Abstract
In vast areas of the ocean, the scarcity of iron controls the growth and productivity of phytoplankton1,2. Although most dissolved iron in the marine environment is complexed with organic molecules3, picomolar amounts of labile inorganic iron species (labile iron) are maintained within the euphotic zone4 and serve as an important source of iron for eukaryotic phytoplankton and particularly for diatoms5. Genome-enabled studies of labile iron utilization by diatoms have previously revealed novel iron-responsive transcripts6,7, including the ferric iron-concentrating protein ISIP2A8, but the mechanism behind the acquisition of picomolar labile iron remains unknown. Here we show that ISIP2A is a phytotransferrin that independently and convergently evolved carbonate ion-coordinated ferric iron binding. Deletion of ISIP2A disrupts high-affinity iron uptake in the diatom Phaeodactylum tricornutum, and uptake is restored by complementation with human transferrin. ISIP2A is internalized by endocytosis, and manipulation of the seawater carbonic acid system reveals a second-order dependence on the concentrations of labile iron and carbonate ions. In P. tricornutum, the synergistic interaction of labile iron and carbonate ions occurs at environmentally relevant concentrations, revealing that carbonate availability co-limits iron uptake. Phytotransferrin sequences have a broad taxonomic distribution8 and are abundant in marine environmental genomic datasets9,10, suggesting that acidification-driven declines in the concentration of seawater carbonate ions will have a negative effect on this globally important eukaryotic iron acquisition mechanism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Moore, C. M. et al. Processes and patterns of oceanic nutrient limitation. Nat. Geosci. 6, 701–710 (2013)
Boyd, P. W. et al. Mesoscale iron enrichment experiments 1993–2005: synthesis and future directions. Science 315, 612–617 (2007)
Gledhill, M. & Buck, K. N. The organic complexation of iron in the marine environment: a review. Front. Microbiol. 3, 69 (2012)
Barbeau, K., Rue, E. L., Bruland, K. W. & Butler, A. Photochemical cycling of iron in the surface ocean mediated by microbial iron(iii)-binding ligands. Nature 413, 409–413 (2001)
Morel, F. M. M., Kustka, A. B. & Shaked, Y. The role of unchelated Fe in the iron nutrition of phytoplankton. Limnol. Oceanogr. 53, 400–404 (2008)
Allen, A. E. et al. Whole-cell response of the pennate diatom Phaeodactylum tricornutum to iron starvation. Proc. Natl Acad. Sci. USA 105, 10438–10443 (2008)
Lommer, M. et al. Genome and low-iron response of an oceanic diatom adapted to chronic iron limitation. Genome Biol. 13, R66 (2012)
Morrissey, J. et al. A novel protein, ubiquitous in marine phytoplankton, concentrates iron at the cell surface and facilitates uptake. Curr. Biol. 25, 364–371 (2015)
Marchetti, A. et al. Comparative metatranscriptomics identifies molecular bases for the physiological responses of phytoplankton to varying iron availability. Proc. Natl Acad. Sci. USA 109, E317–E325 (2012)
Bertrand, E. M. et al. Phytoplankton–bacterial interactions mediate micronutrient colimitation at the coastal Antarctic sea ice edge. Proc. Natl Acad. Sci. USA 112, 9938–9943 (2015)
Lambert, L. A., Perri, H., Halbrooks, P. J. & Mason, A. B. Evolution of the transferrin family: conservation of residues associated with iron and anion binding. Comp. Biochem. Physiol. 142, 129–141 (2005)
Aisen, P., Leibman, A. & Zweier, J. Stoichiometric and site characteristics of the binding of iron to human transferrin. J. Biol. Chem. 253, 1930–1937 (1978)
Baker, H. M., Anderson, B. F. & Baker, E. N. Dealing with iron: common structural principles in proteins that transport iron and heme. Proc. Natl Acad. Sci. USA 100, 3579–3583 (2003)
Fisher, M., Gokhman, I., Pick, U. & Zamir, A. A structurally novel transferrin-like protein accumulates in the plasma membrane of the unicellular green alga Dunaliella salina grown in high salinities. J. Biol. Chem. 272, 1565–1570 (1997)
Anderson, M. A. & Morel, F. M. The influence of aqueous iron chemistry on the uptake of iron by the coastal diatom Thalassiosira weissflogii. Limnol. Oceanogr. 27, 789–813 (1982)
Bruns, C. M. et al. Structure of Haemophilus influenzae Fe+3-binding protein reveals convergent evolution within a superfamily. Nat. Struct. Biol. 4, 919–924 (1997)
