Abstract
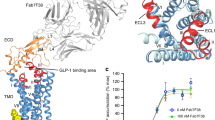
The class B glucagon-like peptide-1 (GLP-1) G protein-coupled receptor is a major target for the treatment of type 2 diabetes and obesity1. Endogenous and mimetic GLP-1 peptides exhibit biased agonism—a difference in functional selectivity—that may provide improved therapeutic outcomes1. Here we describe the structure of the human GLP-1 receptor in complex with the G protein-biased peptide exendin-P5 and a Gαs heterotrimer, determined at a global resolution of 3.3 Å. At the extracellular surface, the organization of extracellular loop 3 and proximal transmembrane segments differs between our exendin-P5-bound structure and previous GLP-1-bound GLP-1 receptor structure2. At the intracellular face, there was a six-degree difference in the angle of the Gαs–α5 helix engagement between structures, which was propagated across the G protein heterotrimer. In addition, the structures differed in the rate and extent of conformational reorganization of the Gαs protein. Our structure provides insights into the molecular basis of biased agonism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
de Graaf, C . et al. Glucagon-like peptide-1 and its class B G protein-coupled receptors: a long march to therapeutic successes. Pharmacol. Rev. 68, 954–1013 (2016)
Zhang, Y. et al. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 546, 248–253 (2017)
Hager, M. V., Clydesdale, L., Gellman, S. H., Sexton, P. M. & Wootten, D. Characterization of signal bias at the GLP-1 receptor induced by backbone modification of GLP-1. Biochem. Pharmacol. 136, 99–108 (2017)
Koole, C. et al. Allosteric ligands of the glucagon-like peptide 1 receptor (GLP-1R) differentially modulate endogenous and exogenous peptide responses in a pathway-selective manner: implications for drug screening. Mol. Pharmacol. 78, 456–465 (2010)
Wootten, D. et al. Differential activation and modulation of the glucagon-like peptide-1 receptor by small molecule ligands. Mol. Pharmacol. 83, 822–834 (2013)
Zhang, H. et al. Autocrine selection of a GLP-1R G-protein biased agonist with potent antidiabetic effects. Nat. Commun. 6, 8918 (2015)
Mann, R. et al. Peptide binding at the GLP-1 receptor. Biochem. Soc. Trans. 35, 713–716 (2007)
Manandhar, B. & Ahn, J. M. Glucagon-like peptide-1 (GLP-1) analogs: recent advances, new possibilities, and therapeutic implications. J. Med. Chem. 58, 1020–1037 (2015)
Liang, Y. L. et al. Phase-plate cryo-EM structure of a class B GPCR–G-protein complex. Nature 546, 118–123 (2017)
Danev, R., Buijsse, B., Khoshouei, M., Plitzko, J. M. & Baumeister, W. Volta potential phase plate for in-focus phase contrast transmission electron microscopy. Proc. Natl Acad. Sci. USA 111, 15635–15640 (2014)
Khoshouei, M., Radjainia, M., Baumeister, W. & Danev, R. Cryo-EM structure of haemoglobin at 3.2 Å determined with the Volta phase plate. Nat. Commun. 8, 16099 (2017)
Khoshouei, M. et al. Volta phase plate cryo-EM of the small protein complex Prx3. Nat. Commun. 7, 10534 (2016)
Siu, F. Y. et al. Structure of the human glucagon class B G-protein-coupled receptor. Nature 499, 444–449 (2013)
Hollenstein, K. et al. Structure of class B GPCR corticotropin-releasing factor receptor 1. Nature 499, 438–443 (2013)
Jazayeri, A. et al. Crystal structure of the GLP-1 receptor bound to a peptide agonist. Nature 546, 254–258 (2017)
Runge, S., Thøgersen, H., Madsen, K., Lau, J. & Rudolph, R. Crystal structure of the ligand-bound glucagon-like peptide-1 receptor extracellular domain. J. Biol. Chem. 283, 11340–11347 (2008)
Wootten, D. et al. The extracellular surface of the GLP-1 receptor is a molecular trigger for biased agonism. Cell 165, 1632–1643 (2016)
Coopman, K. et al. Residues within the transmembrane domain of the glucagon-like peptide-1 receptor involved in ligand binding and receptor activation: modelling the ligand-bound receptor. Mol. Endocrinol. 25, 1804–1818 (2011)
Dods, R. L. & Donnelly, D. The peptide agonist-binding site of the glucagon-like peptide-1 (GLP-1) receptor based on site-directed mutagenesis and knowledge-based modelling. Biosci. Rep. 36, e00285 (2015)
