Abstract
Farming was first introduced to Europe in the mid-seventh millennium bc, and was associated with migrants from Anatolia who settled in the southeast before spreading throughout Europe. Here, to understand the dynamics of this process, we analysed genome-wide ancient DNA data from 225 individuals who lived in southeastern Europe and surrounding regions between 12000 and 500 bc. We document a west–east cline of ancestry in indigenous hunter-gatherers and, in eastern Europe, the early stages in the formation of Bronze Age steppe ancestry. We show that the first farmers of northern and western Europe dispersed through southeastern Europe with limited hunter-gatherer admixture, but that some early groups in the southeast mixed extensively with hunter-gatherers without the sex-biased admixture that prevailed later in the north and west. We also show that southeastern Europe continued to be a nexus between east and west after the arrival of farmers, with intermittent genetic contact with steppe populations occurring up to 2,000 years earlier than the migrations from the steppe that ultimately replaced much of the population of northern Europe.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Tringham, R. E. in The Transition to Agriculture in Prehistoric Europe (ed. Price, D. ) 19–56 (Cambridge Univ. Press, 2000)
Bellwood, P. First Farmers: The Origins of Agricultural Societies 2nd edn (Wiley–Blackwell, 2004)
Golitko, M. in Ancient Europe, 8000 b.c. to a.d. 1000: An Encyclopedia of the Barbarian World (eds Bogucki, P. & Crabtree, P. J. ) 259–266 (Charles Scribners & Sons, 2003)
VanderLinden, M. in Investigating Archaeological Cultures: Material Culture, Variability, and Transmission (eds Roberts, B. W. & VanderLinden, M. ) 289–319 (Springer, 2012)
Bramanti, B. et al. Genetic discontinuity between local hunter-gatherers and central Europe’s first farmers. Science 326, 137–140 (2009)
Skoglund, P. et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012)
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015)
Cassidy, L. M. et al. Neolithic and Bronze Age migration to Ireland and establishment of the insular Atlantic genome. Proc. Natl Acad. Sci. USA 113, 368–373 (2016)
Hofmanová, Z. et al. Early farmers from across Europe directly descended from Neolithic Aegeans. Proc. Natl Acad. Sci. USA 113, 6886–6891 (2016)
Mathieson, I. et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528, 499–503 (2015)
Omrak, A. et al. Genomic evidence establishes Anatolia as the source of the European Neolithic gene pool. Curr. Biol. 26, 270–275 (2016)
Müller, J., Rassmann, K. & Videiko, M. Trypillia Mega-Sites and European Prehistory: 4100–3400 bce (Routledge, 2016)
Anthony, D. W. The Horse, the Wheel and Language (Princeton Univ. Press, 2007)
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015)
Allentoft, M. E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015)
Fu, Q. et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514, 445–449 (2014)
Fu, Q. et al. The genetic history of Ice Age Europe. Nature 534, 200–205 (2016)
Gallego Llorente, M. et al. Ancient Ethiopian genome reveals extensive Eurasian admixture in Eastern Africa. Science 350, 820–822 (2015)
Jones, E. R. et al. Upper Palaeolithic genomes reveal deep roots of modern Eurasians. Nat. Commun. 6, 8912 (2015)
Keller, A. et al. New insights into the Tyrolean Iceman’s origin and phenotype as inferred by whole-genome sequencing. Nat. Commun. 3, 698 (2012)
Kılınç, G. M. et al. The demographic development of the first farmers in Anatolia. Curr. Biol. 26, 2659–2666 (2016)
Lazaridis, I. et al. Genomic insights into the origin of farming in the ancient Near East. Nature 536, 419–424 (2016)
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014)
Olalde, I. et al. Derived immune and ancestral pigmentation alleles in a 7,000-year-old Mesolithic European. Nature 507, 225–228 (2014)
Raghavan, M. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014)
Lazaridis, I. et al. Genetic origins of the Minoans and Mycenaeans. Nature 548, 214–218 (2017)
