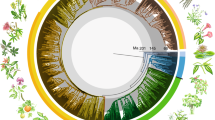
Abstract
High species diversity may result from recent rapid speciation in a ‘cradle’ and/or the gradual accumulation and preservation of species over time in a ‘museum’1,2. China harbours nearly 10% of angiosperm species worldwide and has long been considered as both a museum, owing to the presence of many species with hypothesized ancient origins3,4, and a cradle, as many lineages have originated as recent topographic changes and climatic shifts—such as the formation of the Qinghai–Tibetan Plateau and the development of the monsoon—provided new habitats that promoted remarkable radiation5. However, no detailed phylogenetic study has addressed when and how the major components of the Chinese angiosperm flora assembled to form the present-day vegetation. Here we investigate the spatio-temporal divergence patterns of the Chinese flora using a dated phylogeny of 92% of the angiosperm genera for the region, a nearly complete species-level tree comprising 26,978 species and detailed spatial distribution data. We found that 66% of the angiosperm genera in China did not originate until early in the Miocene epoch (23 million years ago (Mya)). The flora of eastern China bears a signature of older divergence (mean divergence times of 22.04–25.39 Mya), phylogenetic overdispersion (spatial co-occurrence of distant relatives) and higher phylogenetic diversity. In western China, the flora shows more recent divergence (mean divergence times of 15.29–18.86 Mya), pronounced phylogenetic clustering (co-occurrence of close relatives) and lower phylogenetic diversity. Analyses of species-level phylogenetic diversity using simulated branch lengths yielded results similar to genus-level patterns. Our analyses indicate that eastern China represents a floristic museum, and western China an evolutionary cradle, for herbaceous genera; eastern China has served as both a museum and a cradle for woody genera. These results identify areas of high species richness and phylogenetic diversity, and provide a foundation on which to build conservation efforts in China.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
McKenna, D. D. & Farrell, B. D. Tropical forests are both evolutionary cradles and museums of leaf beetle diversity. Proc. Natl Acad. Sci. USA 103, 10947–10951 (2006)
Moreau, C. S. & Bell, C. D. Testing the museum versus cradle tropical biological diversity hypothesis: phylogeny, diversification, and ancestral biogeographic range evolution of the ants. Evolution 67, 2240–2257 (2013)
Blackmore, S., Hong, D.-Y., Raven, P. H. & Wortley, A. H. in Plants of China: A Companion to the Flora of China (eds Hong, D.-Y. & Blackmore, S. ) 1–6 (Science Press, 2013)
Sun, G., Dilcher, D. L., Zheng, S.-L. & Zhou, Z.-K. In search of the first flower: a Jurassic angiosperm, Archaefructus, from northeast China. Science 282, 1692–1695 (1998)
Wen, J., Zhang, J.-Q., Nie, Z.-L., Zhong, Y. & Sun, H. Evolutionary diversifications of plants on the Qinghai–Tibetan Plateau. Front. Genet. 5, 4 (2014)
Xing, Y.-W. & Ree, R. H. Uplift-driven diversification in the Hengduan Mountains, a temperate biodiversity hotspot. Proc. Natl Acad. Sci. USA 114, E3444–E3451 (2017)
Klak, C., Reeves, G. & Hedderson, T. Unmatched tempo of evolution in Southern African semi-desert ice plants. Nature 427, 63–65 (2004)
Mishler, B. D. et al. Phylogenetic measures of biodiversity and neo- and paleo-endemism in Australian Acacia. Nat. Commun. 5, 4473 (2014)
Richardson, J. E., Pennington, R. T., Pennington, T. D. & Hollingsworth, P. M. Rapid diversification of a species-rich genus of neotropical rain forest trees. Science 293, 2242–2245 (2001)
