Abstract
A few commonly used non-antibiotic drugs have recently been associated with changes in gut microbiome composition, but the extent of this phenomenon is unknown. Here, we screened more than 1,000 marketed drugs against 40 representative gut bacterial strains, and found that 24% of the drugs with human targets, including members of all therapeutic classes, inhibited the growth of at least one strain in vitro. Particular classes, such as the chemically diverse antipsychotics, were overrepresented in this group. The effects of human-targeted drugs on gut bacteria are reflected on their antibiotic-like side effects in humans and are concordant with existing human cohort studies. Susceptibility to antibiotics and human-targeted drugs correlates across bacterial species, suggesting common resistance mechanisms, which we verified for some drugs. The potential risk of non-antibiotics promoting antibiotic resistance warrants further exploration. Our results provide a resource for future research on drug–microbiome interactions, opening new paths for side effect control and drug repurposing, and broadening our view of antibiotic resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kåhrström, C. T., Pariente, N. & Weiss, U. Intestinal microbiota in health and disease. Nature 535, 47 (2016)
Forslund, K. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528, 262–266 (2015)
Imhann, F. et al. Proton pump inhibitors affect the gut microbiome. Gut 65, 740–748 (2016)
Jackson, M. A. et al. Proton pump inhibitors alter the composition of the gut microbiota. Gut 65, 749–756 (2016)
Rogers, M. A. & Aronoff, D. M. The influence of non-steroidal anti-inflammatory drugs on the gut microbiome. Clin. Microbiol. Infect. 22, 171–179 (2016)
Flowers, S. A., Evans, S. J., Ward, K. M., McInnis, M. G. & Ellingrod, V. L. Interaction between atypical antipsychotics and the gut microbiome in a bipolar disease cohort. Pharmacotherapy 37, 261–267 (2017)
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016)
Tramontano, M. et al. Nutritional preferences of the human gut bacteria reveal their metabolic idiosyncasies. Nat. Microbiol. https://doi.org/10.1038/s41564-018-0123-9 (2018)
Ejim, L. et al. Combinations of antibiotics and nonantibiotic drugs enhance antimicrobial efficacy. Nat. Chem. Biol. 7, 348–350 (2011)
Taber, H. W., Mueller, J. P., Miller, P. F. & Arrow, A. S. Bacterial uptake of aminoglycoside antibiotics. Microbiol. Rev. 51, 439–457 (1987)
Blaser, M. J. Antibiotic use and its consequences for the normal microbiome. Science 352, 544–545 (2016)
Rani, N., Sharma, A. & Singh, R. Imidazoles as promising scaffolds for antibacterial activity: a review. Mini Rev. Med. Chem. 13, 1812–1835 (2013)
Harbut, M. B. et al. Auranofin exerts broad-spectrum bactericidal activities by targeting thiol-redox homeostasis. Proc. Natl Acad. Sci. USA 112, 4453–4458 (2015)
Farha, M. A. et al. Antagonism screen for inhibitors of bacterial cell wall biogenesis uncovers an inhibitor of undecaprenyl diphosphate synthase. Proc. Natl Acad. Sci. USA 112, 11048–11053 (2015)
Pasolli, E., Truong, D. T., Malik, F., Waldron, L. & Segata, N. Machine learning meta-analysis of large metagenomic datasets: tools and biological insights. PLOS Comput. Biol. 12, e1004977 (2016)
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Bäckhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016)
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011)
Donaldson, G. P., Lee, S. M. & Mazmanian, S. K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016)
