Abstract
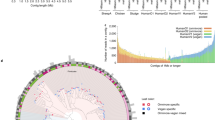
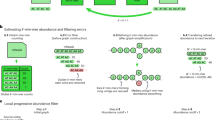
Whole genome amplification by the multiple displacement amplification (MDA) method allows sequencing of DNA from single cells of bacteria that cannot be cultured. Assembling a genome is challenging, however, because MDA generates highly nonuniform coverage of the genome. Here we describe an algorithm tailored for short-read data from single cells that improves assembly through the use of a progressively increasing coverage cutoff. Assembly of reads from single Escherichia coli and Staphylococcus aureus cells captures >91% of genes within contigs, approaching the 95% captured from an assembly based on many E. coli cells. We apply this method to assemble a genome from a single cell of an uncultivated SAR324 clade of Deltaproteobacteria, a cosmopolitan bacterial lineage in the global ocean. Metabolic reconstruction suggests that SAR324 is aerobic, motile and chemotaxic. Our approach enables acquisition of genome assemblies for individual uncultivated bacteria using only short reads, providing cell-specific genetic information absent from metagenomic studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Accessions
BioProject
GenBank/EMBL/DDBJ
NCBI Reference Sequence
Sequence Read Archive
References
Rusch, D.B. et al. The Sorcerer II Global Ocean Sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol. 5, e77 (2007).
Gill, S.R. et al. Metagenomic analysis of the human distal gut microbiome. Science 312, 1355–1359 (2006).
Raghunathan, A. et al. Genomic DNA amplification from a single bacterium. Appl. Environ. Microbiol. 71, 3342–3347 (2005).
Dean, F.B. et al. Comprehensive human genome amplification using multiple displacement amplification. Proc. Natl. Acad. Sci. USA 99, 5261–5266 (2002).
Dean, F.B., Nelson, J.R., Giesler, T.L. & Lasken, R.S. Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 11, 1095–1099 (2001).
Hosono, S. et al. Unbiased whole-genome amplification directly from clinical samples. Genome Res. 13, 954–964 (2003).
Lasken, R.S. Single cell genomic sequencing using Multiple Displacement Amplification. Curr. Opin. Microbiol. 10, 510–516 (2007).
Ishoey, T., Woyke, T., Stepanauskas, R., Novotny, M. & Lasken, R.S. Genomic sequencing of single microbial cells from environmental samples. Curr. Opin. Microbiol. 11, 198–204 (2008).
Zhang, K. et al. Sequencing genomes from single cells by polymerase cloning. Nat. Biotechnol. 24, 680–686 (2006).
Lasken, R.S. & Stockwell, T.B. Mechanism of chimera formation during the Multiple Displacement Amplification reaction. BMC Biotechnol. 7, 19 (2007).
Lasken, R.S. et al. Multiple displacement amplification from single bacterial cells in Whole Genome Amplification: Methods Express (eds. Hughes, S. & Lasken, R.) 119–147 (Scion Publishing Ltd., UK, 2005).
Kvist, T., Ahring, B.K., Lasken, R.S. & Westermann, P. Specific single-cell isolation and genomic amplification of uncultured microorganisms. Appl. Microbiol. Biotechnol. 74, 926–935 (2007).
Mussmann, M. et al. Insights into the genome of large sulfur bacteria revealed by analysis of single filaments. PLoS Biol. 5, e230 (2007).
Marcy, Y. et al. Dissecting biological “dark matter” with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc. Natl. Acad. Sci. USA 104, 11889–11894 (2007).
Podar, M. et al. Targeted access to the genomes of low abundance organisms in complex microbial communities. Appl. Environ. Microbiol. 73, 3205–3214 (2007).
Hongoh, Y. et al. Complete genome of the uncultured Termite Group 1 bacteria in a single host protist cell. Proc. Natl. Acad. Sci. USA 105, 5555–5560 (2008).
Rodrigue, S. et al. Whole genome amplification and de novo assembly of single bacterial cells. PLoS ONE 4, e6864 (2009).
Woyke, T. et al. Assembling the marine metagenome, one cell at a time. PLoS ONE 4, e5299 (2009).
Zerbino, D.R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Pevzner, P.A., Tang, H. & Waterman, M.S. An Eulerian path approach to DNA fragment assembly. Proc. Natl. Acad. Sci. USA 98, 9748–9753 (2001).
Simpson, J.T. et al. ABySS: a parallel assembler for short read sequence data. Genome Res. 19, 1117–1123 (2009).
Chaisson, M.J. & Pevzner, P.A. Short read fragment assembly of bacterial genomes. Genome Res. 18, 324–330 (2008).
Diep, B.A. et al. Complete genome sequence of USA300, an epidemic clone of community-acquired meticillin-resistant Staphylococcus aureus. Lancet 367, 731–739 (2006).
Wright, T.D., Vergin, K.L., Boyd, P.W. & Giovannoni, S.J. A novel delta-subdivision proteobacterial lineage from the lower ocean surface layer. Appl. Environ. Microbiol. 63, 1441–1448 (1997).
Noguchi, H., Park, J. & Takagi, T. MetaGene: prokaryotic gene finding from environmental genome shotgun sequences. Nucleic Acids Res. 34, 5623–5630 (2006).
Tatusov, R.L. et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics 4, 41 (2003).
Goldman, B.S. et al. Evolution of sensory complexity recorded in a myxobacterial genome. Proc. Natl. Acad. Sci. USA 103, 15200–15205 (2006).
