Abstract
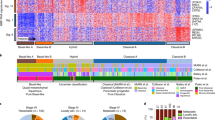
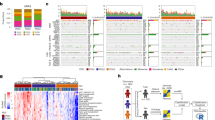
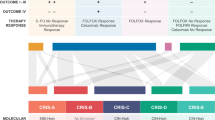
Cancer is often viewed as a caricature of normal developmental processes, but the extent to which its cellular heterogeneity truly recapitulates multilineage differentiation processes of normal tissues remains unknown. Here we implement single-cell PCR gene-expression analysis to dissect the cellular composition of primary human normal colon and colon cancer epithelia. We show that human colon cancer tissues contain distinct cell populations whose transcriptional identities mirror those of the different cellular lineages of normal colon. By creating monoclonal tumor xenografts from injection of a single (n = 1) cell, we demonstrate that the transcriptional diversity of cancer tissues is largely explained by in vivo multilineage differentiation and not only by clonal genetic heterogeneity. Finally, we show that the different gene-expression programs linked to multilineage differentiation are strongly associated with patient survival. We develop two-gene classifier systems (KRT20 versus CA1, MS4A12, CD177, SLC26A3) that predict clinical outcomes with hazard ratios superior to those of pathological grade and comparable to those of microarray-derived multigene expression signatures.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Reya, T., Morrison, S.J., Clarke, M.F. & Weissman, I.L. Stem cells, cancer, and cancer stem cells. Nature 414, 105–111 (2001).
Jordan, C.T., Guzman, M.L. & Noble, M. Cancer stem cells. N. Engl. J. Med. 355, 1253–1261 (2006).
Dalerba, P., Cho, R.W. & Clarke, M.F. Cancer stem cells: models and concepts. Annu. Rev. Med. 58, 267–284 (2007).
Shackleton, M., Quintana, E., Fearon, E.R. & Morrison, S.J. Heterogeneity in cancer: cancer stem cells versus clonal evolution. Cell 138, 822–829 (2009).
Campbell, L.L. & Polyak, K. Breast tumor heterogeneity: cancer stem cells or clonal evolution? Cell Cycle 6, 2332–2338 (2007).
Marusyk, A. & Polyak, K. Tumor heterogeneity: causes and consequences. Biochim. Biophys. Acta 1805, 105–117 (2010).
Kirkland, S.C. Clonal origin of columnar, mucous, and endocrine cell lineages in human colorectal epithelium. Cancer 61, 1359–1363 (1988).
Odoux, C. et al. A stochastic model for cancer stem cell origin in metastatic colon cancer. Cancer Res. 68, 6932–6941 (2008).
Vermeulen, L. et al. Single-cell cloning of colon cancer stem cells reveals a multi-lineage differentiation capacity. Proc. Natl. Acad. Sci. USA 105, 13427–13432 (2008).
Sato, T. et al. Paneth cells constitute the niche for Lgr5 stem cells in intestinal crypts. Nature 469, 415–418 (2011).
Warren, L., Bryder, D., Weissman, I.L. & Quake, S.R. Transcription factor profiling in individual hematopoietic progenitors by digital RT-PCR. Proc. Natl. Acad. Sci. USA 103, 17807–17812 (2006).
Guo, G. et al. Resolution of cell fate decisions revealed by single-cell gene-expression analysis from zygote to blastocyst. Dev. Cell 18, 675–685 (2010).
White, A.K. et al. High-throughput microfluidic single-cell RT-qPCR. Proc. Natl. Acad. Sci. USA 108, 13999–14004 (2011).
Jiao, Y.F., Nakamura, S., Sugai, T., Yamada, N. & Habano, W. Serrated adenoma of the colorectum undergoes a proliferation versus differentiation process: new conceptual interpretation of morphogenesis. Oncology 74, 127–134 (2008).
Wielenga, V.J. et al. Expression of CD44 in Apc and Tcf mutant mice implies regulation by the WNT pathway. Am. J. Pathol. 154, 515–523 (1999).
Prall, F. et al. CD66a (BGP), an adhesion molecule of the carcinoembryonic antigen family, is expressed in epithelium, endothelium, and myeloid cells in a wide range of normal human tissues. J. Histochem. Cytochem. 44, 35–41 (1996).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Becker, L., Huang, Q. & Mashimo, H. Immunostaining of Lgr5, an intestinal stem cell marker, in normal and premalignant human gastrointestinal tissue. Scientific World Journal 8, 1168–1176 (2008).