Hens, B., Brouwers, J., Corsetti, M. & Augustijns, P. Supersaturation and precipitation of posaconazole upon entry in the upper small intestine in humans. J. Pharm. Sci. 105, 2677–2684 (2016)
Bailey, C. J., Wilcock, C. & Scarpello, J. H. Metformin and the intestine. Diabetologia 51, 1552–1553 (2008)
Schloissnig, S. et al. Genomic variation landscape of the human gut microbiome. Nature 493, 45–50 (2013)
Kuhn, M., Letunic, I., Jensen, L. J. & Bork, P. The SIDER database of drugs and side effects. Nucleic Acids Res. 44, D1075–D1079 (2016)
Bodet, C. A., III, Jorgensen, J. H. & Drutz, D. J. Antibacterial activities of antineoplastic agents. Antimicrob. Agents Chemother. 28, 437–439 (1985)
Stringer, A. M., Gibson, R. J., Bowen, J. M. & Keefe, D. M. Chemotherapy-induced modifications to gastrointestinal microflora: evidence and implications of change. Curr. Drug Metab. 10, 79–83 (2009)
Sharma, S. & Singh, A. Phenothiazines as anti-tubercular agents: mechanistic insights and clinical implications. Expert Opin. Investig. Drugs 20, 1665–1676 (2011)
Morgan, A. P. et al. The antipsychotic olanzapine interacts with the gut microbiome to cause weight gain in mouse. PLoS One 9, e115225 (2014)
Li, X. Z., Plésiat, P. & Nikaido, H. The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin. Microbiol. Rev. 28, 337–418 (2015)
Nagy, E., Urbán, E., Nord, C. E. & ESCMID Study Group on Antimicrobial Resistance in Anaerobic Bacteria Antimicrobial susceptibility of Bacteroides fragilis group isolates in Europe: 20 years of experience. Clin. Microbiol. Infect. 17, 371–379 (2011)
Cacace, E., Kritikos, G. & Typas, A. Chemical genetics in drug discovery. Curr. Op. Syst. Biol. 4, 35–42 (2017)
Morita, Y. et al. NorM, a putative multidrug efflux protein, of Vibrio parahaemolyticus and its homolog in Escherichia coli. Antimicrob. Agents Chemother. 42, 1778–1782 (1998)
Sulavik, M. C. et al. Antibiotic susceptibility profiles of Escherichia coli strains lacking multidrug efflux pump genes. Antimicrob. Agents Chemother. 45, 1126–1136 (2001)
Nichols, R. J. et al. Phenotypic landscape of a bacterial cell. Cell 144, 143–156 (2011)
Nasie, I., Steiner-Mordoch, S. & Schuldiner, S. New substrates on the block: clinically relevant resistances for EmrE and homologues. J. Bacteriol. 194, 6766–6770 (2012)
Ariza, R. R., Li, Z., Ringstad, N. & Demple, B. Activation of multiple antibiotic resistance and binding of stress-inducible promoters by Escherichia coli Rob protein. J. Bacteriol. 177, 1655–1661 (1995)
Gustafsson, C. & Persson, B. C. Identification of the rrmA gene encoding the 23S rRNA m1G745 methyltransferase in Escherichia coli and characterization of an m1G745-deficient mutant. J. Bacteriol. 180, 359–365 (1998)
Roldán, M. D., Pérez-Reinado, E., Castillo, F. & Moreno-Vivián, C. Reduction of polynitroaromatic compounds: the bacterial nitroreductases. FEMS Microbiol. Rev. 32, 474–500 (2008)
Matthews, D. A. et al. Dihydrofolate reductase: x-ray structure of the binary complex with methotrexate. Science 197, 452–455 (1977)
Clemente, J. C. et al. The microbiome of uncontacted Amerindians. Sci. Adv. 1, e1500183 (2015)
Spanogiannopoulos, P., Bess, E. N., Carmody, R. N. & Turnbaugh, P. J. The microbial pharmacists within us: a metagenomic view of xenobiotic metabolism. Nat. Rev. Microbiol. 14, 273–287 (2016)
Kritikos, G. et al. A tool named Iris for versatile high-throughput phenotyping in microorganisms. Nat. Microbiol. 2, 17014 (2017)
Rettedal, E. A., Gumpert, H. & Sommer, M. O. Cultivation-based multiplex phenotyping of human gut microbiota allows targeted recovery of previously uncultured bacteria. Nat. Commun. 5, 4714 (2014)
