Abstract
Although it has previously been shown that Neanderthals contributed DNA to modern humans1,2, not much is known about the genetic diversity of Neanderthals or the relationship between late Neanderthal populations at the time at which their last interactions with early modern humans occurred and before they eventually disappeared. Our ability to retrieve DNA from a larger number of Neanderthal individuals has been limited by poor preservation of endogenous DNA3 and contamination of Neanderthal skeletal remains by large amounts of microbial and present-day human DNA3,4,5. Here we use hypochlorite treatment6 of as little as 9 mg of bone or tooth powder to generate between 1- and 2.7-fold genomic coverage of five Neanderthals who lived around 39,000 to 47,000 years ago (that is, late Neanderthals), thereby doubling the number of Neanderthals for which genome sequences are available. Genetic similarity among late Neanderthals is well predicted by their geographical location, and comparison to the genome of an older Neanderthal from the Caucasus2,7 indicates that a population turnover is likely to have occurred, either in the Caucasus or throughout Europe, towards the end of Neanderthal history. We find that the bulk of Neanderthal gene flow into early modern humans originated from one or more source populations that diverged from the Neanderthals that were studied here at least 70,000 years ago, but after they split from a previously sequenced Neanderthal from Siberia2 around 150,000 years ago. Although four of the Neanderthals studied here post-date the putative arrival of early modern humans into Europe, we do not detect any recent gene flow from early modern humans in their ancestry.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
European Nucleotide Archive
NCBI Reference Sequence
Change history
28 March 2018
In the original PDF version of this Letter, Extended Data Tables 1-4 were corrupted; this has been corrected online.
References
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010)
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014)
Pääbo, S. et al. Genetic analyses from ancient DNA. Annu. Rev. Genet. 38, 645–679 (2004)
Gilbert, M. T., Bandelt, H. J., Hofreiter, M. & Barnes, I. Assessing ancient DNA studies. Trends Ecol. Evol. 20, 541–544 (2005)
Krause, J. et al. A complete mtDNA genome of an early modern human from Kostenki, Russia. Curr. Biol. 20, 231–236 (2010)
Korlevic´, P. et al. Reducing microbial and human contamination in DNA extractions from ancient bones and teeth. Biotechniques 59, 87–93 (2015)
Prüfer, K. et al. A high-coverage Neandertal genome from Vindija Cave in Croatia. Science 358, 655–658 (2017)
Hublin, J.-J. The modern human colonization of western Eurasia: when and where? Quat. Sci. Rev. 118, 194–210 (2015)
Higham, T. et al. The timing and spatiotemporal patterning of Neanderthal disappearance. Nature 512, 306–309 (2014)
Fu, Q. et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514, 445–449 (2014)
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015)
Rougier, H. et al. Neandertal cannibalism and Neandertal bones used as tools in Northern Europe. Sci. Rep. 6, 29005 (2016)
Semal, P. et al. New data on the late Neandertals: direct dating of the Belgian Spy fossils. Am. J. Phys. Anthropol. 138, 421–428 (2009)
Soressi, M. et al. in Préhistoire entre Vienne et Charente — Hommes et sociétés du Paléolithique mémoire 38 (eds Buisson-Catil, J. & Primault, J. ) 221–234 (Association des Publications Chauvinoises, 2010)
Pinhasi, R., Higham, T. F., Golovanova, L. V. & Doronichev, V. B. Revised age of late Neanderthal occupation and the end of the Middle Paleolithic in the northern Caucasus. Proc. Natl Acad. Sci. USA 108, 8611–8616 (2011)
Briggs, A. W. et al. Patterns of damage in genomic DNA sequences from a Neandertal. Proc. Natl Acad. Sci. USA 104, 14616–14621 (2007)
Meyer, M. et al. A mitochondrial genome sequence of a hominin from Sima de los Huesos. Nature 505, 403–406 (2014)
