Abstract
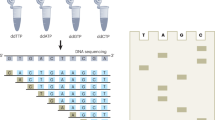
DNA sequencing-by-synthesis (SBS) technology, using a polymerase or ligase enzyme as its core biochemistry, has already been incorporated in several second-generation DNA sequencing systems with significant performance. Notwithstanding the substantial success of these SBS platforms, challenges continue to limit the ability to reduce the cost of sequencing a human genome to $100,000 or less. Achieving dramatically reduced cost with enhanced throughput and quality will require the seamless integration of scientific and technological effort across disciplines within biochemistry, chemistry, physics and engineering. The challenges include sample preparation, surface chemistry, fluorescent labels, optimizing the enzyme-substrate system, optics, instrumentation, understanding tradeoffs of throughput versus accuracy, and read-length/phasing limitations. By framing these challenges in a manner accessible to a broad community of scientists and engineers, we hope to solicit input from the broader research community on means of accelerating the advancement of genome sequencing technology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schloss, J. How to get genomes at one ten-thousandth the cost. Nat. Biotechnol. 26, 1113–1115 (2008).
Dohm, J.C., Lottaz, C., Borodina, T. & Himmelbauer, H. Substantial biases in ultra-short read data sets from high-throughput DNA sequencing. Nucleic Acids Res. 36, e105, 2008.
Craig, D.W. et al. Identification of genetic variants using bar-coded multiplexed sequencing. Nat. Methods 5, 887–893 (2008).
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008).
Morozova, O. & Marra, M.A. Applications of next-generation sequencing technologies in functional genomics. Genomics 92, 255–264 (2008).
Mardis, E. Next-generation DNA sequencing methods. Annu. Rev. Genomics Hum. Genet. 9, 387–402 (2008).
Mardis, E.R. The impact of next-generation sequencing technology on genetics. Trends Genet. 24, 133–141 (2008).
Chi, K.R. The year of sequencing. Nat. Methods 5, 11–14 (2008).
Marguerat, S., Wilhelm, B.T. & Bähler, J. Next-generation sequencing: applications beyond genomes. Biochem. Soc. Trans. 36, 1091–1096 (2008).
Smith, D.R. et al. Rapid whole-genome mutational profiling using next-generation sequencing technologies. Genome Res. 18, 1638–1642 (2008).
Quinn, N.L. et al. Assessing the feasibility of GS FLX Pyrosequencing for sequencing the Atlantic salmon genome. BMC Genomics 9, 404 (2008).
Sarin, S. et al. Caenorhabditis elegans mutant allele identification by whole-genome sequencing. Nat. Methods 5, 865–867 (2008).
Holt, R.A. & Jones, S.J.M. The new paradigm of flow cell sequencing. Genome Res. 18, 839–846 (2008).
Rothberg, J.M. & Leamon, J.H. The development and impact of 454 sequencing. Nat. Biotechnol. 26, 1117–1124 (2008).
Kahvejian, A., Quackenbush, J. & Thompson, J.F. What would you do if you could sequence everything? Nat. Biotechnol. 26, 1125–1133 (2008).
Shendure, J. & Hanlee, J. Next-generation DNA sequencing. Nat. Biotechnol. 26, 1135–1145 (2008).
Branton, D. et al. Nanopore sequencing. Nat. Biotechnol. 26, 1146–1153 (2008).
Levene, M.J. et al. Zero-mode waveguides for single-molecule analysis at high concentrations. Science 299, 682–686 (2003).
Korlach, J. et al. Selective aluminum passivation for targeted immobilization of single DNA polymerase molecules in zero-mode waveguide nanostructures. Proc. Natl. Acad. Sci. USA 105, 1176–1181 (2008).
Williams, J. System and method for nucleic acid sequencing by polymerase synthesis. US patent application US20030194740 (2003).
Willams, J. & Anderson, J. Field-switch sequencing. US patent application US20050266456 (2005).
