Abstract
The cells of multicellular organisms receive extracellular signals using surface receptors. The extracellular domains (ECDs) of cell surface receptors function as interaction platforms, and as regulatory modules of receptor activation1,2. Understanding how interactions between ECDs produce signal-competent receptor complexes is challenging because of their low biochemical tractability3,4. In plants, the discovery of ECD interactions is complicated by the massive expansion of receptor families, which creates tremendous potential for changeover in receptor interactions5. The largest of these families in Arabidopsis thaliana consists of 225 evolutionarily related leucine-rich repeat receptor kinases (LRR-RKs)5, which function in the sensing of microorganisms, cell expansion, stomata development and stem-cell maintenance6,7,8,9. Although the principles that govern LRR-RK signalling activation are emerging1,10, the systems-level organization of this family of proteins is unknown. Here, to address this, we investigated 40,000 potential ECD interactions using a sensitized high-throughput interaction assay3, and produced an LRR-based cell surface interaction network (CSILRR) that consists of 567 interactions. To demonstrate the power of CSILRR for detecting biologically relevant interactions, we predicted and validated the functions of uncharacterized LRR-RKs in plant growth and immunity. In addition, we show that CSILRR operates as a unified regulatory network in which the LRR-RKs most crucial for its overall structure are required to prevent the aberrant signalling of receptors that are several network-steps away. Thus, plants have evolved LRR-RK networks to process extracellular signals into carefully balanced responses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
04 July 2018
In this Letter, the wrong version of the Supplementary Information was used; details of refs. 59-65 were missing, and some of the figure citations in the Supplementary Discussion were incorrect. This has been corrected online.
References
Belkhadir, Y., Yang, L., Hetzel, J., Dangl, J. L. & Chory, J. The growth–defense pivot: crisis management in plants mediated by LRR-RK surface receptors. Trends Biochem. Sci. 39, 447–456 (2014)
Jaillais, Y., Belkhadir, Y., Balsemão-Pires, E., Dangl, J. L. & Chory, J. Extracellular leucine-rich repeats as a platform for receptor/coreceptor complex formation. Proc. Natl Acad. Sci. USA 108, 8503–8507 (2011)
Özkan, E. et al. An extracellular interactome of immunoglobulin and LRR proteins reveals receptor–ligand networks. Cell 154, 228–239 (2013)
Braun, P. et al. An experimentally derived confidence score for binary protein–protein interactions. Nat. Methods 6, 91–97 (2009)
Sun, J., Li, L., Wang, P., Zhang, S. & Wu, J. Genome-wide characterization, evolution, and expression analysis of the leucine-rich repeat receptor-like protein kinase (LRR-RLK) gene family in Rosaceae genomes. BMC Genomics 18, 763 (2017)
Shiu, S. H. et al. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16, 1220–1234 (2004)
Hohmann, U., Lau, K. & Hothorn, M. The structural basis of ligand perception and signal activation by receptor kinases. Annu. Rev. Plant Biol. 68, 109–137 (2017)
Zipfel, C. & Oldroyd, G. E. Plant signalling in symbiosis and immunity. Nature 543, 328–336 (2017)
Soyars, C. L., James, S. R. & Nimchuk, Z. L. Ready, aim, shoot: stem cell regulation of the shoot apical meristem. Curr. Opin. Plant Biol. 29, 163–168 (2016)
Song, W., Han, Z., Wang, J., Lin, G. & Chai, J. Structural insights into ligand recognition and activation of plant receptor kinases. Curr. Opin. Struct. Biol. 43, 18–27 (2017)
Boutrot, F. & Zipfel, C. Function, discovery, and exploitation of plant pattern recognition receptors for broad-spectrum disease resistance. Annu. Rev. Phytopathol. 55, 257–286 (2017)
