Abstract
The initiation of eukaryotic DNA replication occurs in two discrete stages1: first, the minichromosome maintenance (MCM) complex assembles as a head-to-head double hexamer that encircles duplex replication origin DNA during G1 phase; then, ‘firing factors’ convert each double hexamer into two active Cdc45–MCM–GINS helicases (CMG) during S phase. This second stage requires separation of the two origin DNA strands and remodelling of the double hexamer so that each MCM hexamer encircles a single DNA strand. Here we show that the MCM complex, which hydrolyses ATP during double-hexamer formation2,3, remains stably bound to ADP in the double hexamer. Firing factors trigger ADP release, and subsequent ATP binding promotes stable CMG assembly. CMG assembly is accompanied by initial DNA untwisting and separation of the double hexamer into two discrete but inactive CMG helicases. Mcm10, together with ATP hydrolysis, then triggers further DNA untwisting and helicase activation. After activation, the two CMG helicases translocate in an ‘N terminus-first’ direction, and in doing so pass each other within the origin; this requires that each helicase is bound entirely to single-stranded DNA. Our experiments elucidate the mechanism of eukaryotic replicative helicase activation, which we propose provides a fail-safe mechanism for bidirectional replisome establishment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bell, S. P. & Labib, K. Chromosome duplication in Saccharomyces cerevisiae. Genetics 203, 1027–1067 (2016)
Coster, G., Frigola, J., Beuron, F., Morris, E. P. & Diffley, J. F. X. Origin licensing requires ATP binding and hydrolysis by the MCM replicative helicase. Mol. Cell 55, 666–677 (2014)
Kang, S., Warner, M. D. & Bell, S. P. Multiple functions for Mcm2–7 ATPase motifs during replication initiation. Mol. Cell 55, 655–665 (2014)
Ilves, I., Petojevic, T., Pesavento, J. J. & Botchan, M. R. Activation of the Mcm2–7 helicase by association with Cdc45 and GINS proteins. Mol. Cell 37, 247–258 (2010)
Georgescu, R. E. et al. Mechanism of asymmetric polymerase assembly at the eukaryotic replication fork. Nat. Struct. Mol. Biol. 21, 664–670 (2014)
Yeeles, J. T., Deegan, T. D., Janska, A., Early, A. & Diffley, J. F. X. Regulated eukaryotic DNA replication origin firing with purified proteins. Nature 519, 431–435 (2015)
Dean, F. B. & Hurwitz, J. Simian virus 40 large T antigen untwists DNA at the origin of DNA replication. J. Biol. Chem. 266, 5062–5071 (1991)
Lõoke, M., Maloney, M. F. & Bell, S. P. Mcm10 regulates DNA replication elongation by stimulating the CMG replicative helicase. Genes Dev. 31, 291–305 (2017)
Remus, D. et al. Concerted loading of Mcm2–7 double hexamers around DNA during DNA replication origin licensing. Cell 139, 719–730 (2009)
Li, N. et al. Structure of the eukaryotic MCM complex at 3.8 Å. Nature 524, 186–191 (2015)
Noguchi, Y. et al. Cryo-EM structure of Mcm2–7 double hexamer on DNA suggests a lagging-strand DNA extrusion model. Proc. Natl Acad. Sci. USA 114, E9529–E9538 (2017)
Abid Ali, F. et al. Cryo-EM structure of a licensed DNA replication origin. Nat. Commun. 8, 2241 (2017)
McGeoch, A. T., Trakselis, M. A., Laskey, R. A. & Bell, S. D. Organization of the archaeal MCM complex on DNA and implications for the helicase mechanism. Nat. Struct. Mol. Biol. 12, 756–762 (2005)
Costa, A. et al. DNA binding polarity, dimerization, and ATPase ring remodeling in the CMG helicase of the eukaryotic replisome. eLife 3, e03273 (2014)
Froelich, C. A., Kang, S., Epling, L. B., Bell, S. P. & Enemark, E. J. A conserved MCM single-stranded DNA binding element is essential for replication initiation. eLife 3, e01993 (2014)
Georgescu, R. et al. Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation. Proc. Natl Acad. Sci. USA 114, E697–E706 (2017)
Sun, J. et al. The architecture of a eukaryotic replisome. Nat. Struct. Mol. Biol. 22, 976–982 (2015)
Zhou, J. C. et al. CMG-Pol ε dynamics suggests a mechanism for the establishment of leading-strand synthesis in the eukaryotic replisome. Proc. Natl Acad. Sci. USA 114, 4141–4146 (2017)
Abid Ali, F. et al. Cryo-EM structures of the eukaryotic replicative helicase bound to a translocation substrate. Nat. Commun. 7, 10708 (2016)
Duderstadt, K. E., Chuang, K. & Berger, J. M. DNA stretching by bacterial initiators promotes replication origin opening. Nature 478, 209–213 (2011)