Och, L. M. & Shields-Zhou, G. A. The Neoproterozoic oxygenation event: environmental perturbations and biogeochemical cycling. Earth Sci. Rev. 110, 26–57 (2012)
Smith, S. R. et al. Transcriptional orchestration of the global cellular response of a model pennate diatom to diel light cycling under iron limitation. PLoS Genet. 12, e1006490 (2016)
Karas, B. J. et al. Designer diatom episomes delivered by bacterial conjugation. Nat. Commun. 6, 6925 (2015)
Harding, C., Heuser, J. & Stahl, P. Receptor-mediated endocytosis of transferrin and recycling of the transferrin receptor in rat reticulocytes. J. Cell Biol. 97, 329–339 (1983)
Fisher, M., Zamir, A. & Pick, U. Iron uptake by the halotolerant alga Dunaliella is mediated by a plasma membrane transferrin. J. Biol. Chem. 273, 17553–17558 (1998)
Schlabach, M. R. & Bates, G. W. The synergistic binding of anions and Fe3+ by transferrin. Implications for the interlocking sites hypothesis. J. Biol. Chem. 250, 2182–2188 (1975)
Halbrooks, P. J., Mason, A. B., Adams, T. E., Briggs, S. K. & Everse, S. J. The oxalate effect on release of iron from human serum transferrin explained. J. Mol. Biol. 339, 217–226 (2004)
Doney, S. C ., Fabry, V. J ., Feely, R. A. & Kleypas, J. A. Ocean acidification: the other CO2 problem. Annu. Rev. Mar. Sci. 1, 169–192 (2009)
Sunda, W. & Huntsman, S. Effect of pH, light, and temperature on Fe–EDTA chelation and Fe hydrolysis in seawater. Mar. Chem. 84, 35–47 (2003)
Shi, D., Xu, Y., Hopkinson, B. M. & Morel, F. M. M. Effect of ocean acidification on iron availability to marine phytoplankton. Science 327, 676–679 (2010)
Allen, M. D., del Campo, J. A., Kropat, J. & Merchant, S. S. FEA1, FEA2, and FRE1, encoding two homologous secreted proteins and a candidate ferrireductase, are expressed coordinately with FOX1 and FTR1 in iron-deficient Chlamydomonas reinhardtii. Eukaryot. Cell 6, 1841–1852 (2007)
Sasaki, T., Kurano, N. & Miyachi, S. Cloning and characterization of high-CO2-specific cDNAs from a marine microalga, Chlorococcum littorale, and effect of CO2 concentration and iron deficiency on the gene expression. Plant Cell Physiol. 39, 131–138 (1998)
Saito, M. A., Goepfert, T. J. & Ritt, J. T. Some thoughts on the concept of colimitation: three definitions and the importance of bioavailability. Limnol. Oceanogr. 53, 276–290 (2008)
Lis, H., Shaked, Y., Kranzler, C., Keren, N. & Morel, F. M. M. Iron bioavailability to phytoplankton: an empirical approach. ISME J. 9, 1003–1013 (2015)
Caron, D. A. et al. Probing the evolution, ecology and physiology of marine protists using transcriptomics. Nat. Rev. Microbiol. 15, 6–20 (2017)
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013)
Gouy, M., Guindon, S. & Gascuel, O. SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 27, 221–224 (2010)
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014)
Lartillot, N., Lepage, T. & Blanquart, S. PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics 25, 2286–2288 (2009)
Berney, C. & Pawlowski, J. A molecular time-scale for eukaryote evolution recalibrated with the continuous microfossil record. Proc. R. Soc. B 273, 1867–1872 (2006)
Parfrey, L. W., Lahr, D. J., Knoll, A. H. & Katz, L. A. Estimating the timing of early eukaryotic diversification with multigene molecular clocks. Proc. Natl Acad. Sci. USA 108, 13624–13629 (2011)
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012)
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat. Methods 8, 785–786 (2011)
Emanuelsson, O., Nielsen, H., Brunak, S. & von Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J. Mol. Biol. 300, 1005–1016 (2000)
Nielsen, H., Engelbrecht, J., Brunak, S. & von Heijne, G. Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng. 10, 1–6 (1997)
Möller, S., Croning, M. D. R. & Apweiler, R. Evaluation of methods for the prediction of membrane spanning regions. Bioinformatics 17, 646–653 (2001)
Pierleoni, A., Martelli, P. L. & Casadio, R. PredGPI: a GPI-anchor predictor. BMC Bioinformatics 9, 392 (2008)
Sunda, W. G., Price, N. M. & Morel, F. M. M. in Algal Culturing Techniques (ed. Anderson. R. A. ) 35–63 (Elsevier, 2005)
Zaslavskaia, L. A., Lippmeier, J. C., Kroth, P. G., Grossman, A. R. & Apt, K. E. Transformation of the diatom Phaeodactylum tricornutum (Bacillariophyceae) with a variety of selectable marker and reporter genes. J. Phycol. 36, 379–386 (2000)
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009)