Santiago, J., Henzler, C. & Hothorn, M. Molecular mechanism for plant steroid receptor activation by somatic embryogenesis co-receptor kinases. Science 341, 889–892 (2013)
Sun, Y. et al. Structural basis for flg22-induced activation of the Arabidopsis FLS2–BAK1 immune complex. Science 342, 624–628 (2013)
Sun, Y. et al. Structure reveals that BAK1 as a co-receptor recognizes the BRI1-bound brassinolide. Cell Res. 23, 1326–1329 (2013)
Tang, J. et al. Structural basis for recognition of an endogenous peptide by the plant receptor kinase PEPR1. Cell Res. 25, 110–120 (2015)
Braun, P. Interactome mapping for analysis of complex phenotypes: insights from benchmarking binary interaction assays. Proteomics 12, 1499–1518 (2012)
Mukhtar, M. S. et al. Independently evolved virulence effectors converge onto hubs in a plant immune system network. Science 333, 596–601 (2011)
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis. thaliana. Science 301, 653–657 (2003)
Belkhadir, Y. et al. Brassinosteroids modulate the efficiency of plant immune responses to microbe-associated molecular patterns. Proc. Natl Acad. Sci. USA 109, 297–302 (2012)
Mott, G. A. et al. Genomic screens identify a new phytobacterial microbe-associated molecular pattern and the cognate Arabidopsis receptor-like kinase that mediates its immune elicitation. Genome Biol. 17, 98 (2016)
Shinohara, H., Mori, A., Yasue, N., Sumida, K. & Matsubayashi, Y. Identification of three LRR-RKs involved in perception of root meristem growth factor in Arabidopsis. Proc. Natl Acad. Sci. USA 113, 3897–3902 (2016)
Liu, W., Pellegrini, M. & Wang, X. Detecting communities based on network topology. Sci. Rep. 4, 5739 (2014)
Li, X. Q., Xing, T. & Du, D. Identification of top-ranked proteins within a directional protein interaction network using the PageRank algorithm: applications in humans and plants. Curr. Issues Mol. Biol. 20, 13–28 (2016)
Tian, L., Bashan, A., Shi, D. N. & Liu, Y. Y. Articulation points in complex networks. 8, 14223, doi:10.1038/ncomms14223 (2017)
Schwessinger, B. et al. Phosphorylation-dependent differential regulation of plant growth, cell death, and innate immunity by the regulatory receptor-like kinase BAK1. PLoS Genet. 7, e1002046 (2011)
Couto, D. & Zipfel, C. Regulation of pattern recognition receptor signalling in plants. Nat. Rev. Immunol. 16, 537–552 (2016)
McWilliam, H. et al. Analysis Tool Web Services from the EMBL-EBI. Nucleic Acids Res. 41, W597–W600 (2013)
Gou, X. et al. Genome-wide cloning and sequence analysis of leucine-rich repeat receptor-like protein kinase genes in Arabidopsis thaliana. BMC Genomics 11, 19 (2010)
Malo, N., Hanley, J. A., Cerquozzi, S., Pelletier, J. & Nadon, R. Statistical practice in high-throughput screening data analysis. Nat. Biotechnol. 24, 167–175 (2006)
Brideau, C., Gunter, B., Pikounis, B. & Liaw, A. Improved statistical methods for hit selection in high-throughput screening. J. Biomol. Screen. 8, 634–647 (2003)
Halter, T. et al. The leucine-rich repeat receptor kinase BIR2 is a negative regulator of BAK1 in plant immunity. Curr. Biol. 24, 134–143 (2014)
Eyüboglu, B. et al. Molecular characterisation of the STRUBBELIG-RECEPTOR FAMILY of genes encoding putative leucine-rich repeat receptor-like kinases in Arabidopsis thaliana. BMC Plant Biol. 7, 16 (2007)
Jordá, L. et al. ERECTA and BAK1 receptor like kinases interact to regulate immune responses in Arabidopsis. Front. Plant Sci. 7, 897 (2016)
Russinova, E. et al. Heterodimerization and endocytosis of Arabidopsis brassinosteroid receptors BRI1 and AtSERK3 (BAK1). Plant Cell 16, 3216–3229 (2004)
Roux, M. et al. The Arabidopsis leucine-rich repeat receptor-like kinases BAK1/SERK3 and BKK1/SERK4 are required for innate immunity to hemibiotrophic and biotrophic pathogens. Plant Cell 23, 2440–2455 (2011)