Forest, F. et al. Preserving the evolutionary potential of floras in biodiversity hotspots. Nature 445, 757–760 (2007)
Verboom, G. A. et al. Origin and diversification of the Greater Cape flora: ancient species repository, hot-bed of recent radiation, or both? Mol. Phylogenet. Evol. 51, 44–53 (2009)
Thornhill, A. H. et al. Continental-scale spatial phylogenetics of Australian angiosperms provides insights into ecology, evolution and conservation. J. Biogeogr. 43, 2085–2098 (2016)
Donoghue, M. J. A phylogenetic perspective on the distribution of plant diversity. Proc. Natl Acad. Sci. USA 105, 11549–11555 (2008)
Svenning, J.-C., Eiserhardt, W. L., Normand, S., Ordonez, A. & Sandel, B. The influence of paleoclimate on present-day patterns in biodiversity and ecosystems. Annu. Rev. Ecol. Evol. Syst. 46, 551–572 (2015)
Wu, Z.-Y., Raven, P. H. & Hong, D.-Y. (eds) Flora of China, Vol. 1–25 (Science Press & Missouri Botanical Garden Press, 1994–2013)
Qian, H. & Ricklefs, R. E. A comparison of the taxonomic richness of vascular plants in China and the United States. Am. Nat. 154, 160–181 (1999)
López-Pujol, J., Zhang, F.-M., Sun, H.-Q., Ying, T.-S. & Ge, S. Centres of plant endemism in China: places for survival or for speciation? J. Biogeogr. 38, 1267–1280 (2011)
Qian, H. Environmental determinants of woody plant diversity at a regional scale in China. PLoS ONE 8, e75832 (2013)
Wang, Z.-H., Fang, J.-Y., Tang, Z.-Y. & Lin, X. Patterns, determinants and models of woody plant diversity in China. Proc. R. Soc. Lond. B 278, 2122–2132 (2011)
The Angiosperm Phylogeny Group. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 181, 1–20 (2016)
Soltis, D. E. et al. Angiosperm phylogeny: 17 genes, 640 taxa. Am. J. Bot. 98, 704–730 (2011)
Magallón, S., Gómez-Acevedo, S., Sánchez-Reyes, L. L. & Hernández-Hernández, T. A metacalibrated time-tree documents the early rise of flowering plant phylogenetic diversity. New Phytol. 207, 437–453 (2015)
Zanne, A. E. et al. Three keys to the radiation of angiosperms into freezing environments. Nature 506, 89–92 (2014)
Sun, X.-J. & Wang, P.-X. How old is the Asian monsoon system?—Palaeobotanical records from China. Palaeogeogr. Palaeoclimatol. Palaeoecol. 222, 181–222 (2005)
Favre, A. et al. The role of the uplift of the Qinghai–Tibetan Plateau for the evolution of Tibetan biotas. Biol. Rev. Camb. Philos. Soc. 90, 236–253 (2015)
Axelrod, D. I., Al-Shehbaz, I. A. & Raven, P. H. in Floristic Characteristics and Diversity of East Asian Plants (eds Zhang, A.-L. & Wu, S.-G. ) 43–55 (China Higher Education, 1996)
Benton, M. J. The Fossil Record 2 (Chapman & Hall, 1993)
Wu, Z.-Y., Sun, H., Zhou, Z.-K., Li, D.-Z. & Peng, H. Floristics of Seed Plants from China (Science Press, 2010)
Smith, S. A. & Beaulieu, J. M. Life history influences rates of climatic niche evolution in flowering plants. Proc. R. Soc. Lond. B 276, 4345–4352 (2009)
Zhang, Z.-J., He, J.-S., Li, J.-S. & Tang, Z.-Y. Distribution and conservation of threatened plants in China. Biol. Conserv. 192, 454–460 (2015)
Chen, Z.-D. et al. Tree of life for the genera of Chinese vascular plants. J. Syst. Evol. 54, 277–306 (2016)
Smith, S. A. & O’Meara, B. C. treePL: divergence time estimation using penalized likelihood for large phylogenies. Bioinformatics 28, 2689–2690 (2012)
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014)
Miller, M. A ., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Gateway Computing Environments Workshop, IEEE. http://ieeexplore.ieee.org/document/5676129/ (2010)
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012)
