Abstract
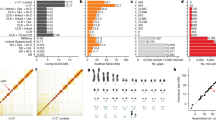
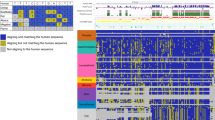
Multiple sequence alignment is a difficult computational problem. There have been compelling pleas for methods to assess whole-genome multiple sequence alignments and compare the alignments produced by different tools. We assess the four ENCODE alignments, each of which aligns 28 vertebrates on 554 Mbp of total input sequence. We measure the level of agreement among the alignments and compare their coverage and accuracy. We find a disturbing lack of agreement among the alignments not only in species distant from human, but even in mouse, a well-studied model organism. Overall, the assessment shows that Pecan produces the most accurate or nearly most accurate alignment in all species and genomic location categories, while still providing coverage comparable to or better than that of the other alignments in the placental mammals. Our assessment reveals that constructing accurate whole-genome multiple sequence alignments remains a significant challenge, particularly for noncoding regions and distantly related species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kent, W. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Woolfe, A. et al. Highly conserved non-coding sequences are associated with vertebrate development. PLoS Biol. 3, e7 (2005).
Xie, X. et al. Systematic discovery of regulatory motifs in conserved regions of the human genome, including thousands of CTCF insulator sites. Proc. Natl. Acad. Sci. USA 104, 7145–7150 (2007).
Gross, S.S. & Brent, M.R. Using multiple alignments to improve gene prediction. J. Comput. Biol. 13, 379–393 (2006).
Siepel, A. et al. Targeted discovery of novel human exons by comparative genomics. Genome Res. 17, 1763–1773 (2007).
Pedersen, J.S. et al. Identification and classification of conserved RNA secondary structures in the human genome. PLOS Comput. Biol. 2, e33 (2006).
Washietl, S., Hofacker, I.L., Lukasser, M., Hüttenhofer, A. & Stadler, P.F. Mapping of conserved RNA secondary structures predicts thousands of functional noncoding RNAs in the human genome. Nat. Biotechnol. 23, 1383–1390 (2005).
Cooper, G.M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005).
Margulies, E. et al. Identification and characterization of multi-species conserved sequences. Genome Res. 13, 2507–2518 (2003).
Margulies, E.H. et al. Analyses of deep mammalian sequence alignments and constraint predictions for 1% of the human genome. Genome Res. 17, 760–774 (2007).
Pollard, K.S., Hubisz, M.J., Rosenbloom, K.R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010).
Prabhakar, S. et al. Close sequence comparisons are sufficient to identify human cis-regulatory elements. Genome Res. 16, 855–863 (2006).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Felsenstein, J. Inferring Phylogenies (Sinauer Associates, 2004).
Wong, K.M., Suchard, M.A. & Huelsenbeck, J.P. Alignment uncertainty and genomic analysis. Science 319, 473–476 (2008).
Siepel, A. & Haussler, D. Phylogenetic estimation of context-dependent substitution rates by maximum likelihood. Mol. Biol. Evol. 21, 468–488 (2004).
Murphy, W.J. et al. Resolution of the early placental mammal radiation using Bayesian phylogenetics. Science 294, 2348–2351 (2001).
Nikolaev, S. et al. Early history of mammals is elucidated with the ENCODE multiple species sequencing data. PLoS Genet. 3, e2 (2007).
Bird, C.P. et al. Fast-evolving noncoding sequences in the human genome. Genome Biol. 8, R118 (2007).
Kim, S. & Pritchard, J. Adaptive evolution of conserved non-coding elements in mammals. PLoS Genet. 3, e147 (2007).
Nielsen, R. et al. A scan for positively selected genes in the genomes of humans and chimpanzees. PLoS Biol. 3, e170 (2005).
Pollard, K.S. et al. An RNA gene expressed during cortical development evolved rapidly in humans. Nature 443, 167–172 (2006).
Prabhakar, S., Noonan, J.P., Pääbo, S. & Rubin, E.M. Accelerated evolution of conserved noncoding sequences in humans. Science 314, 786 (2006).