Hazak, O. et al. Perception of root-active CLE peptides requires CORYNE function in the phloem vasculature. EMBO Rep. 18, 1367–1381 (2017)
Meng, X. et al. Differential function of Arabidopsis SERK family receptor-like kinases in stomatal patterning. Curr. Biol. 25, 2361–2372 (2015)
Nodine, M. D., Yadegari, R. & Tax, F. E. RPK1 and TOAD2 are two receptor-like kinases redundantly required for Arabidopsis embryonic pattern formation. Dev. Cell 12, 943–956 (2007)
Wang, X. et al. IDL6-HAE/HSL2 impacts pectin degradation and resistance to Pseudomonas syringae pv tomato DC3000 in Arabidopsis leaves. Plant J. 89, 250–263 (2017)
Meng, X. et al. Ligand-induced receptor-like kinase complex regulates floral organ abscission in Arabidopsis. Cell Reports 14, 1330–1338 (2016)
Santiago, J. et al. Mechanistic insight into a peptide hormone signaling complex mediating floral organ abscission. Elife 5, e15075 (2016)
Nimchuk, Z. L. CLAVATA1 controls distinct signaling outputs that buffer shoot stem cell proliferation through a two-step transcriptional compensation loop. PLoS Genet. 13, e1006681 (2017)
Cole, S. J. & Diener, A. C. Diversity in receptor-like kinase genes is a major determinant of quantitative resistance to Fusarium oxysporum f.sp. matthioli. New Phytol. 200, 172–184 (2013)
Agusti, J., Lichtenberger, R., Schwarz, M., Nehlin, L. & Greb, T. Characterization of transcriptome remodeling during cambium formation identifies MOL1 and RUL1 as opposing regulators of secondary growth. PLoS Genet. 7, e1001312 (2011)
Xiao, D. et al. SENESCENCE-SUPPRESSED PROTEIN PHOSPHATASE directly interacts with the cytoplasmic domain of SENESCENCE-ASSOCIATED RECEPTOR-LIKE KINASE and negatively regulates leaf senescence in Arabidopsis. Plant Physiol. 169, 1275–1291 (2015)
Kang, Y. H. & Hardtke, C. S. Arabidopsis MAKR5 is a positive effector of BAM3-dependent CLE45 signaling. EMBO Rep. 17, 1145–1154 (2016)
Albert, I. et al. An RLP23–SOBIR1–BAK1 complex mediates NLP-triggered immunity. Nat. Plants 1, 15140 (2015)
Valon, C., Smalle, J., Goodman, H. M. & Giraudat, J. Characterization of an Arabidopsis thaliana gene (TMKL1) encoding a putative transmembrane protein with an unusual kinase-like domain. Plant Mol. Biol. 23, 415–421 (1993)
Tarutani, Y. et al. Molecular characterization of two highly homologous receptor-like kinase genes, RLK902 and RKL1, in Arabidopsis thaliana. Biosci. Biotechnol. Biochem. 68, 1935–1941 (2004)
Chang, C. et al. The TMK1 gene from Arabidopsis codes for a protein with structural and biochemical characteristics of a receptor protein kinase. Plant Cell 4, 1263–1271 (1992)
Wang, T. et al. A receptor heteromer mediates the male perception of female attractants in plants. Nature 531, 241–244 (2016)
Sun, W. et al. Probing the Arabidopsis flagellin receptor: FLS2–FLS2 association and the contributions of specific domains to signaling function. Plant Cell 24, 1096–1113 (2012)
Yeh, Y. H. & Panzeri, D. The Arabidopsis malectin-like/LRR-RLK IOS1 is critical for BAK1-dependent and BAK1-independent pattern-triggered immunity. Plant Cell 28, 1701–1721 (2016)
Igarashi, D., Tsuda, K. & Katagiri, F. The peptide growth factor, phytosulfokine, attenuates pattern-triggered immunity. Plant J. 71, 194–204 (2012)
Mosher, S. et al. The tyrosine-sulfated peptide receptors PSKR1 and PSY1R modify the immunity of Arabidopsis to biotrophic and necrotrophic pathogens in an antagonistic manner. Plant J. 73, 469–482 (2013)
Wang, J. et al. Allosteric receptor activation by the plant peptide hormone phytosulfokine. Nature 525, 265–268 (2015)
Postel, S. et al. The multifunctional leucine-rich repeat receptor kinase BAK1 is implicated in Arabidopsis development and immunity. Eur. J. Cell Biol. 89, 169–174 (2010)