Goodman, A. L. et al. Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proc. Natl Acad. Sci. USA 108, 6252–6257 (2011)
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010)
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012)
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012)
Nielsen, H. B. et al. Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nat. Biotechnol. 32, 822–828 (2014)
Mende, D. R., Sunagawa, S., Zeller, G. & Bork, P. Accurate and universal delineation of prokaryotic species. Nat. Methods 10, 881–884 (2013)
Kultima, J. R. et al. MOCAT2: a metagenomic assembly, annotation and profiling framework. Bioinformatics 32, 2520–2523 (2016)
Kruschke, J. K. Bayesian estimation supersedes the t test. J. Exp. Psychol. Gen. 142, 573–603 (2013)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995)
Kuhn, M. et al. STITCH 4: integration of protein-chemical interactions with user data. Nucleic Acids Res. 42, D401–D407 (2014)
Deghou, S. et al. CART-a chemical annotation retrieval toolkit. Bioinformatics 32, 2869–2871 (2016)
Mudie, D. M. et al. Quantification of gastrointestinal liquid volumes and distribution following a 240 mL dose of water in the fasted state. Mol. Pharm. 11, 3039–3047 (2014)
Law, V. et al. DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res. 42, D1091–D1097 (2014)
Kim, E. R. & Rhee, P. L. How to interpret a functional or motility test—colon transit study. J. Neurogastroenterol. Motil. 18, 94–99 (2012)
Pritchard, S. E. et al. Fasting and postprandial volumes of the undisturbed colon: normal values and changes in diarrhea-predominant irritable bowel syndrome measured using serial MRI. Neurogastroenterol. Motil. 26, 124–130 (2014)
Turnidge, J. & Paterson, D. L. Setting and revising antibacterial susceptibility breakpoints. Clin. Microbiol. Rev. 20, 391–408 (2007)
Campillos, M., Kuhn, M., Gavin, A. C., Jensen, L. J. & Bork, P. Drug target identification using side-effect similarity. Science 321, 263–266 (2008)
Otsuka, Y. et al. GenoBase: comprehensive resource database of Escherichia coli K-12. Nucleic Acids Res. 43, D606–D617 (2015)
Mori, H. et al. Identification of essential genes and synthetic lethal gene combinations in Escherichia coli K-12. Methods Mol. Biol. 1279, 45–65 (2015)
Wu, H. et al. Metformin alters the gut microbiome of individuals with treatment-naive type 2 diabetes, contributing to the therapeutic effects of the drug. Nat. Med. 23, 850–858 (2017)
Steinbeck, C. et al. Recent developments of the chemistry development kit (CDK) - an open-source java library for chemo- and bioinformatics. Curr. Pharm. Des. 12, 2111–2120 (2006)
Kim, S. et al. PubChem substance and compound databases. Nucleic Acids Res. 44, D1202–D1213 (2016)
Acknowledgements
We thank P. Beltrao (EBI), K. C. Huang (Stanford) and F. Cabreiro (UCL) for feedback on the manuscript; F. Rippmann (Merck KGaA) for pointing to the delayed onset of antipsychotics; S. Wicha (University of Hamburg) for discussions on drug concentrations; J. Overington (Medicines Discovery Catapult) for help with drug plasma concentrations, and members of all four laboratories for fruitful discussions (in particular T. Hodges for suggestions on the manuscript and M. Driessen for experimental support). We thank the EMBL mechanical workshop for the custom-made incubator. We acknowledge funding from EMBL and the Microbios grant (ERC-AdG-669830). L.M. and M.P. were supported by the EMBL Interdisciplinary Postdoc (EIPOD) programme under Marie Sklodowska Curie Actions COFUND (grant 291772). A.Te. and A.R.B. were supported by a Sofja Kovaleskaja Award of the Alexander von Humboldt Foundation to A.Ty.