Lipson, M. et al. Parallel palaeogenomic transects reveal complex genetic history of early European farmers. Nature 551, 368–372 (2017)
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016)
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009)
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012)
Gronenborn, D. & Dolukhanov, P. in The Oxford Handbook of Neolithic Europe (eds Fowler, C., Harding, J. & Hofmann, D. ) 195–214 (Oxford Univ. Press, 2015)
Telegin, D. Y. Neolithic cultures of Ukraine and adjacent areas and their chronology. J. World Prehist. 1, 307–331 (1987)
Telegin, D. Y. & Potekhina, I. D. Neolithic Cemeteries and Populations in the Dnieper Basin (British Archaeological Reports, 1987)
Jones, E. R. et al. The Neolithic transition in the Baltic was not driven by admixture with early European farmers. Curr. Biol. 27, 576–582 (2017)
Mittnik, A. et al. The genetic prehistory of the Baltic Sea region. Nat. Commun. 9, 443 (2018)
Saag, L. et al. Extensive farming in Estonia started through a sex-biased migration from the Steppe. Curr. Biol. 27, 2185–2193.e6 (2017)
Maier, A. The Central European Magdalenian: Regional Diversity and Internal Variability (Springer, 2015)
Boric´, D. & Price, T. D. Strontium isotopes document greater human mobility at the start of the Balkan Neolithic. Proc. Natl Acad. Sci. USA 110, 3298–3303 (2013)
Krauß, R., Marinova, E., De Brue, H. & Weninger, B. The rapid spread of early farming from the Aegean into the Balkans via the Sub-Mediterranean–Aegean vegetation Zone. Quat. Int. https://doi.org/10.1016/j.quaint.2017.01.019 (2017)
Bacvarov, K. in Moments in Time: Papers Presented to Pál Raczky on his 60th Birthday (eds Anders, A. & Kulcsár, G. ) 29–34 (L’Harmattan, 2013)
Gurova, M. & Bonsall, C. ‘Pre-Neolithic’ in Southeast Europe: a Bulgarian perspective. Documenta Praehistorica 41, 95–109 (2014)
Brandt, G. et al. Ancient DNA reveals key stages in the formation of central European mitochondrial genetic diversity. Science 342, 257–261 (2013)
Borić, D. in The Oxford Handbook of Neolithic Europe (eds Fowler C., Harding, J. & Hofmann, D. ) 927–957 (Oxford Univ. Press, 2015)
Szmyt, M. in Transition to the Bronze Age (Archaeolingua 30) (eds Heyd, V., Kulcsár, G. & Szeverényi, V. ) 93–111 (Archaeolingua, 2013)
Olalde, I. et al. A common genetic origin for early farmers from Mediterranean Cardial and central European LBK cultures. Mol. Biol. Evol. 32, 3132–3142 (2015)
Posth, C. et al. Pleistocene mitochondrial genomes suggest a single major dispersal of non-Africans and a Late Glacial population turnover in Europe. Curr. Biol. 26, 827–833 (2016)
Anthony, D. W. & Ringe, D. The Indo-European homeland from linguistic and archaeological perspectives. Annu. Rev. Linguist. 1, 199–219 (2015)
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl Acad. Sci. USA 110, 15758–15763 (2013)
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. https://doi.org/10.1101/pdb.prot5448 (2010)
Rohland, N., Harney, E., Mallick, S., Nordenfelt, S. & Reich, D. Partial uracil–DNA–glycosylase treatment for screening of ancient DNA. Phil. Trans. R. Soc. Lond. B 370, 20130624 (2015)
Korlević, P. et al. Reducing microbial and human contamination in DNA extractions from ancient bones and teeth. Biotechniques 59, 87–93 (2015)
Briggs, A. W. et al. Removal of deaminated cytosines and detection of in vivo methylation in ancient DNA. Nucleic Acids Res. 38, e87 (2010)
DeAngelis, M. M., Wang, D. G. & Hawkins, T. L. Solid-phase reversible immobilization for the isolation of PCR products. Nucleic Acids Res. 23, 4742–4743 (1995)
Rohland, N. & Reich, D. Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res. 22, 939–946 (2012)
Maricic, T., Whitten, M. & Pääbo, S. Multiplexed DNA sequence capture of mitochondrial genomes using PCR products. PLoS ONE 5, e14004 (2010)
Kircher, M., Sawyer, S. & Meyer, M. Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res. 40, e3 (2012)
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010)
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013)