Sakamoto, T. et al. The tomato RLK superfamily: phylogeny and functional predictions about the role of the LRRII-RLK subfamily in antiviral defense. BMC Plant Biol. 12, 229 (2012)
Acknowledgements
We thank A. Pasha for uploading the CSILRR interaction dataset to the Bio-Analytic Resource for Plant Biology, Y. Dagdas for comments on the manuscript, C. K. Garcia for the pECIA2/14 vectors, E. Özkan for his protocols, Y. Saijo, K. Tori and M. Butenko for the pepr1/2, erl2 and hsl2 mutant lines, respectively, and A. Bindeus for help with programming software. This work was supported by grants from the Austrian Academy of Sciences through the Gregor Mendel Institute (Y.B. and W.B.); the Natural Sciences and Engineering Research Council of Canada Discovery Grants to D.S.G. and D.D.; a Canada Research Chair in Plant-Microbe Systems Biology (D.D.) or Comparative Genomics (D.S.G.); and the Centre for the Analysis of Genome Evolution and Function (D.D. and D.S.G.). This research was also funded by the Gatsby Charitable Foundation (C.Z.) and the European Research Council (grant ‘PHOSPHinnATE’) (C.Z.). E.S.-L. is supported by a Hertha Firnberg Programme post-doctoral fellowship (T-947) from the FWF Austrian Science Fund. M.S. is supported by a post-doctoral fellowship (STE 2448/1) from the Deutsche Forschungsgemeinschaft (DFG). This work was supported by the National Science Foundation (IOS-1557796) to M.S.M. P.S.-B. acknowledges funding from the Austrian Federal Ministry of Science, Research & Economy, and the City of Vienna through the Vienna Biocenter Core Facilities (VBCF). We would like to thank P. Serrano Drozdowskyj, A. Aszodi and A. Gyenesei from the VBCF Biocomputing facility for developing the Platero software. We also thank the VBCF Plant Sciences facilities for the plant growth chambers.
Author information
Authors and Affiliations
Contributions
E.S.-L., G.A.M., K.G. and Y.B conceived and designed the experiments for the CSI screen. J.N. and A.L. cloned and expressed all the ECDs with inputs from E.S.-L., P.S.-B. and Y.B.; E.S.-L. performed all the ECD interaction assays. T.C.H. conceived, designed and performed the Y2H assays under the supervision of M.S.M.; G.A.M., E.S.-L. and K.P. characterized and tested all the T-DNA insertion lines under the supervision of D.D., D.S.G. and Y.B.; G.A.M., E.S.-L., K.P., N.W., K.G. and J.K. genotyped and bulked all the T-DNA insertion lines. G.A.M. analysed and implemented the computational and statistical analysis of all the data with inputs from D.S.G. and Y.B.; M.L. and G.A.M. conceived, designed and performed the network analysis with inputs from D.D., D.S.G., N.J.P. and Y.B.; M.S. designed and performed the BAK1–FLS2 co-immunoprecipitation assays under the supervision of C.Z.; K.P. organized and performed the APEX–PEPRs co-immunoprecipitation experiment with guidance from E.S.-L.; E.S.-L. and K.G. generated the apex bak1 double-mutant and 35S::APEX transgenic lines. S.B.S. and W.B. contributed and characterized the fir T-DNA mutant. E.S.-L. conceived, organized and performed the physiological assays with brassinosteroids, Pep2 and flg22. D.M. and M.M. devised the synthesis of the flg22 and Pep2 peptides. G.A.M. and Y.B. wrote the manuscript with major input from E.S.-L., M.S.M., C.Z., D.D. and D.S.G.; all authors commented and agreed on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The Gregor Mendel Institute, the University of Toronto and The Sainsbury Laboratory have filed a patent application on the use of the technical and computational approaches described in this work, in which E.S.-L., G.A.M., M.L., K.G., C.Z., D.D., D.S.G. and Y.B. are listed as inventors.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Expression profiles of LRR-RK ECDs produced as recombinant baits with the Drosophila S2 cells protein expression system.
