Abstract
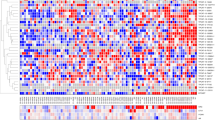
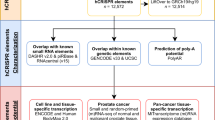
Noncoding RNAs (ncRNAs) are emerging as key molecules in human cancer, with the potential to serve as novel markers of disease and to reveal uncharacterized aspects of tumor biology. Here we discover 121 unannotated prostate cancer–associated ncRNA transcripts (PCATs) by ab initio assembly of high-throughput sequencing of polyA+ RNA (RNA-Seq) from a cohort of 102 prostate tissues and cells lines. We characterized one ncRNA, PCAT-1, as a prostate-specific regulator of cell proliferation and show that it is a target of the Polycomb Repressive Complex 2 (PRC2). We further found that patterns of PCAT-1 and PRC2 expression stratified patient tissues into molecular subtypes distinguished by expression signatures of PCAT-1–repressed target genes. Taken together, our findings suggest that PCAT-1 is a transcriptional repressor implicated in a subset of prostate cancer patients. These findings establish the utility of RNA-Seq to identify disease-associated ncRNAs that may improve the stratification of cancer subtypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Metzker, M.L. Sequencing technologies—the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Guttman, M. et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat. Biotechnol. 28, 503–510 (2010).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Robertson, G. et al. De novo assembly and analysis of RNA-seq data. Nat. Methods 7, 909–912 (2010).
Zerbino, D.R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010).
Orom, U.A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
Rinn, J.L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Gupta, R.A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Pasmant, E. et al. Characterization of a germ-line deletion, including the entire INK4/ARF locus, in a melanoma-neural system tumor family: identification of ANRIL, an antisense noncoding RNA whose expression coclusters with ARF. Cancer Res. 67, 3963–3969 (2007).
Yap, K.L. et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol. Cell 38, 662–674 (2010).
Tsai, M.C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010).
Kotake, Y. et al. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene 30, 1956–1962 (2011).
de Kok, J.B. et al. DD3(PCA3), a very sensitive and specific marker to detect prostate tumors. Cancer Res. 62, 2695–2698 (2002).
Li, J. et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275, 1943–1947 (1997).
Prensner, J.R. & Chinnaiyan, A.M. Oncogenic gene fusions in epithelial carcinomas. Curr. Opin. Genet. Dev. 19, 82–91 (2009).
Tomlins, S.A. et al. Distinct classes of chromosomal rearrangements create oncogenic ETS gene fusions in prostate cancer. Nature 448, 595–599 (2007).
Tomlins, S.A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Thierry-Mieg, D. & Thierry-Mieg, J. AceView: a comprehensive cDNA-supported gene and transcripts annotation. Genome Biol. 7 (suppl. 1), S11–S14 (2006).
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
The FANTOM Consortium. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
He, Y., Vogelstein, B., Velculescu, V.E., Papadopoulos, N. & Kinzler, K.W. The antisense transcriptomes of human cells. Science 322, 1855–1857 (2008).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Yu, J. et al. An integrated network of androgen receptor, polycomb, and TMPRSS2-ERG gene fusions in prostate cancer progression. Cancer Cell 17, 443–454 (2010).
Day, D.S., Luquette, L.J., Park, P.J. & Kharchenko, P.V. Estimating enrichment of repetitive elements from high-throughput sequence data. Genome Biol. 11, R69 (2010).
Kim, T.K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Rubin, M.A. et al. Alpha-methylacyl coenzyme A racemase as a tissue biomarker for prostate cancer. J. Am. Med. Assoc. 287, 1662–1670 (2002).
Dhanasekaran, S.M. et al. Delineation of prognostic biomarkers in prostate cancer. Nature 412, 822–826 (2001).
van Bakel, H., Nislow, C., Blencowe, B.J. & Hughes, T.R. Most “dark matter” transcripts are associated with known genes. PLoS Biol. 8, e1000371 (2010).
Tomlins, S.A. et al. The role of SPINK1 in ETS rearrangement-negative prostate cancers. Cancer Cell 13, 519–528 (2008).
Bjartell, A.S. et al. Association of cysteine-rich secretory protein 3 and beta-microseminoprotein with outcome after radical prostatectomy. Clin. Cancer Res. 13, 4130–4138 (2007).
Oosumi, T., Belknap, W.R. & Garlick, B. Mariner transposons in humans. Nature 378, 672 (1995).
Robertson, H.M., Zumpano, K.L., Lohe, A.R. & Hartl, D.L. Reconstructing the ancient mariners of humans. Nat. Genet. 12, 360–361 (1996).
Kleer, C.G. et al. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc. Natl. Acad. Sci. USA 100, 11606–11611 (2003).
Varambally, S. et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 419, 624–629 (2002).
Ahmadiyeh, N. et al. 8q24 prostate, breast, and colon cancer risk loci show tissue-specific long-range interaction with MYC. Proc. Natl. Acad. Sci. USA 107, 9742–9746 (2010).