Author information
Authors and Affiliations
Contributions
The study was conceived by K.R.P., P.B. and A.Ty., designed by L.M., M.P., G.Z., A.R.B. and A.Ty., and supervised by K.R.P., P.B. and A.Ty. In vitro screening was established by M.P. and performed by L.M., M.P., A.Te. and K.C.F. Follow-up and validation experiments were conducted by L.M., M.P. and E.E.A. H.D. and H.M. constructed and provided the Transbac library. Data preprocessing was performed by M.K. and G.Z.; statistical analyses by M.K.; data curation by L.M., M.K. and E.E.A.; data interpretation by L.M., M.P., M.K., G.Z., K.R.P., P.B. and A.Ty. L.M., M.K., G.Z., K.R.P., P.B. and A.Ty. wrote the manuscript with input from M.P. and A.R.B.; L.M., M.K. and G.Z. designed figures with input from K.R.P., P.B. and A.Ty. All authors approved the final version for publication.
Corresponding authors
Ethics declarations
Competing interests
EMBL has filed two patent applications on repurposing compounds identified in this study for the treatment of infections and for modulating the composition of the gut microbiome, and on the use of the in vitro model of the human gut microbiome to study the impact of xenobiotics (Tentative European patent application numbers EP 18156520 and EP 18155278, respectively). Authors L.M., M.P., M.K., G.Z., K.R.P., P.B. and A.T. are listed as inventors.
Additional information
Reviewer Information Nature thanks K. Lewis, H. B. Nielsen, G. Wright and R. Xavier for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Screen set-up and species selection.
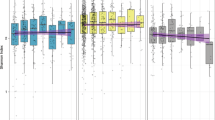
a, Drugs from the Prestwick Chemical Library (arranged in either 96- or 384-well format) were diluted in growth medium (usually mGAM) and pre-reduced in a Coy anaerobic chamber before inoculation with one of forty different human gut microbes. Bacterial growth was monitored for 16–24 h at 37 °C. Growth curves were acquired at least in triplicate for each drug–microbe interaction (see Methods). b, Species with a minimum relative abundance of 1% in at least one sample and a prevalence of 50% across samples (the latter estimated by rarefying to 10,000 reads mapping to taxonomic markers) were included in the set of core species. Boxplots show relative abundances of core species grouped by genus (according to NCBI taxonomy) and coloured by phylum (see key). The inner box indicates the IQR, with the median as black vertical line; the outer bars extend to the 5th and 95th percentiles; circles, outliers. To the right of the boxplots, prevalence is depicted by bars, and next to this the species diversity is shown; grey boxes indicate species represented in the screen with box widths corresponding to mean relative abundance within the genus. c, Relative abundance of genera of which at least one species was represented in the screen cumulates to 78% of the assignable fraction of reads (median across all samples, upper panel); first four boxplots show abundance within each study identified by country codes underneath (DK: Denmark; ES: Spain; US: United States; CN: China) 43,44,45,46. When directly cumulating the relative abundance of represented species the corresponding median is 60% (lower panel). Boxes span the IQR and whiskers extend to the most extreme data points up to a maximum of 1.5 times the IQR. d, Core species are shown in the order of their median abundances across all samples. Relative abundance boxplots and prevalence bars are defined as in b and grey boxes underneath indicate species screened in this study. Numbers in brackets correspond to specI cluster identifiers (version 1)47.
Extended Data Figure 2 Data analysis pipeline for identifying compounds with anticommensal activity.
a, Schematic overview of the data analysis pipeline. All steps (determination of time cutoff and removal of noisy points; normalization and selection of reference compounds; baseline correction and AUC calculation; and hit calling) are explained in detail in the Methods. On the first panel, dashed curves in the righthand plot depict the three possible effects that a drug can have on the growth of a microbe: increase the lag phase, decrease the growth rate or the stationary phase plateau. All effects are captured by cutting off the growth curves upon transition to stationary phase for most compounds (most drugs do not affect growth). On second panel, median growth rates for two drugs on same plate are depicted and normalized, whereas baseline correction (third panel) is applied at the individual wells. b, Growth curves (top, normalized OD) of Bacteroides ovatus in three exemplary drug cases for the three biological replicates (meclofenamic acid (red), moricizine (green) and diacerein (blue)). Light and dark grey shades represent the 50% and 90% confidence intervals for normal growth, respectively. Bottom, normalized AUC histograms for all drugs in the three biological replicates for B. ovatus. Meclofenamic acid is just below the hit threshold, moricizine is a hit with partial but strong growth inhibition, and diacerein almost completely inhibits the growth of B. ovatus. c, For most species, correlation between replicates is very high (median: 0.88). d, For both controls and reference compounds, P values were approximately uniformly distributed. Determining the background distribution of uninhibited growth using reference compounds is validated by their very similar behaviour with control wells. Other drugs (that is, drugs not used as reference compounds) show clear enrichment of low P values.