Merlos-Suarez, A. et al. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell Stem Cell 8, 511–524 (2011).
Sahoo, D., Dill, D.L., Gentles, A.J., Tibshirani, R. & Plevritis, S.K. Boolean implication networks derived from large scale, whole genome microarray datasets. Genome Biol. 9, R157 (2008).
Hoglund, P. et al. Mutations of the down-regulated in adenoma (DRA) gene cause congenital chloride diarrhoea. Nat. Genet. 14, 316–319 (1996).
Fischer, H., Stenling, R., Rubio, C. & Lindblom, A. Differential expression of aquaporin 8 in human colonic epithelial cells and colorectal tumors. BMC Physiol. 1, 1 (2001).
Koslowski, M., Sahin, U., Dhaene, K., Huber, C. & Tureci, O. MS4A12 is a colon-selective store-operated calcium channel promoting malignant cell processes. Cancer Res. 68, 3458–3466 (2008).
Noah, T.K., Kazanjian, A., Whitsett, J. & Shroyer, N.F. SAM pointed domain ETS factor (SPDEF) regulates terminal differentiation and maturation of intestinal goblet cells. Exp. Cell Res. 316, 452–465 (2010).
Gregorieff, A. et al. The ets-domain transcription factor Spdef promotes maturation of goblet and paneth cells in the intestinal epithelium. Gastroenterology 137, 1333–1345 (2009).
van der Flier, L.G. et al. Transcription factor achaete scute-like 2 controls intestinal stem cell fate. Cell 136, 903–912 (2009).
Ezhkova, E. et al. Ezh2 orchestrates gene expression for the stepwise differentiation of tissue-specific stem cells. Cell 136, 1122–1135 (2009).
Park, I.K. et al. Bmi-1 is required for maintenance of adult self-renewing haematopoietic stem cells. Nature 423, 302–305 (2003).
Sangiorgi, E. & Capecchi, M.R. Bmi1 is expressed in vivo in intestinal stem cells. Nat. Genet. 40, 915–920 (2008).
Zeng, Y.A. & Nusse, R. Wnt proteins are self-renewal factors for mammary stem cells and promote their long-term expansion in culture. Cell Stem Cell 6, 568–577 (2010).
Beider, K., Abraham, M. & Peled, A. Chemokines and chemokine receptors in stem cell circulation. Front. Biosci. 13, 6820–6833 (2008).
Jensen, K.B. et al. Lrig1 expression defines a distinct multipotent stem cell population in mammalian epidermis. Cell Stem Cell 4, 427–439 (2009).
Dalla-Favera, R., Wong-Staal, F. & Gallo, R.C. Onc gene amplification in promyelocytic leukaemia cell line HL-60 and primary leukaemic cells of the same patient. Nature 299, 61–63 (1982).
Hoey, T. et al. DLL4 blockade inhibits tumor growth and reduces tumor-initiating cell frequency. Cell Stem Cell 5, 168–177 (2009).
Park, S.Y., Gonen, M., Kim, H.J., Michor, F. & Polyak, K. Cellular and genetic diversity in the progression of in situ human breast carcinomas to an invasive phenotype. J. Clin. Invest. 120, 636–644 (2010).
Losi, L., Baisse, B., Bouzourene, H. & Benhattar, J. Evolution of intratumoral genetic heterogeneity during colorectal cancer progression. Carcinogenesis 26, 916–922 (2005).
Dalerba, P. et al. Phenotypic characterization of human colorectal cancer stem cells. Proc. Natl. Acad. Sci. USA 104, 10158–10163 (2007).
Oien, K.A. Pathologic evaluation of unknown primary cancer. Semin. Oncol. 36, 8–37 (2009).
Lugli, A., Tzankov, A., Zlobec, I. & Terracciano, L.M. Differential diagnostic and functional role of the multi-marker phenotype CDX2/CK20/CK7 in colorectal cancer stratified by mismatch repair status. Mod. Pathol. 21, 1403–1412 (2008).
Sahoo, D., Dill, D.L., Tibshirani, R. & Plevritis, S.K. Extracting binary signals from microarray time-course data. Nucleic Acids Res. 35, 3705–3712 (2007).
Jorissen, R.N. et al. Metastasis-associated gene expression changes predict poor outcomes in patients with dukes stage B and C colorectal cancer. Clin. Cancer Res. 15, 7642–7651 (2009).