Al Olama, A.A. et al. Multiple loci on 8q24 associated with prostate cancer susceptibility. Nat. Genet. 41, 1058–1060 (2009).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Gudmundsson, J. et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat. Genet. 39, 631–637 (2007).
Sotelo, J. et al. Long-range enhancers on 8q24 regulate c-Myc. Proc. Natl. Acad. Sci. USA 107, 3001–3005 (2010).
Taylor, B.S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
Huttenhower, C. et al. Exploring the human genome with functional maps. Genome Res. 19, 1093–1106 (2009).
Laxman, B. et al. A first-generation multiplex biomarker analysis of urine for the early detection of prostate cancer. Cancer Res. 68, 645–649 (2008).
Hessels, D. et al. DD3(PCA3)-based molecular urine analysis for the diagnosis of prostate cancer. Eur. Urol. 44, 8–16 (2003).
Maher, C.A. et al. Chimeric transcript discovery by paired-end transcriptome sequencing. Proc. Natl. Acad. Sci. USA 106, 12353–12358 (2009).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Dennis, G. Jr. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 4, 3 (2003).
Acknowledgements
We thank K. Ramnarayanan and R. Morey for technical assistance with next generation sequencing. We thank R.J. Lonigro, S. Kaylana-Sundaram, T. Barrette, and M. Quist for help with sequencing data analysis, and R. Mehra, B. Han and K. Suleman for prostate tissue specimens. We thank C. Trapnell and G. Pertea for assistance with computational analyses. We thank S. Tomlins, Y.-M. Wu, S. Roychowdhury and members of the Chinnaiyan laboratory for advice and discussions. We thank R. Beroukhim for guidance. This work was supported in part by the US National Institutes of Health (NIH) Prostate Specialized Program of Research Excellence grant P50CA69568, the Early Detection Research Network grant UO1 CA111275 (to A.M.C.), the NIH R01CA132874-01A1 (to A.M.C.), the Department of Defense grant W81XWH-10-0652 and W81XWH-11-1-0337 (to A.M.C.) and the National Center for Functional Genomics supported by the Department of Defense (to A.M.C.). A.M.C. is supported by the Doris Duke Charitable Foundation Clinical Scientist Award, a Burroughs Wellcome Foundation Award in Clinical Translational Research and the Prostate Cancer Foundation. A.M.C. is an American Cancer Society Research Professor. N.P. was supported by a University of Michigan Prostate SPORE Career Development Award. C.A.M. was supported by the American Association of Cancer Research Amgen Fellowship in Clinical/Translational Research, the Canary Foundation and American Cancer Society Early Detection Postdoctoral Fellowship, and a Prostate Cancer Foundation Young Investigator Award. Q.C. was supported by a Department of Defense Postdoctoral Fellowship grant PC094725. J.R.P. was supported by the NIH Cancer Biology Training grant CA009676-18 and the Department of Defense Predoctoral Fellowship W81XWH-10-1-0551. M.K.I. was supported by the Department of Defense Predoctoral Fellowship W81XWH-11-1-0136. J.R.P. and M.K.I. are Fellows of the University of Michigan Medical Scientist Training Program.
Author information
Authors and Affiliations
Contributions
M.K.I., J.R.P. and A.M.C. designed the project and directed experimental studies. M.K.I., O.A.B., C.S.G. and C.A.M. developed computational platforms and performed sequencing data analysis. M.K.I., O.A.B. and H.K.I. performed statistical analyses. J.R.P., S.M.D., J.C.B., Q.C., N.P., H.D.K., B.L., X.W., I.A.A., X.C., X.J. and D.R. performed experimental studies. J.S. and J.T.W. coordinated biospecimens. M.K.I., J.R.P. and A.M.C. interpreted data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The University of Michigan has filed for a patent on the detection of gene fusions in prostate cancer, on which A.M.C. is a co-inventor. The diagnostic field of use for ETS gene fusions has been licensed to GenProbe Inc. The University of Michigan has a sponsored research agreement with GenProbe, which is unrelated to this study. GenProbe has had no role in the design or experimentation of this study, nor has it participated in the writing of the manuscript.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–11, Supplementary Figures 1–28, Supplementary Methods and Supplementary Discussion (PDF 9445 kb)
Rights and permissions
About this article
Cite this article
Prensner, J., Iyer, M., Balbin, O. et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat Biotechnol 29, 742–749 (2011). https://doi.org/10.1038/nbt.1914
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1914
This article is cited by
-
Long non-coding RNA ENST00000503625 is a potential prognostic biomarker and metastasis suppressor gene in prostate cancer
Journal of Cancer Research and Clinical Oncology (2023)
-
Crosstalk between long noncoding RNA and microRNA in Cancer
Cellular Oncology (2023)
-
Role of different non-coding RNAs as ovarian cancer biomarkers
Journal of Ovarian Research (2022)
-
CircSCAF8 promotes growth and metastasis of prostate cancer through the circSCAF8-miR-140-3p/miR-335-LIF pathway
Cell Death & Disease (2022)
-
miR-32 promotes MYC-driven prostate cancer
Oncogenesis (2022)