Koole, C. et al. Second extracellular loop of human glucagon-like peptide-1 receptor (GLP-1R) has a critical role in GLP-1 peptide binding and receptor activation. J. Biol. Chem. 287, 3642–3658 (2012)
Wootten, D. et al. Key interactions by conserved polar amino acids located at the transmembrane helical boundaries in class B GPCRs modulate activation, effector specificity and biased signalling in the glucagon-like peptide-1 receptor. Biochem. Pharmacol. 118, 68–87 (2016)
Yang, D. et al. Structural determinants of binding the seven-transmembrane domain of the glucagon-like peptide-1 receptor (GLP-1R). J. Biol. Chem. 291, 12991–13004 (2016)
Moon, M. J. et al. Ligand binding pocket formed by evolutionarily conserved residues in the glucagon-like peptide-1 (GLP-1) receptor core domain. J. Biol. Chem. 290, 5696–5706 (2015)
Wootten, D. et al. A hydrogen-bonded polar network in the core of the glucagon-like peptide-1 receptor is a fulcrum for biased agonism: lessons from class B crystal structures. Mol. Pharmacol. 89, 335–347 (2016)
Wootten, D., Simms, J., Miller, L. J., Christopoulos, A. & Sexton, P. M. Polar transmembrane interactions drive formation of ligand-specific and signal pathway-biased family B G protein-coupled receptor conformations. Proc. Natl Acad. Sci. USA 110, 5211–5216 (2013)
Furness, S. G. B. et al. Ligand-dependent modulation of G protein conformation alters drug efficacy. Cell 167, 739–749.e11 (2016)
Gregorio, G. G. et al. Single-molecule analysis of ligand efficacy in β2AR-G-protein activation. Nature 547, 68–73 (2017)
Cleator, J. H., Mehta, N. D., Kurtz, D. T. & Hildebrandt, J. D. The N54 mutant of Gαs has a conditional dominant negative phenotype which suppresses hormone-stimulated but not basal cAMP levels. FEBS Lett. 443, 205–208 (1999)
Lee, E., Taussig, R. & Gilman, A. G. The G226A mutant of Gsα highlights the requirement for dissociation of G protein subunits. J. Biol. Chem. 267, 1212–1218 (1992)
Iiri, T., Bell, S. M., Baranski, T. J., Fujita, T. & Bourne, H. R. A Gsα mutant designed to inhibit receptor signaling through Gs. Proc. Natl Acad. Sci. USA 96, 499–504 (1999)
Berlot, C. H. A highly effective dominant negative αs construct containing mutations that affect distinct functions inhibits multiple Gs-coupled receptor signaling pathways. J. Biol. Chem. 277, 21080–21085 (2002)
Berlot, C. H. & Bourne, H. R. Identification of effector-activating residues of Gsα. Cell 68, 911–922 (1992)
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005)
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017)
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D . et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Savage, E. E., Wootten, D., Christopoulos, A., Sexton, P. M. & Furness, S. G. A simple method to generate stable cell lines for the analysis of transient protein–protein interactions. Biotechniques 54, 217–221 (2013)
Hager, M. V. J., Johnson, L. M., Wootten, D., Sexton, P. M. & Gellman, S. H. β-Arrestin-biased agonists of the GLP-1 receptor from β-amino acid residue incorporation into GLP-1 analogues. J. Am. Chem. Soc. 138, 14970–14979 (2016)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Ehlert, F. J. The relationship between muscarinic receptor occupancy and adenylate cyclase inhibition in the rabbit myocardium. Mol. Pharmacol. 28, 410–421 (1985)
Acknowledgements
The work was supported by the Monash University Ramaciotti Centre for Cryo-Electron Microscopy, the National Health and Medical Research Council of Australia (NHMRC) project grants (1061044, 1065410, 1120919 and 1126857), NHMRC program grant (1055134), Strategic Priority Research Program of the Chinese Academy of Sciences (XDA12020347) and Shanghai Science and Technology Development Fund (15DZ2291600). P.M.S., A.C., D.W. and C.K. are NHMRC Principal Research, Senior Principal Research, Career Development and CJ Martin Fellows, respectively. S.L. received the Postgraduate Overseas Study Fellowship from CAS. We thank J. Plitzko, G. Christopoulos, V. Julita, J. Michaelis, X. Zhang, P. Thompson and M. Liu for assay and technical support and B. Kobilka for technical advice and comments on the manuscript.