Britton, T., Anderson, C. L., Jacquet, D., Lundqvist, S. & Bremer, K. Estimating divergence times in large phylogenetic trees. Syst. Biol. 56, 741–752 (2007)
Anderson, C. L. Dating Divergence Times in Phylogenies. PhD thesis, Uppsala Univ. (2007)
R Core Team. R: a language and environment for statistical computing. http://R-project.org/ (2014)
Zachos, J., Pagani, M., Sloan, L., Thomas, E. & Billups, K. Trends, rhythms, and aberrations in global climate 65 Ma to present. Science 292, 686–693 (2001)
Smith, S. A. & Donoghue, M. J. Rates of molecular evolution are linked to life history in flowering plants. Science 322, 86–89 (2008)
Jetz, W. & Rahbek, C. Geographic range size and determinants of avian species richness. Science 297, 1548–1551 (2002)
Rahbek, C . et al. Predicting continental-scale patterns of bird species richness with spatially explicit models. Proc. R. Soc. Lond. B 274, 165–174 (2007)
Lennon, J. J., Koleff, P., Greenwood, J. J. D. & Gaston, K. J. Contribution of rarity and commonness to patterns of species richness. Ecol. Lett. 7, 81–87 (2004)
Iglewicz, B. & Hoaglin, D. How to Detect and Handle Outliers (ASQC Quality, 1993)
Rousseeuw, P. J. & Croux, C. Alternatives to the median absolute deviation. J. Am. Stat. Assoc. 88, 1273–1283 (1993)
Cleophas, T. J. Clinical trials: robust tests are wonderful for imperfect data. Am. J. Ther. 22, e1–e5 (2015)
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992)
Rodrigues, A. S. L., Brooks, T. M. & Gaston, K. J. in Phylogeny and Conservation (eds Purvis, A., Gittleman, J. L., & Brooks, T. ) 101–119 (Cambridge Univ. Press, 2005)
Webb, C. O., Ackerly, D. D., McPeek, M. A. & Donoghue, M. J. Phylogenies and community ecology. Annu. Rev. Ecol. Syst. 33, 475–505 (2002)
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. 25, 1965–1978 (2005)
Qian, H. & Jin, Y. An updated megaphylogeny of plants, a tool for generating plant phylogenies and an analysis of phylogenetic community structure. J. Plant Ecol. 9, 233–239 (2016)
Kuhn, T. S., Mooers, A. Ø. & Thomas, G. H. A simple polytomy resolver for dated phylogenies. Methods Ecol. Evol. 2, 427–436 (2011)
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012)
Acknowledgements
We thank J.-Y. Fang, D.-Z. Li, K.-P. Ma, S.-Z. Zhang, H. Sun, J.-Q. Liu, Z.-H. Wang, X.-Q. Wang and H.-Z. Kong for help initiating this study. This research was supported by the National Key Basic Research Program of China (2014CB954100), the National Natural Science Foundation of China (31590822), the Chinese Academy of Sciences International Institution Development Program (SAJC201613), the National Natural Science Foundation of China and US National Science Foundation Dimensions Collaboration Project (31461123001), the US National Science Foundation (Open Tree of Life: DEB-1207915, DEB-1208428; ABI DBI-1458466 and DBI-1458640; iDigBio: EF-1115210 and DBI-1547229; US–China Dimensions of Biodiversity: DEB-1442280) and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
Z.-D.C., P.S.S., D.E.S and J.-H.L. conceived the paper. L.-M.L., L.-F.M., T.Y., J.-F.Y., B.L., H.-L.L. and M.S. analysed the data. L.-M.L., L.-F.M., T.Y., J.-F.Y., B.L., J.T.M., S.M., P.S.S., D.E.S., J.-H.L. and Z.-D.C. wrote the first draft and finalized the manuscript. H.-H.H., Y.-T.N., D.-X.P., M.C., K.-L.X., C.-T.L. and V.-C.D. contributed data. J.T.M., A.-M.L., Y.-H.C., S.A.S., P.S.S., D.E.S., J.-H.L. and Z.-D.C. contributed substantially to revisions. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks R. Colwell, V. Savolainen and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Dated megaphylogeny of the Chinese angiosperms.