DeLong, E.F. et al. Community genomics among stratified microbial assemblages in the ocean's interior. Science 311, 496–503 (2006).
Rich, V.I., Pham, V.D., Eppley, J., Shi, Y. & Delong, E.F. Time-series analyses of Monterey Bay coastal microbial picoplankton using a 'genome proxy' microarray. Environ. Microbiol. 13, 116–134 (2010).
Yooseph, S. et al. Genomic and functional adaptation in surface ocean planktonic prokaryotes. Nature 468, 60–66 (2010).
Iizuka, T. et al. Plesiocystis pacifica gen. nov., sp. nov., a marine myxobacterium that contains dihydrogenated menaquinone, isolated from the Pacific coasts of Japan. Int. J. Syst. Evol. Microbiol. 53, 189–195 (2003).
Callister, S.J. et al. Comparative bacterial proteomics: analysis of the core genome concept. PLoS ONE 3, e1542 (2008).
Mitreva, M. Bacterial core gene set. <http://www.hmpdacc.org/doc/sops/reference_genomes/metrics/Bacterial_CoreGenes_SOP.pdf> (2008).
Nelson, K.E. et al. A catalog of reference genomes from the human microbiome. Science 328, 994–999 (2010).
Woyke, T. et al. One bacterial cell, one complete genome. PLoS ONE 5, e10314 (2010).
King, G.M. Microbial carbon monoxide consumption in salt marsh sediments. FEMS Microbiol. Ecol. 59, 2–9 (2007).
Schloss, P.D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Wilgenbusch, J.C. & Swofford, D. Inferring evolutionary trees with PAUP*. Curr. Prot. Bioinformatics, Unit 6.4 6.4.1–6.4.28 (2003).
Hernandez, D., Francois, P., Farinelli, L., Ostera, M. & Schrenzel, J. De novo bacterial genome sequencing: Millions of very short reads assembled on a desktop computer. Genome Res. 18, 802–809 (2008).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Mao, F., Dam, P., Chou, J., Olman, V. & Xu, Y. DOOR: a database for prokaryotic operons. Nucleic Acids Res. 37, D459–D463 (2009).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Tanenbaum, D.M. et al. The JCVI standard operating procedure for annotating prokaryotic metagenomic shotgun sequencing data. Stand. Genomic Sci. 2, 229–237 (2010).
Ramirez-Flandes, S. & Ulloa, O. Bosque: integrated phylogenetic analysis software. Bioinformatics 24, 2539–2541 (2008).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A.C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, W182–185 (2007).
Acknowledgements
This work was partially supported by grants to R.S.L. from the National Human Genome Research Institute (NIH-2 R01 HG003647) and the Alfred P. Sloan Foundation (Sloan Foundation-2007-10-19), and by a grant to P.A.P. and G.T. from the US National Institutes of Health (NIH grant 3P41RR024851-02S1). We thank M. Kim (J. Craig Venter Institute) for bioinformatics support.
Author information
Authors and Affiliations
Contributions
All authors analyzed data. H.C. and G.T. wrote software. M.N., J.L.Y.-G., M.-J.L. and L.J.F. performed wet lab experiments. Illumina sequencing was performed at Illumina Cambridge Ltd. O.S.-T. analyzed sequencing data at Illumina. H.C., J.L.Y.-G., G.T., C.L.D., M.-J.L., L.J.F., N.A.G., P.A.P. and R.S.L. wrote the manuscript. H.C., G.T., M.-J.L., C.L.D., J.H.B., D.B.R. and N.A.G. created figures and tables. R.S.L. and M.-J.L. supervised the JCVI group. P.A.P. and G.T. supervised the UCSD group. N.A.G. and D.J.E. supervised the Illumina group. G.P.S. initiated the Illumina-JCVI collaboration.
Corresponding author
Ethics declarations
Competing interests
L.J.F., N.A.G., O.S.-T., G.P.S. and D.J.E. are employees of Illumina, the commercial source of Illumina sequencing, which is evaluated in this manuscript.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–5, Supplementary Methods, Supplementary Data 3 and Supplementary Figures 1–13 (PDF 2029 kb)
Supplementary Data 1
Velvet-SC source code (TGZ 4047 kb)
Supplementary Data 2
EULER-SR Error correction source code (TGZ 129 kb)
Rights and permissions
About this article
Cite this article
Chitsaz, H., Yee-Greenbaum, J., Tesler, G. et al. Efficient de novo assembly of single-cell bacterial genomes from short-read data sets. Nat Biotechnol 29, 915–921 (2011). https://doi.org/10.1038/nbt.1966
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1966
This article is cited by
-
Mitogenome-wise codon usage pattern from comparative analysis of the first mitogenome of Blepharipa sp. (Muga uzifly) with other Oestroid flies
Scientific Reports (2022)
-
Metapangenomics reveals depth-dependent shifts in metabolic potential for the ubiquitous marine bacterial SAR324 lineage
Microbiome (2021)
-
Using single-cell sequencing technology to detect circulating tumor cells in solid tumors
Molecular Cancer (2021)
-
Functional genomics by integrated analysis of transcriptome of sweet potato (Ipomoea batatas (L.) Lam.) during root formation
Genes & Genomics (2020)
-
Studying the gut virome in the metagenomic era: challenges and perspectives
BMC Biology (2019)