a–o, Western blot analyses of raw supernatants from S2 cells transfected with ECD expression vectors. Blots were cropped and arranged to match the phylogenetic tree of the LRR-RK gene family. The family subclasses and Arabidopsis gene initiative (AGI) identifiers are indicated at the top. For lanes showing no obvious anti-V5 signals, a mild concentration of the S2 cell media and/or purification on protein-A-coated 96-well plates allowed for confirmation of expression and secretion of the ECDs. This experiment was conducted once with the full set of 200 ECDs. The expression of 130 independently expressed ECDs was tested one additional time with similar results.
Extended Data Figure 2 Calibration of the CSILRR screen conditions on ligand-dependent (FLS2–BAK1) and ligand-independent (BAK1–BIR4) interaction pairs.
a, b, Western blot analyses of raw supernatants from S2 cells transfected with prey and bait expression vectors for the ECD of FLS2 (bait, western blot: anti-V5 antibody; prey, western blot: anti-Flag antibody). S2 cells left untreated (−) or treated with CuSO4 (+). Days post transfection (dpt) are indicated on top. The experiment was repeated independently twice with similar results. c, Binding of the FLS2 ECD to the protein-A-coated 96-well plates. A fourfold dilution (4×) of the insect cell medium containing the ECD of FLS2 saturates the binding sites of protein-A-coated wells as indicated by immunoblots of the flow-through (FT). The experiment was repeated independently twice with similar results. d–f, As in a–c but for BAK1. The experiment was repeated independently twice with similar results. g, Plate interaction assays between the ECDs of BAK1 (prey) and FLS2 (bait) represented as cumulative absorbance (Abs 650 nm) over 18 h. Dots represent individual observations at each hour from five technical replicates. Box plots display the first and third quartiles, split by the median (red line); whiskers extend to include the maximum and minimum values. The presence of flg22 (+) in fourfold-diluted CSILRR screening conditions weakly promotes the interaction between the two ECDs. h, Technical replicates and box plots are as in g, but with BAK1 (bait) and FLS2 (prey). i, Technical replicates and box plots are as in g but with BAK1 (prey eightfold diluted) and FLS2 (bait fourfold diluted). In these conditions, the binding between the ECDs of BAK1 and FLS2 is largely enhanced by the presence of flg22 (+), indicating that the proteins produced in our expression system can interact in a ligand-dependent manner and are thus functional. j, Technical replicates and box plots as in g, but using a prey variant of BAK1 that can no longer pentamerize owing to the deletion of the COMP domain (BAK1 mono-prey). Binding between the two ECDs is still observed, but at a reduced level, thus indicating the importance of the pentamerization motif for detecting transient and low affinity interactions in the absence of ligand. k, l, Binding of FLS2 and BAK1 ECDs to protein-A-coated 96-well plates (as indicated by immunoblots of the flow-through) when proteins are produced from S2 cells growing either at 21 °C or 27 °C. Immunoblots show a slight increase in protein production at 27 °C with similar binding capacities to the protein-A-coated plate. The protein expression levels at the two temperatures were assessed more than three times with similar results. The plate saturation experiment for proteins produced at 27 °C was conducted once. m, Plate interaction assays between BAK1 (prey) and FLS2 (bait) (in fourfold-diluted conditions) represented as cumulative absorbance (Abs 650 nm) over a 150-min time course. Dots represent individual observations made every 10 min from four technical replicates. Box plots as in g. Although slightly more abundant, proteins produced at 27 °C do not interact as well when produced at 21 °C. Protein expression for the CSILRR screen was performed at 21 °C. n, The FLS2–BAK1 interaction is insensitive to changes in pH conditions. Left, the interaction between FLS2 (bait) and BAK1 (prey) was observed in the pH range from 5.5 to 7.5. This experiment was conducted once. Right, plate interaction assays between BAK1 (prey) and FLS2 (bait) (in fourfold-diluted conditions) represented as cumulative absorbance over a 3-h time course. Dots represent individual observations at each hour from one technical replicate. The CSILRR screen was performed at the pH of the conditioned S2 cells supernatant (~pH 7.5). o, Plate interaction assays between BAK1 (as mono-prey (blue dots) or penta-prey (black dots)) and BIR4 represented as cumulative absorbance over a 3-h time course. Dots represent individual observations at each hour from one technical replicate. This experiment was conducted once. The data indicate that the pentamerization of the prey is a key requirement for enhancing the interaction detection sensitivity, without disrupting the functionality of the ECDs. BAK1 and BIR4 are ligand-independent interaction partners and the screening conditions used are also appropriate to detect this interaction.