Dalén, L. et al. Partial genetic turnover in Neandertals: continuity in the East and population replacement in the West. Mol. Biol. Evol. 29, 1893–1897 (2012)
Mendez, F. L., Poznik, G. D., Castellano, S. & Bustamante, C. D. The divergence of Neandertal and modern human Y chromosomes. Am. J. Hum. Genet. 98, 728–734 (2016)
Meyer, M. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012)
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016)
Cavalli-Sforza, L. L., Menozzi, P. & Piazza, A. The History and Geography of Human Genes (Princeton Univ. Press, 1994)
Svensson, A. et al. A 60 000 year Greenland stratigraphic ice core chronology. Clim. Past 4, 47–57 (2008)
Hublin, J.-J. & Roebroeks, W. Ebb and flow or regional extinctions? On the character of Neandertal occupation of northern environments. C. R. Palevol 8, 503–509 (2009)
Müller, U. C. et al. The role of climate in the spread of modern humans into Europe. Quat. Sci. Rev. 30, 273–279 (2011)
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014)
Fu, Q. et al. The genetic history of Ice Age Europe. Nature 534, 200–205 (2016)
Kuhlwilm, M. et al. Ancient gene flow from early modern humans into Eastern Neanderthals. Nature 530, 429–433 (2016)
Overmann, K. & Coolidge, F. Human species and mating systems: Neandertal–Homo sapiens reproductive isolation and the archaeological and fossil records. J. Anthropol. Sci. 91, 91–110 (2013)
Karmin, M. et al. A recent bottleneck of Y chromosome diversity coincides with a global change in culture. Genome Res. 25, 459–466 (2015)
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl Acad. Sci. USA 110, 15758–15763 (2013)
Gansauge, M. T. & Meyer, M. Single-stranded DNA library preparation for the sequencing of ancient or damaged DNA. Nat. Protoc. 8, 737–748 (2013)
Dabney, J. & Meyer, M. Length and GC-biases during sequencing library amplification: a comparison of various polymerase-buffer systems with ancient and modern DNA sequencing libraries. Biotechniques 52, 87–94 (2012)
Kircher, M., Sawyer, S. & Meyer, M. Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res. 40, e3 (2012)
Maricic, T., Whitten, M. & Pääbo, S. Multiplexed DNA sequence capture of mitochondrial genomes using PCR products. PLoS ONE 5, e14004 (2010)
Fu, Q. et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proc. Natl Acad. Sci. USA 110, 2223–2227 (2013)
Welker, F. et al. Palaeoproteomic evidence identifies archaic hominins associated with the Châtelperronian at the Grotte du Renne. Proc. Natl Acad. Sci. USA 113, 11162–11167 (2016)
Renaud, G., Kircher, M., Stenzel, U. & Kelso, J. freeIbis: an efficient basecaller with calibrated quality scores for Illumina sequencers. Bioinformatics 29, 1208–1209 (2013)
Renaud, G., Stenzel, U. & Kelso, J. leeHom: adaptor trimming and merging for Illumina sequencing reads. Nucleic Acids Res. 42, e141 (2014)
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Sawyer, S., Krause, J., Guschanski, K., Savolainen, V. & Pääbo, S. Temporal patterns of nucleotide misincorporations and DNA fragmentation in ancient DNA. PLoS ONE 7, e34131 (2012)
Green, R. E. et al. A complete Neandertal mitochondrial genome sequence determined by high-throughput sequencing. Cell 134, 416–426 (2008)
Briggs, A. W. et al. Targeted retrieval and analysis of five Neandertal mtDNA genomes. Science 325, 318–321 (2009)
Skoglund, P. et al. Separating endogenous ancient DNA from modern day contamination in a Siberian Neandertal. Proc. Natl Acad. Sci. USA 111, 2229–2234 (2014)
Brown, S. et al. Identification of a new hominin bone from Denisova Cave, Siberia using collagen fingerprinting and mitochondrial DNA analysis. Sci. Rep. 6, 23559 (2016)
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015)
Gansauge, M. T. & Meyer, M. Selective enrichment of damaged DNA molecules for ancient genome sequencing. Genome Res. 24, 1543–1549 (2014)
Posth, C. et al. Deeply divergent archaic mitochondrial genome provides lower time boundary for African gene flow into Neanderthals. Nat. Commun. 8, 16046 (2017)