Author information
Authors and Affiliations
Contributions
Y.-L.L. established the GLP-1R complex expression and purification strategy, expressed and purified the complex, and performed negative stain EM and data acquisition/analysis; Y.-L.L. and M.R. performed preliminary cryo-EM screening; M.K. performed cryo-sample preparation and phase plate imaging to acquire EM data and performed EM map calculations; A.G. built the model and performed refinement; A.G., C.K. and D.M.T. performed pharmacological assays; L.C., T.T.T. and S.L. performed the mutagenesis studies; S.G.B.F. and P.Z. designed and performed the G protein BRET assays; R.D. and W.B. organized and developed the Volta phase plate cryo-EM data acquisition strategy; Y.-L.L., M.K., A.G., S.G.B.F., P.Z., L.C., C.K., D.M.T., T.T.T., S.L., A.C., P.M.S. and D.W. performed data analysis; S.G.B.F., P.Z., C.K., A.C., L.J.M., M.-W.W. and A.C. assisted with data interpretation and preparation of the manuscript; Y.L.L., M.K., A.G., P.M.S. and D.W. interpreted data and wrote the manuscript; P.M.S. and D.W. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks R. Glaeser, F. Marshall and J. Mayer for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 GLP-1R pharmacology.
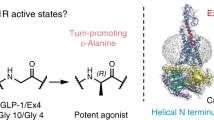
a, Peptide sequences. b, Pharmacology of untagged GLP-1R (WT GLP-1R) and the purification construct (HA–GLP-1R). c, Insect cell pharmacology of HA–GLP-1R. Top, radioligand competition binding. Bottom, GTPγS binding. Left, no Gs protein. ExP5 has lower affinity than GLP-1 and exendin-4 and does not bind GTPγS. Middle, wild-type Gs enhances peptide affinity and promotes GTPγS binding. Right, dominant-negative Gs is similar to wild-type Gs in binding, but does not bind GTPγS. d, Bias factors calculated from concentration–response curves using the Black and Leff operational model5,20,41 (see Methods) confirm that ExP5 is a biased agonist relative to GLP-1. e, Top left, pIC50 of ExP5 is ∼100-fold lower than of GLP-1 (CHOFlpIn whole cell). Top right, GLP-1 and ExP5 have β-arrestin1 coupling with pEC50 ∼30-fold to the right of their pIC50 (dotted lines). ExP5 is more potent than GLP-1 in cAMP signalling (pEC50 relative to pIC50). Bottom left, pIC50:pEC50 ratios for G protein (cAMP) and β-arrestin1 of ExP5 relative to GLP-1 highlights ExP5 bias arises from enhanced Gs coupling, not reduced β-arrestin1 recruitment. Bottom right, ratio of ExP5 efficacy (calculated using the Ehlert method44) relative to GLP-1 in cAMP and β-arrestin1 recruitment confirms that ExP5 bias arises from enhanced Gαs efficacy. Data in b, c are mean ± s.e.m. of three (insect cells) or four (CHOFlpIn cells) independent experiments, conducted in duplicate or triplicate, respectively. Data in d, e are from 11 independent experiments performed in duplicate. *P < 0.05 by one-way analysis of variance and Dunnett’s post-test.
Extended Data Figure 2 Purification and Volta phase plate imaging of the ExP5–GLP-1R–Gs complex.
a, Left, elution profile of the purified complex. Middle, pooled complex fractions, concentrated and analysed by size exclusion chromatography (SEC). Right, SDS–PAGE/Coomassie blue stain and western blot of the complex showing all components. Anti-His antibody detects Flag–GLP-1R–His, Gβ–His and Nb35–His (red) and anti-Gs antibody detects Gαs (green). b, Left, Volta phase plate micrograph of the complex (representative of 2,793). Middle, 2D class averages. Right; ‘gold standard’ Fourier shell correlation (FSC) curves; the overall nominal resolution is 3.26 Å. c, Left, Volta phase plate phase shift history throughout the dataset. Right, histogram of the estimated micrograph resolutions from the CTF.
Extended Data Figure 3 Atomic resolution model of the ExP5–GLP-1R–Gs heterotrimer in the cryo-EM density map.
EM density map and model are shown for all seven transmembrane helices and H8 of the receptor, the ExP5 peptide and the α5α helix of the GαSα Ras-like domain. Bulky residues are highlighted. All transmembrane helices exhibit good density, with TM6—which is flexible—being the least well-resolved.
Extended Data Figure 4 Comparison of class B GPCR structures.
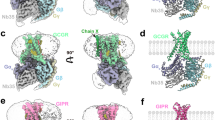
a–c, Agonist-bound full-length structures have distinct NTD orientations. d–f, Side view (d), extracellular view (e) and cytoplasmic view (f) of the conformational reorganization between inactive (GCGR, PDB 4L6R) and active structures (ExP5-bound GLP-1R). Distances are measured from Cα residues 1.33, 6.58, 7.35 and 6.35. Numbering uses the Wootten class B system. g–h, Superimposition of transmembrane domains from sCT–CTR–Gs (grey, PDB 5U27), GLP-1–GLP-1R–Gs (red, PDB 5VAI) and ExP5–GLP-1R–Gs with the inactive GCGR (green, PDB 4L6R). The largest differences in active structures relative to the inactive GCGR occur in TM1, TM6, TM7 and ECL3 (h), but the nature and extent of conformational change varies.