Hardin, S., Gao, X., Briggs, J., Willson, R. & Tu, S.C. Real-time sequence determination. US patent application US20030064366 (2003).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Harris, T.D. et al. Single-molecule DNA sequencing of a viral genome. Science 320, 106–109 (2008).
Milton, J. et al. Modified nucleotides (for polynucleotide sequencing). World and US patent application WO2004/018497, US2007/0166705 (2004).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Turcatti, G., Romieu, A., Fedurco, M. & Tairi A.P. A new class of cleavable fluorescent nucleotides: synthesis and optimization as reversible terminators for DNA sequencing by synthesis. Nucleic Acids Res. 36, e25 (2008).
Guo, J. et al. Four-color DNA sequencing with 3′-O-modified nucleotide reversible terminators and chemically cleavable fluorescent dideoxynucleotides. Proc. Natl. Acad. Sci. USA 105, 9145–9150 (2008).
Ju, J. et al. Four-color DNA sequencing by synthesis using cleavable fluorescent nucleotide reversible terminators. Proc. Natl. Acad. Sci. USA 103, 19635–19640 (2006).
Seo, T.S. et al. Four-color DNA sequencing by synthesis on a chip using photocleavable fluorescent nucleotides. Proc. Natl. Acad. Sci. USA 102, 5926–5931 (2005).
Wu, W. et al. Termination of DNA synthesis by N6-alkylated, not 3′-O-alkylated, photocleavable 2′-deoxyadenosine triphosphates. Nucleic Acids Res. 35, 6339–6349 (2007).
McKernan, K., Blanchard, A., Kotler, L. & Costa, G. Reagents, methods and libraries for bead-based sequencing. PCT patent application WO2006084132 (2007).
Shendure, J. et al. Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732 (2005).
Fuller, C.W. & Nelson, J.R. Method for nucleic acid analysis. PCT patent application WO2005123957 (2005).
Fuller, C.W. Rapid parallel nucleic acid analysis. US patent application US20060051807 (2006).
Ronaghi, M., Uhlén, M. & Nyrén, P. A sequencing method based on real-time pyrophosphate. Science 281, 363–365 (1998).
Scheibye-Alsing, K. et al. Sequence assembly. Comput. Biol. Chem. 33, 121–136 (2009).
Trapnell, C. & Salzberg, S.L. How to map billions of short reads onto genomes. Nat. Biotechnol. 27, 455–457 (2009).
Chetverina, H.V. & Chetverin, A.B. Cloning of RNA molecules in vitro. Nucleic Acids Res. 21, 2349–2353 (1993).
Church, G.M. Replica amplification of nucleic acid arrays. US patent 6,432,360 (2002).
Dressman, D., Yan, H., Traverso, G., Kinzler, K.W. & Vogelstein, B. Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc. Natl. Acad. Sci. USA 100, 8817–8822 (2003).
Embleton, M.J., Gorochov, G., Jones, P.T. & Winter, G.P. In situ recombinant PCR within single cells US Patent 5,830,663 (1998).
Griffiths, A. & Tawfik, D. In vitro sorting method. US patent 6,489,103 (2002).
Holliger, P. & Ghadessy, F. Emulsion compositions. US patent 7,429,467 (2008).
Brenner, S. et al. In vitro cloning of complex mixtures of DNA on microbeads: physical separation of differentially expressed cDNAs. Proc. Natl. Acad. Sci. USA 97, 1665–1670 (2000).
Adessi, C., Kawashima, E., Mayer, P., Mermod, J.J. & Turcatti, G. Methods of nucleic acid amplification and sequencing. PCT patent application WO2000018957 (2000).
Boles, T.C., Kron, S.J. & Adams, C.P. Nucleic acid-containing polymerizable complex. US patent 5,932,711 (1999).
Zhang, K. et al. Long-range polony haplotyping of individual human chromosome molecules. Nat. Genet. 38, 382–387 (2006).