Skoglund, P. et al. Separating endogenous ancient DNA from modern day contamination in a Siberian Neandertal. Proc. Natl Acad. Sci. USA 111, 2229–2234 (2014)
Korneliussen, T. S., Albrechtsen, A. & Nielsen, R. ANGSD: analysis of next generation sequencing data. BMC Bioinformatics 15, 356 (2014)
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006)
Petkova, D., Novembre, J. & Stephens, M. Visualizing spatial population structure with estimated effective migration surfaces. Nat. Genet. 48, 94–100 (2016)
Brown, L. D., Cai, T. T. & DasGupta, A. Interval estimation for a binomial proportion. Stat. Sci. 16, 101–133 (2001)
Agresti, A. & Coull, B. A. Approximate is better than ‘exact’ for interval estimation of binomial proportions. Am. Stat. 52, 119–126 (1998)
Longin, R. New method of collagen extraction for radiocarbon dating. Nature 230, 241–242 (1971)
Brown, T. A., Nelson, D. E., Vogel, J. S. & Southon, J. R. Improved collagen extraction by modified Longin method. Radiocarbon 30, 171–177 (1988)
Kennett, D. J. et al. Archaeogenomic evidence reveals prehistoric matrilineal dynasty. Nat. Commun. 8, 14115 (2017)
van Klinken, G. J. Bone collagen quality indicators for palaeodietary and radiocarbon measurements. J. Archaeol. Sci. 26, 687–695 (1999)
Stuiver, M. & Polach, H. A. Discussion: reporting of 14C data. Radiocarbon 19, 355–363 (1977)
Bronk Ramsey, C. OxCal 4.23 Online Manualhttps://c14.arch.ox.ac.uk/oxcalhelp/hlp_contents.html (2013)
Cook, G. T. et al. A freshwater diet-derived 14C reservoir effect at the Stone Age sites in the Iron Gates gorge. Radiocarbon 43, 453–460 (2001)
Acknowledgements
We thank D. Anthony, I. Lazaridis and M. Lipson for comments on the manuscript, B. Llamas, A. Cooper and A. Furtwängler for contributions to laboratory work, R. Evershed for contributing 14C dates and F. Novotny for assistance with samples. Support for this project was provided by the Human Frontier Science Program fellowship LT001095/2014-L to I.M., by DFG grant AL 287 / 14-1 to K.W.A.; by Irish Research Council grant GOIPG/2013/36 to D.F.; by the NSF Archaeometry program BCS-1460369 to D.J.K. for AMS 14C work; by MEN-UEFISCDI grant, Partnerships in Priority Areas Program – PN II (PN-II-PT-PCCA-2013-4-2302) to C.L.; by Croatian Science Foundation grant IP-2016-06-1450 to M.N. and I.J.; by European Research Council grant ERC CoG 724703 and Deutsche Forschungsgemeinschaft DFG FOR2237 to K.H.; by ERC starting grant ADNABIOARC (263441) to R.P.; and by US National Science Foundation HOMINID grant BCS-1032255, US National Institutes of Health grant GM100233, the Howard Hughes Medical Institute and an Allen Discovery Center grant from the Paul Allen Foundation to D.R.
Author information
Authors and Affiliations
Contributions
S.A.-R., A.S.-N., S.Vai., S.A., K.W.A., R.A., D.A., A.A., N.A., K.B., M.B.G., H.B., M.B., A.Bo., Y.B., A.Bu., J.B., S.C., N.J.C., R.C., M.C., C.C., D.G.D., N.E., M.Fr., B.Gal., G.G., B.Ge., T.Ha., V.H., K.H., T.Hi., S.I., I.J., I.Ka., D.Ko., A.K., D.La., M.La., C.L., M.Le., K.L., D.L.V., D.Lo., I.L., M.Ma., F.M., K.M., H.M., M.Me., P.M., V.M., V.P., T.D.P., A.Si., L.S., M.Š., V.S., P.S., A.St., T.S., M.T.-N., C.T., I.V., F.Va., S.Vas., F.Ve., S.Ve., E.V., B.V., C.V., J.Z., S.Z., P.W.S., G.C., R.K., D.C., G.Z., B.Gay., M.Li., A.G.N., I.P., A.P., D.B., C.B., J.K., R.P. and D.R. assembled and interpreted archaeological material. C.P., A.S.-N., N.R., N.B., F.C., O.C., D.F., M.Fe., B.Gam., G.G.F., W.H., E.H., E.J., D.Ke., B.K.-K., I.Ku., M.Mi., A.M., K.N., M.N., J.O., S.P., K.Si., K.St. and S.Vai. performed laboratory work. I.M., C.P., A.S.-N., S.M., I.O., N.P. and D.R. analysed data. D.J.K., S.T., D.B. and C.B. interpreted 14C dates. J.K., R.P. and D.R. supervised analysis or laboratory work. I.M. and D.R. wrote the paper with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks C. Renfrew, A. Scally and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
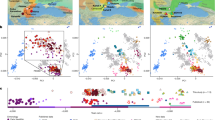
Extended Data Figure 1 Principal components analysis of ancient individuals.