Extended Data Figure 5 ECL3, TM7, TM1 and ICL3 may be associated with GLP-1R biased agonism.
a, Conformational differences in GLP-1R ECL3 between ExP5-bound (blue) and GLP-1-bound (red) GLP-1R structures are supported by density in their respective cryo-EM maps. b, L3887.43A affected the potency of GLP-1 mediated cAMP more than ExP5 (mean + s.e.m. of four independent experiments). c, Right, TM1 overlays from agonist-bound class B GPCR structures reveals a different conformation for GLP-1–GLP-1R. Left, TM1 model overlays of ExP5–GLP-1R and GLP-1–GLP-1R with their associated cryo-EM maps (GLP-1, red ribbon/mesh; ExP5, blue ribbon/surface) reveals limited differences in the TM1 backbone, but potentially distinct side-chain orientations. d, Left, ICL3 backbone conformation in GLP-1–GLP-1R (PDB 5VAI) is supported by density (EMD-3653). Limited density is observed for ICL3 (337–343) in ExP5–GLP-1R.
Extended Data Figure 6 Rearrangement of conserved networks upon GLP-1R binding to ExP5.
Comparison of conserved networks in the inactive (green, GCGR) and activated (blue, ExP5–GLP-1R–Gs) states; central polar network (cyan), cytoplasmic polar networks (orange) and hydrophobic residues (pink). Inactive state interactions are incompatible with peptide binding and reorganize on activation. Upper middle, major rearrangements within the hydrophobic network (top, inactive; bottom, activated); side chains involved in ground state stabilization in green, inactive and active state in pink and active state in blue. Lower left and lower right, reorganization of the central hydrogen bond network and cytoplasmic networks, respectively, where green is inactive and blue is active. Subscript, Wootten numbering. These conformational changes are detailed in Supplementary Video 1.
Extended Data Figure 7 GLP-1R–G protein interactions.
a, GLP-1R forms interactions with GαSRas and Gβ. b–e, Receptor side chains (blue) within 4.5 Å of GαS side chains (gold) or Gβ side chains (cyan). b–d, GαSα5 forms polar and non-polar interactions with the cytoplasmic cavity formed by TM6 opening. Potential interactions also occur between GαSαN and ICL2 of GLP-1R. e, GLP-1R H8 aromatic residues embed within the detergent micelle and polar residues form direct interactions with Gβ. f, Left, the distinct engagement angle of GαSα5 with the receptor (Fig. 3) results in an overall rotation of the GαsRas,β,γ in ExP5–GLP-1R relative to GLP-1–GLP-1R. Right, overlaying Gαs from both structures reveals only minor differences in the G protein upon receptor engagement.
Supplementary information
Supplementary Table
This file contains Supplemental Data Table 1. Cryo-EM data collection, refinement and validation statistics. (PDF 96 kb)
Morph between inactive and active receptor rearrangement of conserved networks upon GLP-1R binding to ExP5.
Morph between inactive GLP-1R (homology model of the GLP-1R inactive TM generated using the closely related GCGR (4L6R) as a template) and the ExP5 active receptor reveals major rearrangement of the central hydrogen bond network, the hydrophobic network and the two ground state stabilising hydrogen bond networks (summarised in Ext. Data Figure 5) are associated with large scale conformational movements within the TM bundle (particularly TM6) upon receptor activation. (MP4 795 kb)
Rights and permissions
About this article
Cite this article
Liang, YL., Khoshouei, M., Glukhova, A. et al. Phase-plate cryo-EM structure of a biased agonist-bound human GLP-1 receptor–Gs complex. Nature 555, 121–125 (2018). https://doi.org/10.1038/nature25773
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25773
This article is cited by
-
Molecular features of the ligand-free GLP-1R, GCGR and GIPR in complex with Gs proteins
Cell Discovery (2024)
-
A framework for Frizzled-G protein coupling and implications to the PCP signaling pathways
Cell Discovery (2024)
-
Applications and prospects of cryo-EM in drug discovery
Military Medical Research (2023)
-
Intermediate-state-trapped mutants pinpoint G protein-coupled receptor conformational allostery
Nature Communications (2023)
-
Two-step structural changes in M3 muscarinic receptor activation rely on the coupled Gq protein cycle
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.