Extended Data Figure 3 Comparison of the primary and retest screens parameters.
a, Geometric mean of the normalized absorbance values for the HCI (red dots) and LCI (yellow dots) obtained from the primary screen (CSI), the validation screen (retest) and the negative controls (NC) associated with the two screens. n denotes numbers of bidirectional interactions: HCI CSI (n = 567), HCI retest (n = 567), LCI CSI (n = 248), LCI retest (n = 248), and NC (n = 618). The box plots contain the first and third quartiles, split by the median (yellow or red lines indicated by the arrow on the left of the boxes); whiskers extend to include the maximum and minimum values. Statistical significance was determined using unbalanced one-way ANOVA by Tukey’s HSD for all pairwise comparisons. Datasets with the same letter are indistinguishable at >95% confidence. b, Plots of a linear regression for the entire set of normalized absorbance values obtained from the retest screens (absorbance retest; y axis) and the corresponding values from the from the primary screen (absorbance CSI; x axis). The thick, straight red line is the linear regression that best describes the entire set of data points (Spearman’s r = 0.7696; indicated on top). The fine red dashed lines represent the 95% confidence intervals of the regression. n = 815 bidirectional interactions. c, Comparison of the geometric mean of normalized absorbance values for selected interactions. Values from the primary screen (absorbance CSI; y axis) and the validation screen (absorbance retest; x axis) are shown for the LCI set (yellow dots) and for 20 interactions selected at random from the HCI set (red dots). The number of interactions shown for each set was selected to approach the numbers present in the entire interaction search space. The red lines show the absorbance values corresponding to the FLS2–BAK1 interaction in both screens. d, Retest assay performance parameters interpreted within the performance window measured by positive reference set (PRS) and LCI calibration. To estimate the reliability of the estimates provided by the retest, the observed rate of interactions found in the HCI and LCI sets were used for a Monte Carlo simulation. n = 100,000 independent sets of observations selected at random from these populations, with the number of observations equal to the number present in the retest sets. These values were used to calculate the mean and s.d. of the samplings, which are presented as error bars.
Extended Data Figure 4 Characterization of BRI1 interaction partners.
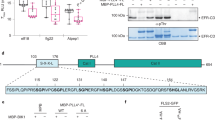
a, qPCR analyses showing altered gene expression in T-DNA lines targeting the interaction partners of BRI1 (Fig. 1b). Genotypes are indicated. Relative expression levels were calculated and ACTIN was used as reference gene to control for cDNA amount in each reaction. The box plots contain the first and third quartiles, split by the median; whiskers extend to include the maximum and minimum values. n = 4 biologically independent mRNA samples for all genotypes, except for bak1-4, skm1 and sobir1 where n = 3. Statistical significance was estimated by an unpaired two-sided t-test and is indicated on top of the boxes: erl2 *P = 0.0012, fir *P = 5.3508 × 10−6, bak1-4 *P = 3.08212 × 10−7, bam3 *P = 1.9378 × 10−5, serk4 *P = 0.0108, hsl2 *P = 2.06945 × 10−5, sark *P = 0.0259, rlk *P = 2.12971 × 10−10, rul1 *P = 7.49918 × 10−5, srf4 *P = 3.08212 × 10−7, skm1 *P = 5.5911 × 10−6, sobir1 *P = 0.0001. ns, not significant. b, T-DNA insertions targeting the HCI (top interactions) and LCI (bottom interaction) partners of BRI1. Morphology of representative seedlings grown for 7 days in the absence (NT) or presence (BL) of 500 nM brassinolide, the most potent brassinosteroid. Genotypes are indicated. The experiment was conducted six times with similar results.