Major clades, including magnoliids, monocots, superrosids and superasterids, as well as the basal eudicot grade, are indicated with different colours. Divergence times were estimated using treePL.
Extended Data Figure 2 The 95% confidence intervals of divergence times and the Spearman’s rank correlation between our dating and those of recent publications.
a, b, Plots of divergence times and 95% confidence intervals (grey bars) for each family (a, n = 273) and genus (b, n = 2,909). The centre values are ages calculated based on the optimal maximum likelihood tree. c, Correlation of nodal ages between treePL and PATHd8 in this study (n = 5,863; r = 0.94, P = 0). d, Correlation of family ages between treePL and ref. 22 (n = 236, r = 0.68, P = 1.17 × 10−33). e, Correlation of family ages between treePL and ref. 23 (n = 257; r = 0.55, P = 4.54 × 10−22). f, Correlation of family ages between ref. 22 and ref. 23 (n = 235; r = 0.75, P = 2.11 × 10−43). The solid line is y = x.
Extended Data Figure 3 Number of angiosperm genera that originated during specified geological timespans.
Column with three colours shows the number of woody (grey), herbaceous (yellow) and mixed genera (light blue) that originated within a specific geological timespan. Number of woody genera, n = 995; number of herbaceous genera, n = 1,569; mixed genera (genera with both woody and herbaceous species), n = 101. The dashed line indicates the accumulated percentage of genera that have originated since the Early Cretaceous. Global temperature changes that have occurred since the Palaeogene are shown by the red curve (from ref. 39; reprinted with permission from AAAS). The x axis indicates the geological period and time in millions of years. The left y axis shows the total number of genera that have originated by any given time period; the right y-axis represents the accumulated percentage of genera that originated within a geological time period.
Extended Data Figure 4 Plot of divergence times of the Chinese angiosperm genera in each grid cell.
Mean and median values of the divergence times are indicated.
Extended Data Figure 5 Histograms and distribution of skewness and kurtosis for divergence times in each grid cell.
a–c, Range of skewness for all genera (a), woody genera (b) and herbaceous genera (c). d–f, Range of kurtosis (computed as the fourth standardized moment) for all genera (d), woody genera (e) and herbaceous genera (f). g–i, Spatial distribution of skewness for all genera (g), woody genera (h) and herbaceous genera (i). j–l, Spatial distribution of kurtosis for all genera (j), woody genera (k) and herbaceous genera (l). Skewness values in most grid cells are positive and around 1–2, which implies that divergence times of genera are slightly right-skewed (there are more young ages in each grid cell). Kurtosis values in most grid cells are within a range of 4–8, larger than the value (3) for a normal distribution, which implies that the distribution of divergence times has more extreme outliers than the normal distribution. For eastern China, kurtosis values of approximately 4 for all genera are consistent with grid cells having a range of divergence times—including very young and very old ages—as expected for an area that is both a cradle and a museum.
Extended Data Figure 6 Geographic patterns of median ages for the Chinese angiosperm genera.
a–i, Median ages for all genera, woody genera and herbaceous genera (from left to right), based on all sampled genera (a–c), the youngest 25% of genera (d–f), and the oldest 25% of genera (g–i) in each grid cell. j–l, Null-model test to identify recent (blue grid cells) and ancient (red grid cells) divergence centres for all genera (j), woody genera (k) and herbaceous genera (l). The analyses include 2,592 angiosperm genera (woody genera, n = 925; herbaceous genera, n = 1,501; genera with both woody and herbaceous species, n = 166). Maps adapted from National Administration of Surveying, Mapping and Geoinformation of China (http://www.sbsm.gov.cn; review drawing number: GS(2016)1576).