Points for 486 ancient individuals are projected onto principal components defined by 777 present-day west Eurasian individuals (grey points). This differs from Fig. 1b in that the plot is not cropped and the present-day individuals are shown.
Extended Data Figure 2 Supervised ADMIXTURE analysis.
Supervised ADMIXTURE analysis modelling each ancient individual (one per row), as a mixture of populations represented by clusters that are constrained to contain northwestern-Anatolian Neolithic (grey), Yamnaya from Samara (yellow), EHG (pink) and WHG (green) populations. Dates in parentheses indicate approximate range of individuals in each population. This differs from Fig. 1d in that it contains some previously published samples7,9,10,19,23,26 and includes sample identification numbers.
Extended Data Figure 4 Genetic spatial structure in hunter-gatherers.
We infer the estimated effective migration surface62, a model of genetic relatedness in which individuals move in a random direction from generation to generation on an underlying grid, such that genetic relatedness is determined by distance. The migration parameter, m, defining the local rate of migration, varies on the grid and is inferred. This plot shows log10(m), scaled relative to the average migration rate, which is arbitrary. Thus log10(m) = 2, for example, implies that the rate of migration at this point on the grid is 100 times higher than average. To restrict the model as much as possible to hunter-gatherer populations, the migration surface is inferred using data from 116 individuals that date to earlier than approximately 5000 bc and have no northwestern-Anatolian-Neolithic-related ancestry. Although the migration surface is sensitive to sampling and fine-scale features may not be interpretable, the migration ‘barrier’ (region of low migration) running north-to-south and separating populations with primarily WHG ancestry from those with primarily EHG ancestry seems to be robust, and consistent with inferred admixture proportions. This analysis suggests that Mesolithic hunter-gatherer population structure was clustered and not smoothly clinal (that is, genetic differentiation did not vary consistently with distance). Superimposed on this background, pie charts show the WHG, EHG and CHG ancestry proportions inferred for populations used to construct the migration surface. This represents another way of visualizing the data in Fig. 2, Supplementary Table 3.1.3; we use two population models if they fit with P > 0.01, and three population models otherwise. Pie charts with only a single colour are those that were fixed to be the source populations.
Extended Data Figure 5 Sex bias in hunter-gatherer admixture.
The log-likelihood surfaces for the proportion of female (x axis) and male (y axis) ancestors that are hunter-gatherer-related for the combined populations analysed in Fig. 3c, and the two populations with the strongest evidence for sex bias. Numbers in parentheses, number of individuals in each group. The log-likelihood scale ranges from 0 to −10, in which 0 is the feasible point with the highest likelihood.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-3. (PDF 11029 kb)
Supplementary Data
This file contains Supplementary Tables 1-5. Supplementary Table 1 shows the details of ancient individuals analysed in this study, Supplementary Table 2 shows the key D-statistics to support statements about population history, Supplementary Table 3 shows the qpAdm models with 7-population outgroup set, Supplementary Table 4 shows the qpAdm models with extended 14-population outgroup set, Supplementary Table 5 shows the qpAdm models for Neolithic populations for chromosome X and Supplementary Table 6 shows the additional 14C dating information. (XLSX 268 kb)
Rights and permissions
About this article
Cite this article
Mathieson, I., Alpaslan-Roodenberg, S., Posth, C. et al. The genomic history of southeastern Europe. Nature 555, 197–203 (2018). https://doi.org/10.1038/nature25778
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25778
This article is cited by
-
The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes
Scientific Data (2024)
-
The Persian plateau served as hub for Homo sapiens after the main out of Africa dispersal
Nature Communications (2024)
-
Accurate detection of identity-by-descent segments in human ancient DNA
Nature Genetics (2024)
-
A complex subsistence regime revealed for Cucuteni–Trypillia sites in Chalcolithic eastern Europe based on new and old macrobotanical data
Vegetation History and Archaeobotany (2024)
-
100 ancient genomes show repeated population turnovers in Neolithic Denmark
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.