Extended Data Figure 3 Anticommensal activity relative to compound- and compartment-specific drug concentrations.
We made a simplified pharmacokinetic estimation of small intestine and colon concentrations by assuming that one dose of an orally administered drug (extracted from Drugs@FDA and Daily Defined Dose (DDD) of the ATC) reaches the intestine and is dissolved or absorbed similarly to the well-absorbed drug posaconazole19 (Supplementary Table 1). After absorption into the liver via the portal circulation, the drug enters circulation through the hepatic veins and reaches its characteristic plasma concentration. The two main routes of drug elimination are either secretion via kidneys and urine or secretion into the intestine via the biliary duct. In the intestine, drugs can be reabsorbed in a circuit called the enterohepatic cycle or excreted in stools. Compounds that are either poorly absorbed in the small intestine or secreted by bile reach the large intestine. Considering the measured excreted fraction of the drug in faeces (both changed and unchanged compound, as we do not know whether drug is metabolized in liver or gut), and assuming a large intestinal transit time of 24 h55 and a volume of distribution in the colon of 0.6 l56, we estimated the colon concentrations of the human-targeted drugs in our screen (Supplementary Table 1). Histograms for drug dose, plasma concentration, estimated small intestine concentration, urinary and fecal excretion and estimated colon concentration depict the respective distributions for human-targeted drugs, colour coded according to their anticommensal behaviour in our screen. Dashed lines indicate medians and vertical lines highlight the drug concentration used in our screen (20 μM). Interactions between drugs and microbiota are possible throughout the entire gastrointestinal tract, with microbial load having a gradient-like distribution (ileum and colon containing the largest numbers); this can be disturbed during disease18. In addition, drugs can be modified at several stages: by host digestive and intestinal epithelial enzymes, by phase I and phase II metabolism in the liver and by microbial enzymes. Some of these processes neutralize each other, resulting in reconversion into the original compound.
Extended Data Figure 4 Effects of metformin in gut microbiota in vivo correlate with its in vitro activity.
a, IC25s of the antidiabetic drug metformin for a selection of 22 strains. Metformin did not inhibit any species in our screen as the concentration used, 20 μM (red line), is below the IC25 of all strains. However, at its estimated small intestinal and colon concentration of 1.5 mM (blue line), metformin would inhibit 3 of 22 tested strains. This exemplifies that more human-targeted drugs would interfere with bacterial growth if doses were to be increased towards drug- and body-site-specific concentrations. b, IC25s of metformin correlate well with its observed effects in humans61, based on the four species that overlapped between the two studies. Significant treatment effects on the species level were mapped to our set of strains for which we had determined IC25s.
Extended Data Figure 5 Validation of the screen and conservative hit-calling.
a, b, Validation of our screen by IC25 determination for 25 selected drugs in a subset of up to 27 strains reveals high precision (94%) and recall (85%). We considered IC25 as the lowest concentration that reduces growth by at least 25% (see Methods). Breakdown into active and inactive compounds for drugs concentrations at the 20 μM concentration, used in our screen. True positives (TP), green; false positives (FP), red; true negatives (TN), grey; false negatives (FN), blue. c, Number of drugs with anticommensal activity versus the applied FDR threshold for all compounds (left) and human-targeted drugs (right). Increasing the FDR threshold from 0.01 to 0.1 (vertical grey lines) would nearly double the fraction of drugs that affect human gut microbes. d, IC25s of 25 drugs in up to 27 individual strains (see also a, b). The white area indicates the drug concentration range tested for each drug. Symbol sizes depict the number of strains with a particular IC25, symbol colours indicate categorization into false negative, false positive, true negative and true positive, and symbol shapes qualify whether actual IC25s were determined or IC25 was deemed to be higher or lower than the highest or lowest concentration tested, respectively. Vertical line indicates the drug concentration used in screen (20 μM). IC25s for all drug–strain pairs are listed in Supplementary Table 4. Particular drugs were responsible for false negatives in our screen (acarbose, loperamide, thioridazine), presumably owing to drug decay.