Falciatore, A., Casotti, R., Leblanc, C., Abrescia, C. & Bowler, C. Transformation of nonselectable reporter genes in marine diatoms. Mar. Biotechnol. 1, 239–251 (1999)
Poulsen, N. & Kröger, N. A new molecular tool for transgenic diatoms: control of mRNA and protein biosynthesis by an inducible promoter–terminator cassette. FEBS J. 272, 3413–3423 (2005)
Weyman, P. D. et al. Inactivation of Phaeodactylum tricornutum urease gene using transcription activator-like effector nuclease-based targeted mutagenesis. Plant Biotechnol. J. 13, 460–470 (2015)
Hudson, R. J. M. & Morel, F. M. M. Distinguishing between extra- and intracellular iron in marine phytoplankton. Limnol. Oceanogr. 34, 1113–1120 (1989)
Dickson, A. G., Sabine, C. L., & Christian, J. R. (eds) Guide to Best Practices for Ocean CO 2 Measurements (North Pacific Marine Science Organization, 2007)
Kustka, A. B., Allen, A. E. & Morel, F. M. Sequence analysis and transcriptional regulation of iron acquisition genes in two marine diatoms. J. Phycol. 43, 715–729 (2007)
Dutta, D., Williamson, C. D., Cole, N. B. & Donaldson, J. G. Pitstop 2 is a potent inhibitor of clathrin-independent endocytosis. PLoS ONE 7, e45799 (2012)
Carter, B. R., Radich, J. A., Doyle, H. L. & Dickson, A. G. An automated system for spectrophotometric seawater pH measurements. Limnol. Oceanogr. Methods 11, 16–27 (2013)
Lewis, E ., Wallace, D. & Allison, L. J. Program developed for CO 2 System Calculations (Carbon Dioxide Information Analysis Center, 1998)
Slinker, B. K. The statistics of synergism. J. Mol. Cell. Cardiol. 30, 723–731 (1998)
R Core Team. R: A Language and Environment for Statistical Computing. http://www.R-project.org/ (R Foundation for Statistical Computing, Vienna, Austria, 2014)
Acknowledgements
We thank J. Badger for early contributions to phylogenetic analyses, A. Dickson for pH analysis, K. Forsch for CSV measurements and E. Bertrand for trace-metal clean techniques. This study was supported by the National Science Foundation (NSF-MCB-1024913, NSF-ANT-1043671 and NSF-OCE-0727997), United States Department of Energy Genomics Science program (DE-SC00006719 and DE-SC0008593), and the Gordon and Betty Moore Foundation grant GBMF3828 (A.E.A.); NSF-1557928 (A.B.K.); and the Czech Science Foundation, project 15-17643S (M.O. and A.H.).
Author information
Authors and Affiliations
Contributions
J.B.M., A.B.K., M.O. and A.E.A. designed the study and interpreted the results. J.B.M., M.O. and A.H. generated and analysed phylogenetic and molecular clock data. J.B.M. and B.J.K. generated mutant cell lines, J.B.M. and A.B.K. with assistance from K.A.B. performed physiology experiments. J.B.M. performed microscopy, and H.Z. performed western blots. T.K. and A.J.A. analysed inorganic carbon species. J.B.M. and J.P.M. conducted statistical analyses. J.B.M. wrote the paper with input from A.E.A., A.B.K., M.O., J.P.M, A.J.A. and K.A.B. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S. Amin, E. DeLong and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Conservation of active site amino acids.
a, Left, conservation of putative iron-coordinating (red) and carbonate-coordinating (green) amino acids among phylogenetic groups, filled circles indicate >85% conservation, unfilled circles indicate <20% conservation; arginine and lysine were permitted at the carbonate-binding site11. Right, logo tag detailing alignment conservation at the anion-binding region. Phosphonate-binding proteins (Bacteria and Thaumarchaea) retain the S/T-rich phosphonate-binding area, whereas transferrin (Euryarchaea and Metazoa) and phytotransferrin have the carbonate-coordinating K/R insertion. A downstream SAG region used to stabilize carbonate in transferrin24 shows some conservation in phytotransferrin, but an upstream threonine (−4 amino acids from the conserved Arg) is not present. b, Active site homology among ISIP2A (PT54465) and two transmembrane-anchored P. tricornutum homologues, PT45708 and PT54466. Amino acid distances based on PT54465, red/green triangles are iron- and carbonate-coordinating amino acids.
Extended Data Figure 2 Estimated divergence times between transferrin, phytotransferrin and PBPs.
Analyses were carried out using PhyloBayes (top) and BEAST (bottom). For the fossil calibration points used in generating the minimum and maximum constraints, see Supplementary Table 1. MYA, million years ago.
Extended Data Figure 3 Bayesian (PhyloBayes) phylogenetic tree contrasted with alternate topology derived using maximum likelihood.
