (2010).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Anonymous. The sequence is dead: long live the genome. Nat. Biotechnol. 29, 463 (2011).
Lee, W. et al. The mutation spectrum revealed by paired genome sequences from a lung cancer patient. Nature 465, 473–477 (2010).
Pleasance, E.D. et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 463, 184–190 (2010).
Pleasance, E.D. et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463, 191–196 (2010).
Dalgliesh, G.L. et al. Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature 463, 360–363 (2010).
Roach, J.C. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639 (2010).
Ahn, S.M. et al. The first Korean genome sequence and analysis: full genome sequencing for a socio-ethnic group. Genome Res. 19, 1622–1629 (2009).
Baranzini, S.E. et al. Genome, epigenome and RNA sequences of monozygotic twins discordant for multiple sclerosis. Nature 464, 1351–1356 (2010).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Drmanac, R. et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327, 78–81 (2010).
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Fujimoto, A. et al. Whole-genome sequencing and comprehensive variant analysis of a Japanese individual using massively parallel sequencing. Nat. Genet. 42, 931–936 (2010).
Kim, J.I. et al. A highly annotated whole-genome sequence of a Korean individual. Nature 460, 1011–1015 (2009).
Kitzman, J.O. et al. Haplotype-resolved genome sequencing of a Gujarati Indian individual. Nat. Biotechnol. 29, 59–63 (2011).
Ley, T.J. et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456, 66–72 (2008).
Lupski, J.R. et al. Whole-genome sequencing in a patient with Charcot-Marie-Tooth neuropathy. N. Engl. J. Med. 362, 1181–1191 (2010).
McKernan, K.J. et al. Sequence and structural variation in a human genome uncovered by short-read, massively parallel ligation sequencing using two-base encoding. Genome Res. 19, 1527–1541 (2009).
Pelak, K. et al. The characterization of twenty sequenced human genomes. PLoS Genet. 6, e1001111 (2010).
Rasmussen, M. et al. Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature 463, 757–762 (2010).
Schuster, S.C. et al. Complete Khoisan and Bantu genomes from southern Africa. Nature 463, 943–947 (2010).
Tong, P. et al. Sequencing and analysis of an Irish human genome. Genome Biol. 11, R91 (2010).
Wheeler, D.A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008).
Rhead, B. et al. The UCSC Genome Browser database: update 2010. Nucleic Acids Res. 38, D613–D619 (2010).
Siva, N. 1000 Genomes project. Nat. Biotechnol. 26, 256 (2008).
Lynch, M. et al. A genome-wide view of the spectrum of spontaneous mutations in yeast. Proc. Natl. Acad. Sci. USA 105, 9272–9277 (2008).
Haag-Liautard, C. et al. Direct estimation of per nucleotide and genomic deleterious mutation rates in Drosophila. Nature 445, 82–85 (2007).
Baranzini, S.E. et al. Genome, epigenome and RNA sequences of monozygotic twins discordant for multiple sclerosis. Nature 464, 1351–1356 (2010).
Cooper, G.M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005).
Penzkofer, T., Dandekar, T. & Zemojtel, T. L1Base: from functional annotation to prediction of active LINE-1 elements. Nucleic Acids Res. 33, D498–D500 (2005).
Leunen, K. et al. Recurrent copy number alterations in BRCA1-mutated ovarian tumors alter biological pathways. Hum. Mutat. 30, 1693–1702 (2009).
Gorringe, K.L. & Campbell, I.G. Large-scale genomic analysis of ovarian carcinomas. Mol. Oncol. 3, 157–164 (2009).
Futreal, P.A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
The Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Muotri, A.R. et al. L1 retrotransposition in neurons is modulated by MeCP2. Nature 468, 443–446 (2010).
Karlsson, H. et al. Retroviral RNA identified in the cerebrospinal fluids and brains of individuals with schizophrenia. Proc. Natl. Acad. Sci. USA 98, 4634–4639 (2001).
Altshuler, D.M. et al. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Weckx, S. et al. novoSNP, a novel computational tool for sequence variation discovery. Genome Res. 15, 436–442 (2005).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc Natl Acad Sci USA 39, 16910–16915 (2010).