Ermini, L. et al. Complete mitochondrial genome sequence of the Tyrolean Iceman. Curr. Biol. 18, 1687–1693 (2008)
Gilbert, M. T. P. et al. Paleo-Eskimo mtDNA genome reveals matrilineal discontinuity in Greenland. Science 320, 1787–1789 (2008)
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013)
Krause, J. et al. The complete mitochondrial DNA genome of an unknown hominin from southern Siberia. Nature 464, 894–897 (2010)
Reich, D. et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468, 1053–1060 (2010)
Sawyer, S. et al. Nuclear and mitochondrial DNA sequences from two Denisovan individuals. Proc. Natl Acad. Sci. USA 112, 15696–15700 (2015)
Horai, S. et al. Man’s place in Hominoidea revealed by mitochondrial DNA genealogy. J. Mol. Evol. 35, 32–43 (1992)
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013)
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013)
Bouckaert, R. et al. BEAST 2: a software platform for Bayesian evolutionary analysis. PLOS Comput. Biol. 10, e1003537 (2014)
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nat. Methods 9, 772 (2012)
Baele, G. et al. Improving the accuracy of demographic and molecular clock model comparison while accommodating phylogenetic uncertainty. Mol. Biol. Evol. 29, 2157–2167 (2012)
Baele, G., Li, W. L. S., Drummond, A. J., Suchard, M. A. & Lemey, P. Accurate model selection of relaxed molecular clocks in Bayesian phylogenetics. Mol. Biol. Evol. 30, 239–243 (2013)
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012)
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006)
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006)
Meyer, M. et al. Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins. Nature 531, 504–507 (2016)
Busing, F. M. T. A., Meijer, E. & Van Der Leeden, R. Delete-m jackknife for unequal m. Stat. Comput. 9, 3–8 (1999)
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987)
Schliep, K. P. phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011)
The Chimpanzee Sequencing and Analysis Consortium. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 437, 69–87 (2005)
Locke, D. P. et al. Comparative and demographic analysis of orang-utan genomes. Nature 469, 529–533 (2011)
Gibbs, R. A. et al. Evolutionary and biomedical insights from the rhesus macaque genome. Science 316, 222–234 (2007)
Moorjani, P. et al. A genetic method for dating ancient genomes provides a direct estimate of human generation interval in the last 45,000 years. Proc. Natl Acad. Sci. USA 113, 5652–5657 (2016)
The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012)
Paten, B., Herrero, J., Beal, K., Fitzgerald, S. & Birney, E. Enredo and Pecan: genome-wide mammalian consistency-based multiple alignment with paralogs. Genome Res. 18, 1814–1828 (2008)
Paten, B. et al. Genome-wide nucleotide-level mammalian ancestor reconstruction. Genome Res. 18, 1829–1843 (2008)
Gravel, S. et al. Demographic history and rare allele sharing among human populations. Proc. Natl Acad. Sci. USA 108, 11983–11988 (2011)
Acknowledgements
We thank A. Weihmann and B. Höber for their help with DNA sequencing, U. Stenzel for computational support and advice for data analysis, R. Barr for the help with the graphics, V. Slon for helpful discussions and comments on the manuscript. Q.F. is funded in part by NSFC (91731303, 41672021, 41630102), CAS (QYZDB-SSW-DQC003, XDB13000000, XDA19050102, XDPB05) and the Howard Hughes Medical Institute (grant number 55008731). D.R. is supported by the US National Science Foundation (grant BCS-1032255) and by an Allen Discovery Center of the Paul Allen Foundation and is an investigator of the Howard Hughes Medical Institute. This study was funded by the Max Planck Society and the European Research Council (grant agreement number 694707 to S.P.). M.So. thanks the owner of Les Cottés, and the French Ministry of Culture for financial support and excavation permits.