Smith, J.J. et al. Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 138, 958–968 (2010).
Guastadisegni, C., Colafranceschi, M., Ottini, L. & Dogliotti, E. Microsatellite instability as a marker of prognosis and response to therapy: a meta-analysis of colorectal cancer survival data. Eur. J. Cancer 46, 2788–2798 (2010).
Bardia, A. et al. Adjuvant chemotherapy for resected stage II and III colon cancer: comparison of two widely used prognostic calculators. Semin. Oncol. 37, 39–46 (2010).
Dalerba, P. et al. Reconstitution of human telomerase reverse transcriptase expression rescues colorectal carcinoma cells from in vitro senescence: evidence against immortality as a constitutive trait of tumor cells. Cancer Res. 65, 2321–2329 (2005).
Ringner, M. What is principal component analysis? Nat. Biotechnol. 26, 303–304 (2008).
O'Doherty, U., Swiggard, W.J. & Malim, M.H. Human immunodeficiency virus type 1 spinoculation enhances infection through virus binding. J. Virol. 74, 10074–10080 (2000).
Wang, G.P. et al. DNA bar coding and pyrosequencing to analyze adverse events in therapeutic gene transfer. Nucleic Acids Res. 36, e49 (2008).
Ishizawa, K. et al. Tumor-initiating cells are rare in many human tumors. Cell Stem Cell 7, 279–282 (2010).
Quintana, E. et al. Efficient tumour formation by single human melanoma cells. Nature 456, 593–598 (2008).
Acknowledgements
This study was supported by National Institutes of Health (NIH) grants U54-CA126524 and P01-CA139490 (to S.R.Q. and M.F.C.), the NIH Director's Pioneer Awards (to S.R.Q.) and a grant from the Ludwig foundation (to M.F.C.). P.D. was supported by a training grant from the California Institute for Regenerative Medicine (CIRM) and by a BD Biosciences Stem Cell Research Grant (Summer 2011). T.K. was supported by a fellowship from the Machiah Foundation. D.S. was supported by NIH grant K99-CA151673, by Department of Defense grant W81XWH-10-1-0500 and a grant from the Siebel Stem Cell Institute and the Thomas and Stacey Siebel Foundation. We wish to thank R. Tibshirani, D. Witten, L. Warren, R.A. White III, E. Gilbert, P. Lovelace, M. Palmor, C. Donkers and S.P. Miranda for helpful discussion and technical support in many moments during the completion of this study.
Author information
Authors and Affiliations
Contributions
P.D., T.K., D.S., M.F.C. and S.R.Q. conceived the study and designed the experiments. P.S.R., M.E.R., A.A.L., M.Z., N.F.N., M.v.d.W. and H.C. provided intellectual guidance in the design of selected experiments. P.D., T.K., D.S., P.S.R., A.A.L., S.S., J.O., D.M.J., D.Q., J.W., and S.H. performed the experiments. P.D., T.K., D.S., N.F.N., Y.S., M.F.C. and S.R.Q. analyzed the data and/or provided intellectual guidance in their interpretation. J.B., A.A.S. and B.V. provided samples and reagents. P.D., T.K., D.S., M.F.C. and S.R.Q. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
S.R.Q. is a founder, consultant and shareholder of Fluidigm. S.R.Q. and M.F.C. are founders, consultants and shareholders of QuantiCell. The authors have filed a patent application based on the results described in this study.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–4, Supplementary Methods and Supplementary Figures 1–24 (PDF 19216 kb)
Rights and permissions
About this article
Cite this article
Dalerba, P., Kalisky, T., Sahoo, D. et al. Single-cell dissection of transcriptional heterogeneity in human colon tumors. Nat Biotechnol 29, 1120–1127 (2011). https://doi.org/10.1038/nbt.2038
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2038
This article is cited by
-
Clinical plasma cells-related genes to aid therapy in colon cancer
BMC Genomics (2023)
-
The DNA glycosylase NEIL2 is protective during SARS-CoV-2 infection
Nature Communications (2023)
-
Single-Cell Sequencing Technology and Its Application in the Study of Central Nervous System Diseases
Cell Biochemistry and Biophysics (2023)
-
Intestinal cellular heterogeneity and disease development revealed by single-cell technology
Cell Regeneration (2022)
-
Semantic clustering analysis of E3-ubiquitin ligases in gastrointestinal tract defines genes ontology clusters with tissue expression patterns
BMC Gastroenterology (2022)