Extended Data Figure 6 IC25 relation to drug concentrations in human body.
For drug–strain pairs with measured IC25s (see also Extended Data Fig. 5), we compared IC25s with plasma and estimated small intestine and colon concentrations by plotting the number of strains that are affected in relation to whether they are above or below relevant body concentrations (colour code). With the exception of oestradiol valerate and 5-FU (only plasma concentrations available), all other drugs with available body concentrations reach concentrations high enough in the body to reach their IC25 for at least one gut microbial species (out of up to 27 species tested for IC25s).
Extended Data Figure 7 Concordance of drug in vitro species susceptibilities and drug-mediated shifts in microbiome composition of patients.
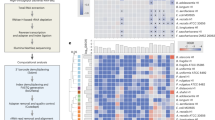
a, Association coefficients between PPI usage and relative taxonomical abundance in faecal microbiomes of PPI users from two studies (twins, UK cohort, green4; three independent cohorts from the Netherlands3, blue) (left) are compared to in vitro growth inhibition of isolates with same taxonomic rank in the presence of PPIs (omeprazole, lansoprazole and rabeprazole) as assessed by FDR-adjusted P values (q values) in our screen (right). Point size in the left panel corresponds to the q value as reported in the original study. Taxa that were reduced in patients (negative association coefficient, left of vertical black line) were mostly inhibited by PPIs in our screen (q value below 0.01, left of vertical black line), whereas enriched taxa were insensitive to PPIs. Box plots show: centre line, median; box limits, upper and lower quartiles; whiskers, 1.5× IQR; points, outliers. For fewer than 10 data points, all points are shown individually. b, Spearman correlation coefficients between association coefficients of faecal microbiome composition after consumption of amoxicillin or azathioprine7 and the screen P values. The histogram represents the background distribution of correlations between the in vitro data for all human-targeted drugs and the in vivo response to these drugs; correlations with amoxicillin or azathioprine are highlighted by triangles c, Comparisons between association coefficients and drugs from different therapeutic classes as assessed by Falony et al.7 and our in vitro data. d, A study of a cohort of patients with bipolar disease6 reported a significant decrease in abundance of Akkermansia upon treatment with atypical antipsychotics (AAP). When we compared distributions of adjusted P values from our screen for different strains, Akkermansia muciniphila was significantly more sensitive than all other strains to antipsychotics in general and APP in particular (P = 0.02 and P = 0.09, two-sided Wilcoxon rank sum test). By contrast, A. muciniphila is relatively more resistant than other strains across all human-targeted drugs (P = 0.0005, two-sided Wilcoxon rank sum test). Violin plot shows estimated density of points with the estimated median as vertical bar. For fewer than 10 data points, all points are also shown individually.
Extended Data Figure 8 Evaluation of anticommensal activity predictions based on side-effects.