Zhang, K. et al. Sequencing genomes from single cells via polymerase clones. Nat. Biotechnol. 24, 680–686 (2006).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Li, M., Diehl, F., Dressman, D., Vogelstein, B. & Kinzler, K.W. BEAMing up for detection and quantification of rare sequence variants. Nat. Methods 3, 95–97 (2006).
Pihlak, A. et al. Rapid genome sequencing with short universal tiling probes. Nat. Biotechnol. 26, 676–684 (2008).
Liu, D., Daubendiek, S.L., Zillman, M.A., Ryan, K. & Kool, E.T. Rolling circle DNA synthesis: small circular oligonucleotides as efficient templates for DNA polymerases. J. Am. Chem. Soc. 118, 1587–1594 (1996).
Baner, J., Nilsson, M., Mendel-Hartvig, M. & Landegren, U. Signal amplification of padlock probes by rolling circle replication. Nucleic Acids Res. 26, 5073–5078 (1998).
Fire, A. & Xu, S.Q. Rolling replication of short DNA circles. Proc. Natl. Acad. Sci. USA 92, 4641–4645 (1995).
Blanco, L. et al. Highly efficient DNA synthesis by the phage phi 29 DNA polymerase. Symmetrical mode of DNA replication. J. Biol. Chem. 264, 8935–8940 (1989).
Dean, F.B., Nelson, J.R., Giesler, T.L. & Lasken, R.S. Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 11, 1095–1099 (2001).
Lizardi, P.M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 19, 225–232 (1998).
Ramanathan, A. et al. An integrative approach for the optical sequencing of single DNA molecules. Anal. Biochem. 330, 227–241 (2004).
Parameswaran, P. et al. A pyrosequencing-tailored nucleotide barcode design unveils opportunities for large-scale sample multiplexing. Nucleic Acids Res. 35, e130 (2007).
Albert, T.J. et al. Direct selection of human genomic loci by microarray hybridization. Nat. Methods 4, 903–905 (2007).
Porreca, G.J. et al. Multiplex amplification of large sets of human exons. Nat. Methods 4, 931–936 (2007).
Kim, J.B. et al. Polony multiplex analysis of gene expression (PMAGE) in mouse hypertrophic cardiomyopathy. Science 316, 1481–1484 (2007).
Williams, J.G. et al. An artificial processivity clamp made with streptavidin facilitates oriented attachment of polymerase-DNA complexes to surfaces. Nucl. Acids Res. 36, e121 (2008).
Barbee, K.D. & Huang, X. Magnetic assembly of high-density DNA arrays for genomic analyses. Anal. Chem. 80, 2149–2154 (2008).
Brenner, S. Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nat. Biotechnol. 18, 630–634 (2000).
Mujumdar, R.B., Ernst, L.A., Mujumdar, S.R. & Waggoner, A.S. Cyanine dye labeling reagents containing isothiocyanate groups. Cytometry 10, 11–19 (1989).
Sood, A. et al. Terminal phosphate-labeled nucleotides with improved substrate properties for homogeneous nucleic acid assays. J. Am. Chem. Soc. 127, 2394–2395 (2005).
Prober, J.M. et al. A system for rapid DNA sequencing with fluorescent chain-terminating dideoxynucleotides. Science 238, 336–341 (1987).
Langer, P.R., Waldrop, A.A. & Ward, D.C. Enzymatic synthesis of biotin-labeled polynucleotides: novel nucleic acid affinity probes. Proc. Natl. Acad. Sci. USA 78, 6633–6637 (1981).
Wu, W. et al. Termination of DNA synthesis by N6-alkylated, not 3′-O-alkylated, photocleavable 2′-deoxyadenosine triphosphates. Nucleic Acids Res. 35, 6339–6349 (2007).
Canard, B. & Sarfati, R.S. DNA polymerase fluorescent substrates with reversible 3′-tags. Gene 148, 1–6 (1994).