Douglas, M. E. & Diffley, J. F. X. Recruitment of Mcm10 to sites of replication initiation requires direct binding to the minichromosome maintenance (MCM) complex. J. Biol. Chem. 291, 5879–5888 (2016)
Robertson, P. D. et al. Domain architecture and biochemical characterization of vertebrate Mcm10. J. Biol. Chem. 283, 3338–3348 (2008)
Marahrens, Y. & Stillman, B. A yeast chromosomal origin of DNA replication defined by multiple functional elements. Science 255, 817–823 (1992)
Coster, G. & Diffley, J. F. X. Bidirectional eukaryotic DNA replication is established by quasi-symmetrical helicase loading. Science 357, 314–318 (2017)
Shore, D. & Baldwin, R. L. Energetics of DNA twisting. II. Topoisomer analysis. J. Mol. Biol. 170, 983–1007 (1983)
Zivanovic, Y., Goulet, I. & Prunell, A. Properties of supercoiled DNA in gel electrophoresis. The V-like dependence of mobility on topological constraint. DNA-matrix interactions. J. Mol. Biol. 192, 645–660 (1986)
Hsieh, T. S. & Wang, J. C. Thermodynamic properties of superhelical DNAs. Biochemistry 14, 527–535 (1975)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
van Heel, M., Harauz, G ., Orlova, E. V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996)
Acknowledgements
We thank K. Labib for anti-Psf1 antibody, G. Kelly (the Francis Crick Institute, Bioinformatics) for help with mathematical modelling, the Francis Crick Institute Fermentation Facility for cell production and L. Collinson, R. Carzaniga (the Francis Crick Institute, Electron Microscopy) and T. Pape (Electron Microscopy Centre, Imperial College) for electron microscopy support. This work was supported by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001065 and FC001066), the UK Medical Research Council (FC001065 and FC001066), and the Wellcome Trust (FC001065 and FC001066). This work was also funded by a Wellcome Trust Senior Investigator Award (106252/Z/14/Z) and a European Research Council Advanced Grant (669424-CHROMOREP) to J.F.X.D.
Author information
Authors and Affiliations
Contributions
All authors conceived the electron microscopy experiments; M.E.D. prepared the samples and F.A.A. performed the imaging. M.E.D. and J.F.X.D. conceived all other experiments, which were carried out by M.E.D. M.E.D. and J.F.X.D. wrote the paper with input from F.A.A. and A.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks A. Leschziner and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 CMG assembly and activation are separable steps.
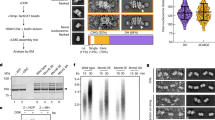
a, To determine when CMG assembly saturates, reactions were carried out on bead-immobilized ARS1 DNA and washed with high-salt buffer (HSW, buffer A + KCl) at the times indicated. The data show that no new CMG assembly takes place after 5 min. b, To confirm this, MCMs were loaded in parallel onto a bead-immobilized ARS1 DNA fragment and a soluble ARS1 plasmid, and phosphorylated with DDK. A firing factor mix, complete except for Mcm10, was added to the soluble reaction only, which was then added to the bead-immobilized MCMs at the times indicated after firing factor addition to the soluble reaction. After 8 min, beads were washed with high-salt buffer and bound proteins were analysed by immunoblotting. Psf1 signal relative to lane 2 is indicated. The experiment confirms that no CMG assembly takes place more than 5 min after firing factors have been added. c, To test whether Mcm10 can trigger DNA unwinding even after CMG assembly has finished, reactions were set up as in Fig. 1d, except Mcm10 was omitted until the times indicated after firing-factor addition. Mcm10 triggered robust unwinding, even when added more than 5 min after firing factors. Mcm10 can therefore activate preassembled CMG for DNA unwinding. d, To test whether Mcm10 can activate preassembled CMG for replication, CMG was assembled on an immobilized ARS1 plasmid with or without Mcm10. Beads were washed with low- (Buffer A + 0.25 M K-glutamate, LSW) or high-salt buffer, and replication proteins with or without Mcm10 and cofactors, including radiolabelled dCTP, were added. Mcm10 enabled DNA replication even when CMG had been washed to remove excess firing factors. e, Schematic outlining the CMG assembly and CMG activation steps described here.