a, IC25s of 26 candidate compounds (P value for enrichment of antibiotic-related side effects <1 × 10−4, using a one-sided Fisher’s exact test) and 16 control compounds (see also d of this Figure) were determined for 18 representative strains; results are depicted as an IC25 heatmap. Drugs are ordered according to their similarity in side effects to antibiotics from left to right (for antibiotic-related side effects see Supplementary Table 5). Qualifiers indicate whether IC25s are higher or lower than the indicated concentration; if no symbol, the box depicts the exact IC25. If highest tested concentrations did not reduce growth of any of the tested strains, the compound was classified as inactive (for example, Topiramate). b, Dose of the tested compounds according to the Defined Daily Dose and Drugs@FDA databases (see also Supplementary Table 1). c, Based on a compound’s recommended dose and its median IC25 for different bacterial strains, we estimated the number of doses need to reach this IC25. This number was plotted against the drug’s P value for enrichment of antibiotic-related side effects. For direct comparison between the two groups, see Fig. 3c. Circles in magenta depict drug–strain pairs for which growth was reduced, showing a clear correlation between P values and the estimated number of doses (magenta line). To rule out the possibility that the tested concentration range is causing this correlation, we also depict the estimated number of doses corresponding to the highest tested concentration (grey line), which exhibits no clear dependency between P value and number of doses. A vertical line across all panels connects all parameters attributable to a particular drug. d, Recommended single drug doses of human-targeted drugs with no anticommensal activity in our screen plotted against enrichment in antibiotic-related side effects (n = 339). Candidate and control drugs selection for testing for anticommensal activity at higher concentrations were selected on the basis of similarity to antibiotic-related side effects (vertical black line depicts prediction threshold) and aiming at drugs used at higher doses than concentration in our screen (horizontal dashed line). Purple and dark grey triangles indicate hits and non-hits from this validation effort, respectively. e, Ratios between IC25 and estimated colon concentrations are significantly lower (P = 0.017, two-sided Wilcoxon rank sum test) for candidate drugs than for control drugs. For candidate drugs, 16 of 52 (31%) IC25s were below the estimated colon concentrations while for control drugs this fraction was only 5 of 50 (10%). Box plots show: centre line, median; box limits, upper and lower quartiles; whiskers, 1.5× IQR; points, outliers.
Extended Data Figure 9 Drug therapeutic classes with anticommensal activity.
a, Fraction of drugs with anticommensal activity by ATC indication area (bars). All first-level indication areas and significantly enriched lower levels are shown (see also Extended Data Fig. 10). Significance (P value, one-sided Fischer’s exact test) is controlled for multiple hypothesis testing (Benjamini–Hochberg) independently at each ATC hierarchy level. b, Heat map of anticommensal activity and chemical similarities of human-targeted drugs within the three significantly ATC indication levels from a (indicated by circled numbers). Colours represent the median of drug pairwise Spearman correlations within and between subgroups depicted, calculated from the growth profiles of the 40 strains in each drug (P values) or their chemical similarity (Tanimoto scores62). Examples of structurally similar (phenothiazines; N05AA-AC) and diverse (N05AF-AX) antipsychotics that elicit similar responses in our screen are marked. c, Antipsychotics exhibit higher similarity in gut microbes they target than that expected on the basis of their structural similarity (P = 2 × 10−19 estimated from random permutations; other classes depicted show no significance difference). Box plots show: centre line, median; box limits, upper and lower quartiles; whiskers, 1.5× IQR; points, outliers. Notches correspond roughly to a 95% confidence interval for comparing medians.
Extended Data Figure 10 Drugs with anticommensal activity for all hierarchy levels of the ATC classification system.
Fraction of drugs with anticommensal activity for all indication areas of the ATC classification scheme with a high fraction of active compounds. Shown are indication areas that contain at least two active compounds and a fraction of at least 50% active compounds, their parent terms and all top-level indication areas. Significance (P value, one-sided Fischer’s exact test) is indicated by the bar colour and corrected for multiple hypothesis testing (Benjamini–Hochberg) independently at each hierarchy level of the ATC. Many smaller classes, including PPIs (A02BC), non-selective calcium channel blockers (C08E), synthetic oestrogens (G03CB), leukotriene receptor antagonists (R03DC) and phenothiazine and other antihistamines (R06AD and R06AX) are enriched, but owing to multiple testing and the small numbers of drugs tested in each group, they do not reach a significant P value.
Extended Data Figure 11 Comparing chemical similarity of drugs and similarity of hit profiles across gut microbes.
a, Heat map of anticommensal activity and chemical similarities for all active human-targeted drugs in our screen. Drugs are clustered according to chemical similarity. Colours represent the median of drug pairwise Spearman correlations within and between subgroups depicted, calculated from the growth profiles of the 40 strains in each drug (P values) or their chemical similarity (Tanimoto scores62). Several prominent groups are colour coded. Only drugs of some classes both share chemical similarity and have similar effects on the 40 strains—for example, phenothiazine antipsychotics and antihistamines (N05A and R06AD), structurally similar dibenzothiazepines and dibenzoxazepines for antipsychotics and antidepressants (N05AH and N06AA), PPIs (A02BC), anti-oestrogens (L02BA), synthetic oestrogens (G03CB) and anti-inflammatory fenamates (M01AG and M02AA06). b, A mild correlation exists between chemical similarity (Tanimoto scores) and anticommensal activity similarity (drug pairwise Spearman correlations): rs = 0.12 (P value of Spearman’s test <2 × 10−16).