Tabor, S. & Richardson, C.C. A single residue in DNA polymerases of the Escherichia coli DNA polymerase I family is critical for distinguishing between deoxy- and dideoxyribonucleotides. Proc. Natl. Acad. Sci. USA 92, 6339–6343 (1995).
Doublié, S., Tabor, S., Long, A.M., Richardson, C.C. & Ellenberger, T. Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 Å resolution. Nature 391, 251–258 (1998).
Kumar, S. et al. Terminal phosphate labeled nucleotides: synthesis, applications, and linker effect on incorporation by DNA polymerases. Nucleosides Nucleotides Nucleic Acids 24, 401–408 (2005).
Mulder, B.A. et al. Nucleotide modification at the gamma-phosphate leads to the improved fidelity of HIV-1 reverse transcriptase. Nucleic Acids Res. 33, 4865–4873 (2005).
Williams, J.G.K., Anderson, J.P., Urlacher, T.M. & Steffens, D.L. Mutant polymerases for sequencing and genotyping. US patent application US20070048748 (2007).
Williams, J.G.K. Polymerases with charge-switch activity and methods of generating such polymers. US published patent application US20040259082 (2004).
Korlach, J. et al. Long, processive enzymatic DNA synthesis using 100% dye-labeled terminal phosphate-linked nucleotides. Nucleosides Nucleotides Nucleic Acids 27, 1072–1083 (2008).
Reynolds, B., Miller, R., Williams, J.G. & Anderson, J.P. Synthesis and stability of novel terminal phosphate-labeled nucleotides. Nucleosides Nucleotides Nucleic Acids 27, 18–30 (2008).
Steffens, D.L. & Williams, J.G. Efficient site-directed saturation mutagenesis using degenerate oligonucleotides. J. Biomol. Tech. 18, 147–149 (2007).
Ryu, J., Hong, S.S., Horn, B.K.P., Freeman, D.M. & Mermelstein, M.S. Multibeam interferometric illumination as the primary source of resolution in optical microscopy. Appl. Phys. Lett. 88, 171112 (2006).
Mico, V. et al. Transverse resolution improvement using rotating-grating time-multiplexing approach. J. Opt. Soc. Am. A Opt. Image Sci. Vis. 25, 1115–1129 (2008).
Pierce, J.R. An Introduction to Information Theory, edn. 2 (Dover Press, Mineola, NY, 1980).
Lundstrom, M. Moore's law forever? Science 299, 210–211 (2001).
Stelzer, E.H.K. Beyond the diffraction limit? Nature 417, 806–807 (2002).
Lloyd, S. Ultimate physical limits to computation. Nature 406, 1047–1054 (2000).
Lloyd, S., Giovannetti, V. & Maccone, L. Physical limits to communication. Phys. Rev. Lett. 93, 100501–100504 (2004).
Stelzer, E.H.K. & Grill, S. The uncertainty principle applied to estimate focal spot dimensions. Opt. Commun. 173, 51–56 (2000).
Moerner, W.E. & Fromm, D.P. Methods of single-molecule fluorescence spectroscopy and microscopy. Rev. Sci. Inst. 74, 3597–3619 (2003).
Basché, Th., Ambrose, W.P. & Moerner, W.E. Optical spectra and kinetics of single impurity molecules in a polymer: spectral diffusion and persistent spectral hole burning. J. Opt. Soc. Am. B 9, 829–836 (1992).
Stevenson, C.L. & Winefordner, J.D. Estimating detection limits in ultratrace analysis. Part I: The variability of estimated detection limits. Applied Spect. 45, 1217–1224 (1991).
Stevenson, C.L. & Winefordner, J.D. Estimating detection limits in ultratrace analysis. Part II: Detecting and counting atoms and molecules. Applied Spect. 46, 407–419 (1992).
Stevenson, C.L. & Winefordner, J.D. Estimating detection limits in ultratrace analysis. Part III: Monitoring atoms and molecules with laser-induced fluorescence. Applied Spect. 46, 715–724 (1992).