Extended Data Figure 2 Characterization of DNA unwinding using small DNA circles.
a, Models of DNA unwinding with or without RPA. b, To define the relative positions of different topoisomers of radiolabelled 616-bp DNA circles containing ARS1 (used to analyse small changes in DNA supercoiling in the unwinding assay), nicked circles (nc, lane 1) were ligated closed in the indicated ethidium bromide (EthBr) concentrations. The supercoiling states of different bands of covalently closed DNA were determined relative to the ground state (α) by tracking the order in which bands peaked as ethidium bromide concentration increased and DNA was increasingly negatively supercoiled (see Methods for further details). Two bands peaked at the same position for α–5, and are likely to represent alternative configurations of the α–5 topoisomer. c, Primer extension reactions reading the T-rich strand of the ARS-consensus sequence (ACS) of ARS1 were carried out using 616-bp ARS1 DNA treated with potassium permanganate as indicated after CMG assembly in the absence of RPA. Reactions were separated on 5% sequencing gels, dried and analysed by autoradiography. Base pair numbering is relative to the 5′ end of the T-rich strand of the ACS. d, As Fig. 2c; lane 1 shows that MCM loading is required for all shifts in topoisomer distribution. Compared with other control samples, such as −DDK, topoisomer distribution was subtly different without MCM; this was not due to loading, which, as shown in Fig. 2b, does not affect topoisomer distribution. e, As Fig. 2a, except Mcm10 was omitted from all reactions. No proteins except Topo I were added to the reaction in lane 1 after MCM loading. There was no detectable change in supercoiling relative to when no firing factors (FF) were added (lane 1) when individual firing factors were omitted, suggesting that DNA untwisting in the absence of Mcm10 takes place during CMG assembly.
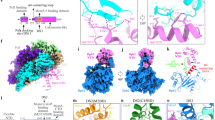
Extended Data Figure 3 Analysis of nucleotide binding and turnover by MCM.
a, Double hexamers assembled on bead-immobilized DNA using [α-32P]ATP were treated with DDK as indicated, and analysed by scintillation counting. Error bars, s.e.m.. b, Immunoblots of CMG-assembly reactions carried out as in Fig. 3d and washed with low-salt buffer. c, ATPase assays using [α-32P]ATP, single-MCM hexamers and Mcm10 as indicated were quantified after thin layer chromatography. Error bars, s.e.m.
Extended Data Figure 4 Characterization of replicative helicase activation using electron microscopy.
a, Examples of micrographs and complete sets of reference-free class averages of the indicated helicase activation reactions, washed with high-salt buffer (buffer A + KCl). In −DDK, +Mcm10: 7,410 of 23,092 total particles were double hexamers. In +DDK, +Mcm10: 14,668 and 10,492 of 43,320 total particles were CMG and double hexamers, respectively. In +DDK, −Mcm10: 3,984 and 2,226 of 12,920 total particles were CMG and double hexamers, respectively. Classes are positioned with respect to the abundance of source particles, with the most abundant class in the top left-hand corner, and abundance decreasing from left to right and from top to bottom. b, As a, with representative source micrographs. 5,032 of 6,815 and 2,049 of 20,904 particles were double hexamers when Dpb11 or Sld3–Sld7 were omitted, respectively. Scale bar, 100 nm. c, Comparison of CMG formed in the indicated conditions. d, as Fig. 4d. e, As Fig. 4e. Arrows, position of CMG. f, Representative crops from micrographs of the indicated samples. Arrows, position of MCM trains. Trains were not observed when either Mcm10 or the protein roadblock was omitted. Scale bar, 100 nm.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1, source images for all data obtained by electrophoretic separation. (PDF 879 kb)
Comparison of CMG and a CMG-polymerase epsilon complex with 2D projections of train ends
This video shows a comparison of CMG and a CMG-polymerase epsilon complex with 2D projections of train ends. (MP4 2089 kb)
Rights and permissions
About this article
Cite this article
Douglas, M., Ali, F., Costa, A. et al. The mechanism of eukaryotic CMG helicase activation. Nature 555, 265–268 (2018). https://doi.org/10.1038/nature25787
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25787
This article is cited by
-
Parental histone transfer caught at the replication fork
Nature (2024)
-
Quantity and quality of minichromosome maintenance protein complexes couple replication licensing to genome integrity
Communications Biology (2024)
-
TopBP1 utilises a bipartite GINS binding mode to support genome replication
Nature Communications (2024)
-
Genome maintenance meets mechanobiology
Chromosoma (2024)
-
Synergism between CMG helicase and leading strand DNA polymerase at replication fork
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.