Author information
Authors and Affiliations
Contributions
M.H., M.M. and S.P. conceived the study. M.Sl., N.P., D.R., K.P., M.M., S.P. and J.Ke. supervised the study. M.H., P.K., S.N. and B.N. performed ancient DNA laboratory work. H.R., I.C., P.Se., M.So., S.T., J.-J.H., I.G., Ž.K., P.R., L.V.G., V.B.D., C.P. and J.Kr. provided and analysed archaeological material. M.H., Q.F., A.H., M.P., F.M., S.G., P.Sk. and V.N. analysed ancient DNA data. M.H., M.M., S.P. and J.Ke. wrote the manuscript with the input of all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks C. Lalueza-Fox, C. Stringer and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
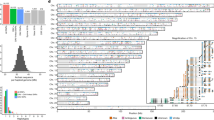
Extended Data Figure 1 Frequency of nucleotide substitutions at the beginning and the end of nuclear alignments for the final dataset of Les Cottés Z4-1514, Goyet Q56-1, Mezmaiskaya 2, Vindija 87 and Spy 94a.
Only fragments of at least 35 bp that mapped to the human reference genome with a mapping quality of at least 25 (MQ ≥ 25) were used for this analysis. Solid lines depict all fragments and dashed lines the fragments that have a C-to-T substitution at the opposing end (‘conditional’ C-to-T substitutions). All other types of substitutions are marked in grey.
Extended Data Figure 2 Fragment size distribution of fragments longer than 35 bp mapped to the human reference genome with MQ ≥ 25 for each of the five late Neanderthals.
All fragments are depicted in solid lines and fragments with C-to-T substitutions to the reference genome (putatively deaminated fragments) are depicted with dashed lines.
Extended Data Figure 3 Sex determination based on the number of fragments aligning to the X chromosome and the autosomes.
The expected ratios of X to (X + autosomal) fragments for a female and a male individual are depicted as dashed lines. The results were concordant for all fragments (in red) and for deaminated fragments only (in grey).
Extended Data Figure 4 Principal component analysis of the genomes of Vindija 33.19, Altai, the Denisovan individual, five late Neanderthals and Mezmaiskaya 1.
Genomes of the high-coverage archaics were used to estimate the eigenvectors of the genetic variation and low-coverage Neanderthals were projected onto the plane. Only transversion polymorphisms and bi-allelic sites were considered for the analysis, to a total of 1,010,417 sites as defined by the high-coverage genomes. PC, principal component.
Supplementary information
Supplementary Information
This file contains Supplementary Sections 1-12, Supplementary Tables and Supplementary References – see contents page for details. (PDF 4507 kb)
Supplementary Table 1
An assessment of the catalog of modern-human-specific fixed derived changes in the late Neandertals. (CSV 1793 kb)
Rights and permissions
About this article
Cite this article
Hajdinjak, M., Fu, Q., Hübner, A. et al. Reconstructing the genetic history of late Neanderthals. Nature 555, 652–656 (2018). https://doi.org/10.1038/nature26151
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature26151
This article is cited by
-
Evolutionary origin of germline pathogenic variants in human DNA mismatch repair genes
Human Genomics (2024)
-
The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes
Scientific Data (2024)
-
The Neanderthal niche space of Western Eurasia 145 ka to 30 ka ago
Scientific Reports (2024)
-
Ancient human DNA recovered from a Palaeolithic pendant
Nature (2023)
-
Anatomically modern human in the Châtelperronian hominin collection from the Grotte du Renne (Arcy-sur-Cure, Northeast France)
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.