Dewey, C.N., Huggins, P.M., Woods, K., Sturmfels, B. & Pachter, L. Parametric alignment of Drosophila genomes. PLOS Comput. Biol. 2, e73 (2006).
Blanchette, M. Computation and analysis of genomic multi-sequence alignments. Annu. Rev. Genomics Hum. Genet. 8, 193–213 (2007).
Kumar, S. & Filipski, A. Multiple sequence alignment: in pursuit of homologous DNA positions. Genome Res. 17, 127–135 (2007).
Lunter, G. et al. Uncertainty in homology inferences: assessing and improving genomic sequence alignment. Genome Res. 18, 298–309 (2008).
Margulies, E.H. Confidence in comparative genomics. Genome Res. 18, 199–200 (2008).
Margulies, E.H. & Birney, E. Approaches to comparative sequence analysis: towards a functional view of vertebrate genomes. Nat. Rev. Genet. 9, 303–313 (2008).
Rokas, A. Lining up to avoid bias. Science 319, 416–417 (2008).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Bray, N. & Pachter, L. MAVID: constrained ancestral alignment of multiple sequences. Genome Res. 14, 693–699 (2004).
Brudno, M. et al. LAGAN and Multi-LAGAN: efficient tools for large-scale multiple alignment of genomic DNA. Genome Res. 13, 721–731 (2003).
Paten, B., Herrero, J., Beal, K., Fitzgerald, S. & Birney, E. Enredo and Pecan: genome-wide mammalian consistency-based multiple alignment with paralogs. Genome Res. 18, 1814–1828 (2008).
Prakash, A. & Tompa, M. Measuring the accuracy of genome-size multiple alignments. Genome Biol. 8, R124 (2007).
Dubchak, I., Poliakov, A., Kislyuk, A. & Brudno, M. Multiple whole-genome alignments without a reference organism. Genome Res. 19, 682–689 (2009).
Prakash, A. & Tompa, M. Assessing the discordance of multiple sequence alignments. IEEE/ACM Trans. Comput. Biol. Bioinformatics 6, 542–551 (2009).
Wang, L. & Jiang, T. On the complexity of multiple sequence alignment. J. Comput. Biol. 1, 337–348 (1994).
Karlin, S. & Altschul, S.F. Applications and statistics for multiple high-scoring segments in molecular sequences. Proc. Natl. Acad. Sci. USA 90, 5873–5877 (1993).
States, D.J., Gish, W. & Altschul, S.F. Improved sensitivity in nucleic acid database searches using application-specific scoring matrices. Methods: A Companion to Methods in Enzymology 3, 66–70 (1991).
Acknowledgements
We thank P. Green, W. Noble, W.L. Ruzzo and especially A. Prakash for helpful discussions and technical advice. We thank the US National Institutes of Health and the Natural Sciences and Engineering Research Council of Canada for financial support.
Author information
Authors and Affiliations
Contributions
X.C., design, implementation, experimentation, analysis; M.T., design, analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Text and Supplementary Figs. 1–4 (PDF 1068 kb)
Rights and permissions
About this article
Cite this article
Chen, X., Tompa, M. Comparative assessment of methods for aligning multiple genome sequences. Nat Biotechnol 28, 567–572 (2010). https://doi.org/10.1038/nbt.1637
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1637
This article is cited by
-
Inferring synteny between genome assemblies: a systematic evaluation
BMC Bioinformatics (2018)
-
Testing robustness of relative complexity measure method constructing robust phylogenetic trees for Galanthus L. Using the relative complexity measure
BMC Bioinformatics (2013)
-
Novel algorithm for phylogenetic analysis of proteins: application to analysis of the evolution of H5N1 influenza viruses
Journal of Mathematical Chemistry (2013)
-
Thermally assisted quantum annealing of a 16-qubit problem
Nature Communications (2013)
-
Rigorous and thorough bioinformatic analyses of olfactory receptor promoters confirm enrichment of O/E and homeodomain binding sites but reveal no new common motifs
BMC Genomics (2011)