Extended Data Figure 12 More complex, bulkier and heavier human-targeted drugs are more effective against Gram-positive bacteria.
Fraction of inhibited Gram-positive (blue, n = 22) or Gram-negative (red, n = 18) strains per drug plotted against different chemical properties of the drugs. Chemical properties, such as complexity (based on atom types, symmetry, computed using the Bertz/Hendrickson/Ihlenfeldt formula), molecular weight, TPSA (an estimate of the area, in Å2), volume (in Å3) and XLogP (distribution coefficient that is a measure of differential solubility in octanol and water) were obtained from PubChem63. For each chemical property, we used a type II ANOVA to test for linear dependency between the fraction of affected species and the chemical property (slope). Additionally, we tested whether this dependency depended on the Gram stain (slope difference). It is possible that there is no significant slope without considering Gram stain, but that there is a significant difference between the slopes for the two Gram stains. Lines show a linear fit to the data, with 95% confidence intervals as shaded area.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion. (PDF 236 kb)
Supplementary Table 1
This file shows the compounds used in this study. Sheet 1 shows the compounds of the Prestwick Chemical Library, their assignment to therapeutic classes according to the ATC classification system, chemical and physical properties, single doses (which are half of the daily doses), plasma concentrations, estimated small intestine concentrations, route of excretion, estimated colon concentrations and conversion of the molar concentration of the screen (20 μM) into μg/ml. Sheet 2 shows the chemicals used in this study. (XLSX 201 kb)
Supplementary Table 2
This file contains the strains used in this study, their medium requirements and starting ODs for drug screening in 96- or 384-format. (XLSX 51 kb)
Supplementary Table 3
This file contains drugs with anticommensal activity in our screen. Sheet 1 shows the adjusted p-values for the impact of 1197 drugs on anaerobic growth of 40 human gut bacteria (see Methods). Sheet 2 shows the literature evidence and lowest measured MICs for antibacterial activity of the top 40 human-targeted drugs that affect more than 10 strains in our screen. (XLSX 496 kb)
Supplementary Table 4
This file contains IC25 determination which shows 25 selected drugs in a subset of up to 27 strains. (XLSX 61 kb)
Supplementary Table 5
This file contains the antibiotic-related side effects in human-targeted drugs (see Methods). (XLSX 49 kb)
Supplementary Table 6
This file contains the screen results of additionally screened strains (Figure 4), which shows the E. coli ΔtolC, its parental background (BW25113) and of B. fragilis HM-20 (NT5057) and B. uniformis HM-715 (NT5065). (XLSX 106 kb)
Rights and permissions
About this article
Cite this article
Maier, L., Pruteanu, M., Kuhn, M. et al. Extensive impact of non-antibiotic drugs on human gut bacteria. Nature 555, 623–628 (2018). https://doi.org/10.1038/nature25979
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25979
This article is cited by
-
Metagenomic gut microbiome analysis of Japanese patients with multiple chemical sensitivity/idiopathic environmental intolerance
BMC Microbiology (2024)
-
Promiscuous, persistent and problematic: insights into current enterococcal genomics to guide therapeutic strategy
BMC Microbiology (2024)
-
High-throughput transcriptomics of 409 bacteria–drug pairs reveals drivers of gut microbiota perturbation
Nature Microbiology (2024)
-
Current understanding of the Alzheimer’s disease-associated microbiome and therapeutic strategies
Experimental & Molecular Medicine (2024)
-
Co-selection for antibiotic resistance by environmental contaminants
npj Antimicrobials and Resistance (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.