Prasinococcus capsulatus, Pyraminomonas obovata and other chlorophyte algae have copies of transferrin and phytotransferrin, whereas all other non-chlorophyte algae have only phytotransferrin. Branches are colour-coded by phylogenetic group. Scale bar, 0.5 substitutions per position.
Extended Data Figure 4 Characterization of ISIP2A.
a, Western blot of wild-type P. tricornutum compared to ISIP2A-knockout strains (ΔISIP2A). The estimated mass of ISIP2A protein is 57 kDa. b, Specific growth rates of wild type compared to Δ ISIP2A. c, Uptake of 59Fe-ferrioxamine B is unaffected in Δ ISIP2A, suggesting an alternate pathway for uptake of iron–siderophore complexes. d, Effect of a clathrin-mediated endocytosis (CME) inhibitor on short-term 59Fe′ uptake rates, wild-type P. tricornutum. e, Upon addition of iron to iron-limited cells, the membrane-impermeable FM1-43 stain is internalized into vesicle-like inclusions. Scale bar, 5 μm. Pink is plastid auto-fluorescence. b, Data are mean ± s.e.m.; two-sided heteroscedastic t-test, n = biological replicates for wild-type and ISIP2A are indicated in brackets as (WT, ISIP2A) and P values are indicated. 12 pM Fe′ (5, 3), P = 0.008; 22 pM Fe′ (7, 5), P = 0.003; 43 pM Fe′ (6, 4), P = 0.006; 83 pM Fe′ (4, 4), P = 0.007; 165 pM Fe′ (3, 3), P = 0.07. d, Data are mean ± s.e.m.; n = 3 biological replicates; two-sided t-test, P = 0.009.
Extended Data Figure 5 Reconciliation of the carbonate effect with the influence of acidification on seawater iron chemistry.
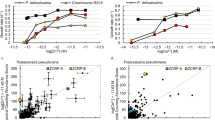
a, Linearized representation of Fig. 3a, with high iron uptake rates (blue triangles) plotted on the secondary y axis, and pH values for each [CO32−] listed at top. Fe–FOB uptake rates decrease with pH, consistent with the findings of ref. 26, whereas Fe′ uptake rates increase with decreasing pH, inconsistent with the hypothesized effects of acidification on iron–EDTA chemistry26. This inconsistency is resolved when uptake rates are plotted as a function of the synergistic interaction between Fe′ and CO32− (Fig. 3b). b, c, Under CO2-induced acidification, the strong influence of pH on [Fe′] results in a significant correlation of uptake to [Fe′], although the interaction Fe′ and CO32− results in a better fit (Fig. 4a, solid line). d, e, When the change in [Fe′] is constrained relative to pH and [CO32−], uptake rates are positively correlated with [CO32−] and uncorrelated to [Fe′], revealing the influence of the carbonate ion on Fe′ uptake rates. For statistical analyses of the linear regressions, see Extended Data Table 4.
Extended Data Figure 6 Derivation of second-order and constitutive rate constants from P. tricornutum resuspended in NaHCO3-manipulated medium.
a, Regression of the interaction product of Fe′ and CO32−. Regression excludes the two observations in which the medium was not supplemented with NaHCO3. b, 59Fe uptake rates for treatments incubated with low (2–5 pM Fe′, open symbols) and high (20–50 pM Fe′, closed symbols) 59Fe versus [CO32−]. c, Demonstration of reproducibility at low [Fe′]. Data are mean ± s.e.m.; n = 3 biological replicates. d, Uptake rates normalized to [Fe′], plotted against [CO32−]. The slope has units of mol Fe cell−1 h−1 (M Fe′)−1 (M CO32−)−1, equivalent to the pseudo-first-order uptake rate with respect to carbonate. e, Rates of uptake calculated as a function of [Fe′] and [CO32−] versus measured rates.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 and Supplementary Tables 1-2. (PDF 332 kb)
Rights and permissions
About this article
Cite this article
McQuaid, J., Kustka, A., Oborník, M. et al. Carbonate-sensitive phytotransferrin controls high-affinity iron uptake in diatoms. Nature 555, 534–537 (2018). https://doi.org/10.1038/nature25982
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25982
This article is cited by
-
Short-term acidification promotes diverse iron acquisition and conservation mechanisms in upwelling-associated phytoplankton
Nature Communications (2023)
-
Gene duplication and functional divergence of new genes contributed to the polar acclimation of Antarctic green algae
Marine Life Science & Technology (2023)
-
Proteomic traits vary across taxa in a coastal Antarctic phytoplankton bloom
The ISME Journal (2022)
-
Genomic adaptation of the picoeukaryote Pelagomonas calceolata to iron-poor oceans revealed by a chromosome-scale genome sequence
Communications Biology (2022)
-
Phycosphere pH of unicellular nano- and micro- phytoplankton cells and consequences for iron speciation
The ISME Journal (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.