Landauer, R. Minimal energy requirements in communication. Science 272, 1914–1918 (1996).
Zurek, W.H. Thermodynamic cost of computation, algorithmic complexity and the information metric. Nature 341, 119–124 (1989).
Simon, S.H., Moustakas, A.L., Stoytchev, M. & Safar, H. Communication in a disordered world. Phys. Today 54, 38–43 (2001).
Méray, L. & Demény, O. Detection limit and decision thresholds in spectrometry. Appl. Spect. 55, 1102–1108 (2001).
Lachmann, M., Newman, M.E.J. & Moore, C. The physical limits of communication. Am. J. Phys. 72, 1290–1293 (2004).
Gordon, J.M. & Feuermann, D. Optical performance at the thermodynamic limit with tailored imaging designs. Appl. Opt. 44, 2327–2331 (2005).
Sheehan, P.E. & Whitman, L.J. Detection limits for nanoscale biosensors. Nano Lett. 5, 803–807 (2005).
Prummer, M. Sick., B. Renn, A. & Wild, U.P. Multiparameter microscopy and spectroscopy for single-molecule analytics. Anal. Chem. 76, 1633–1640 (2004).
Hubaux, A. & Vos, G. Decision and detection limits for linear calibration curves. Anal. Chem. 42, 849–855 (1970).
Lundquist, P.M. et al. Parallel confocal detection of single molecules in real time. Opt. Lett. 33, 1026–1028 (2008).
Bashford, G. et al. Automated bead-trapping apparatus and control system for single-molecule DNA sequencing. Opt. Express 16, 3445–3455 (2008).
Gibson, J.D. Principles of Digital and Analog Communications, edn. 2 (Macmillan Publishing, New York, 1993).
Honigs, D. The sayings of Tomas Hirschfeld. Applied Spect. 40, 11A (1986).
Hirschfeld, T. Limits of analysis. Anal. Chem. 48, 16A–31A (1976).
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Eckmann, J.-P. Trading codes for errors. Proc. Natl. Acad. Sci. USA 105, 8165–8166 (2008).
Acknowledgements
This work was supported in part by the National Human Genome Research Institute, National Institutes of Health.
Author information
Authors and Affiliations
Contributions
C.W.F. and L.R.M. wrote this review, with additions and editorial assistance from S.A.B., G.M.C., T.H., X.H., S.B.J., J.R.N., D.C.S. and D.V.V., who contributed portions of the text and read drafts of the manuscript for accuracy. J.A.S. is the scientific manager of the NHGRI Sequencing Technology Development Program; he proposed the idea for the review, provided a forum to begin its formulation at a program meeting and read the manuscript for accuracy.
Corresponding author
Ethics declarations
Competing interests
G.M.C. plays advisory roles at several companies relevant to this paper and these are listed at http://arep.med.harvard.edu/gmc/tech.html.
Rights and permissions
About this article
Cite this article
Fuller, C., Middendorf, L., Benner, S. et al. The challenges of sequencing by synthesis. Nat Biotechnol 27, 1013–1023 (2009). https://doi.org/10.1038/nbt.1585
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1585
This article is cited by
-
SSR Marker Acquisition and Application from Transcriptome of Captive Chinese Forest Musk Deer (Moschus berezovskii)
Biochemical Genetics (2023)
-
Sequence-dependence of Cy3 and Cy5 dyes in 3ʹ terminally-labeled single-stranded DNA
Scientific Reports (2022)
-
PCR inhibition in qPCR, dPCR and MPS—mechanisms and solutions
Analytical and Bioanalytical Chemistry (2020)
-
Evaluating 18s-rRNA LAMP and selective whole genome amplification (sWGA) assay in detecting asymptomatic Plasmodium falciparum infections in blood donors
Malaria Journal (2019)
-
Gen2Epi: an automated whole-genome sequencing pipeline for linking full genomes to antimicrobial susceptibility and molecular epidemiological data in Neisseria gonorrhoeae
BMC Genomics (2019)