Wingender, E. et al. TRANSFAC: an integrated system for gene expression regulation. Nucleic Acids Res. 28, 316–319 (2000).
Griffith, O.L. et al. ORegAnno: an open-access community-driven resource for regulatory annotation. Nucleic Acids Res. 36, D107–D113 (2008).
Visel, A. et al. VISTA Enhancer Browser–a database of tissue-specific human enhancers. Nucleic Acids Res. 35, D88–D92 (2007).
Felsenstein, J. & Churchill, G.A. A Hidden Markov Model approach to variation among sites in rate of evolution. Mol. Biol. Evol. 13, 93–104 (1996).
Ng, P.C. & Henikoff, S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acid Res. 31, 3812–3814 (2003).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Kaminker, J.S. et al. CanPredict: a computational tool for predicting cancer-associated missense mutations. Nucleic Acids Res. 35, W595–W598 (2007).
Acknowledgements
We appreciate the assistance of M. Veugelers and S. Plaisance (VIB Technology Watch). We acknowledge G. Peuteman, T. Van Brussel, S. Cammaerts, M. Strazisar and the Genetic Service Facility (http://www.vibgeneticservicefacility.be/) for technical assistance. We highly appreciate the helpful comments from the reviewers. The research was supported by the Fund for Scientific Research Flanders (FWO-F) to J.R. and P.V.L., the Agency for Innovation by Science and Technology (IWT) to M.V.D.B., the Stichting tegen Kanker, FWO-F and the KULeuven (KULPFV/10/016-SymBioSysII)) to D.L.
Author information
Authors and Affiliations
Contributions
D.L. and J.D.-F. conceptualized this work. J.R. and P.D.R. wrote algorithms and analyzed data. H.Z. analyzed the Yoruban genome, A.L. assisted with the twin analysis. J.C. and B.H. performed RTG-related analyses. P.V.L. provided the ASCAT algorithm, D.S. performed SNP array experiments. K.C., M.V.D.B., B.S., E.D. and I.V. selected and characterized patient samples. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
B.H. and J.C. are employees of Real Time Genomics and have financial interests in Real Time Genomics.
Supplementary information
Supplementary Text and Figures
Supplementary Notes 1–14 (PDF 5224 kb)
Supplementary Table S1
Validation experiments for the twin and tumor-normal genomes using Sanger sequencing and Sequenom MassARRAY genotyping (XLSX 390 kb)
Supplementary Table S2
Metrics and error rates calculated for each filter combination performed on the twin genome comparison using coverage depth cutoffs of 10 and 20 (XLSX 908 kb)
Supplementary Table S3
Overlap analysis of somatic variants in Tumor 1 and its replicate using three filter settings (XLSX 106 kb)
Supplementary Table S4
Sequenom validation of somatic missense SNVs in the ovarian clear cell tumor genome using three filter settings (XLSX 67 kb)
Supplementary Table S5
Prediction of the effect of the validated somatic mutations and somatic non-coding SNVs in the ovarian serous carcinoma (XLSX 18 kb)
Supplementary Table S6
Effect of filters cumulatively applied to the NA19240 genome, using stringent CG filters versus unfiltered CG data (XLSX 500 kb)
Rights and permissions
About this article
Cite this article
Reumers, J., De Rijk, P., Zhao, H. et al. Optimized filtering reduces the error rate in detecting genomic variants by short-read sequencing. Nat Biotechnol 30, 61–68 (2012). https://doi.org/10.1038/nbt.2053
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2053
This article is cited by
-
Increased prime edit rates in KCNQ2 and SCN1A via single nicking all-in-one plasmids
BMC Biology (2023)
-
Mutated Toll-like receptor 9 increases Alzheimer’s disease risk by compromising innate immunity protection
Molecular Psychiatry (2023)
-
Integrated genetic analyses of immunodeficiency-associated Epstein-Barr virus- (EBV) positive primary CNS lymphomas
Acta Neuropathologica (2023)
-
Genetic mapping, transcriptomic sequencing and metabolic profiling indicated a glutathione S-transferase is responsible for the red-spot-petals in Gossypium arboreum
Theoretical and Applied Genetics (2022)
-
OsWRKY115 on qCT7 links to cold tolerance in rice
Theoretical and Applied Genetics (2022)