Extended Data Figure 7 Spatial distribution of MDTs based on geographic range-size quartiles and the youngest 25% and oldest 25% of genera in China.
a–d, MDT patterns of the first (a), second (b), third (c) and fourth quartiles (d) of the sampled Chinese angiosperm genera. The first, second, third and fourth quartiles range from the narrowest to the widest geographic distribution, and represent 0.6%, 3.5%, 13.7% and 82.1% of 1,409,239 records, respectively. The Spearman’s rank correlation coefficients between the overall MDT (including all genera) and MDT of the first, second, third and fourth geographic quartile are 0.12 (P = 1.46 × 10−3), 0.59 (P = 1.21 × 10−87), 0.43 (P = 2.51 × 10−43) and 0.99 (P = 0), respectively. e, MDT pattern of the youngest 25% of genera in China, showing that there are young genera in both western and eastern China. f, MDT pattern of the oldest 25% of genera in China, confirming that older genera mainly occur in eastern China. Maps adapted from National Administration of Surveying, Mapping and Geoinformation of China (http://www.sbsm.gov.cn; review drawing number: GS(2016)1576).
Extended Data Figure 8 Patterns of generic richness, phylogenetic diversity and phylogenetic structure for the Chinese angiosperm genera.
a–c, Richness for all genera (a), woody genera (b) and herbaceous genera (c). d–f, Phylogenetic diversity for all genera (d), woody genera (e) and herbaceous genera (f). g–i, SES-PD for all genera (g), woody genera (h) and herbaceous genera (i). j–l, NRI for all genera (j), woody genera (k) and herbaceous genera (l). m–o, NTI for all genera (m), woody genera (n) and herbaceous genera (o). The analyses include 2,592 angiosperm genera (woody genera, n = 925; herbaceous genera, n = 1,501; genera with both woody and herbaceous species, n = 166). Maps adapted from National Administration of Surveying, Mapping and Geoinformation of China (http://www.sbsm.gov.cn; review drawing number: GS(2016)1576).
Extended Data Figure 9 Patterns of species-level phylogenetic diversity for all Chinese angiosperms.
a–l, Observed phylogenetic diversity for all species (a, d, g, j), woody species (b, e, h, k) and herbaceous species (c, f, i, l) based on species trees 210, 30, 174 and 461 (species trees were randomly selected from 1,000 post-burn-in trees). m–o, SES-PD for all species (m), woody species (n) and herbaceous species (o) based on species tree 461. The analyses include 26,978 angiosperm species (woody, n = 10,169; herbaceous, n = 16,809). Phylogenetic diversity and SES-PD based on 10 species trees produce similar patterns; Spearman’s rank correlation coefficients, r > 0.99, P < 2.20 × 10−16. Maps adapted from National Administration of Surveying, Mapping and Geoinformation of China (http://www.sbsm.gov.cn; review drawing number: GS(2016)1576).
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Text and Supplementary References. (PDF 606 kb)
Rights and permissions
About this article
Cite this article
Lu, LM., Mao, LF., Yang, T. et al. Evolutionary history of the angiosperm flora of China. Nature 554, 234–238 (2018). https://doi.org/10.1038/nature25485
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25485
This article is cited by
-
Ensemble species distribution modeling and multilocus phylogeography provide insight into the spatial genetic patterns and distribution dynamics of a keystone forest species, Quercus glauca
BMC Plant Biology (2024)
-
Effects of climate and environmental heterogeneity on the phylogenetic structure of regional angiosperm floras worldwide
Nature Communications (2024)
-
Geographical patterns and determinants of insect biodiversity in China
Science China Life Sciences (2024)
-
The Sino-Himalayan flora evolved from lowland biomes dominated by tropical floristic elements
BMC Biology (2023)
-
Human activities and species biological traits drive the long-term persistence of old trees in human-dominated landscapes
Nature Plants (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.