Extended Data Figure 5 Characterization of FLS2 interaction partners.
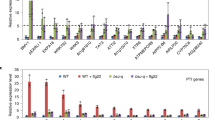
a, qPCR analyses showing altered gene expression in T-DNA lines targeting the interaction partners of FLS2 (Fig. 1c). Genotypes are indicated. Relative expression levels were calculated and ACTIN was used as reference gene to control for cDNA amount in each reaction. n = 9 biologically independent mRNA samples for all tested genotypes. Statistical significance was estimated by an unpaired two-sided t-test: mik1 *P = 8.17192 × 10−6, pskr1 *P = 0.007, pepr2 *P = 0.007, at3g14840 *P = 0.005, at2g01210 *P = 0.0032, pepr1 *P = 1.16519 × 10−5, fei2 *P = 0.005, nik3 *P = 0.0015. b, Oxidative burst represented as total photon counts, triggered by 1 μM flg22 in wild type (black) and mutant lines targeting the HCI (top; red) and LCI (bottom, yellow) partners for FLS2. Genotypes are indicated. Dots represent individual observations from four independent experiments. n denotes numbers of biologically independent leaf discs: WT (n = 36), mik1 (n = 36), fls2 (n = 28), pskr1 (n = 27), pepr2 (n = 38), at3g46350 (n = 39). Statistical significance was determined using linear mixed effect modelling, and symbols indicate the results of a post hoc unpaired two-sided t-test corrected with the Holm method for multiple testing: mik1 *P = 4.32 × 10−2, fls2 *P = 1 × 10−15. c, As in b, except: WT (n = 32), fls2 (n = 27), bak1 (n = 39), at3g14840 (n = 33), at2g01210 (n = 38), pepr1 (n = 40). bak1 *P = 1 × 10−15, fls2 *P = 1 × 10−15. d, As in b and c, except: WT (n = 43), fls2 (n = 29), bam3 (n = 33), fir (n = 39), srf9 (n = 32), fei2 (n = 45), nik3 (n = 32). fir *P = 1.38 × 10−3, fls2 *P = 1.2 × 10−15, nik3 *P = 1.38 × 10−3. The ROS burst assays in b–d were performed on independent plates (set number) and every plate contained wild type and fls2 controls, as well as randomly assigned mutant lines. e, flg22-induced peroxidase (POX) assay in wild-type (black bar) and mutant lines targeting the HCI (top interactions; red) and LCI (bottom interactions, yellow) partners for FLS2. Genotypes are indicated. Leaf disks from 4-week-old plants were treated with water (NT) or 1 μM flg22 (T). The level of flg22-induced POX was normalized to the corresponding non-treated control. The level of POX present in the wild type was set to 100 for easier interpretation. n denotes numbers of biologically independent leaf discs from two independent experiments: WT (n = 44), mik1 (n = 10), fls2 (n = 17), bak1 (n = 31), bam3 (n = 42), srf9 (n = 18), fir (n = 55), pskr1 (n = 24), pepr2 (n = 12), at3g46350 (n = 36), at3g14840 (n = 12), at2g01210 (n = 18), pepr1 (n = 12), fei2 (n = 11), nik3 (n = 15). Statistical significance was estimated using a paired two-sided t-test for each genotype, corrected for multiple tests using the Holm–Bonferroni correction. mik1 *P = 5.71 × 10−4, fls2 *P = 0.046, bak1 *P = 0.0039, fir *P = 0.0048, pskr1 *P = 9.49 × 10−5. All box plots contain the first and third quartiles, split by the median; whiskers extend to include the maximum and minimum values.
Extended Data Figure 6 FIR regulates flg22-induced responses.
a, Seedlings of the genotypes indicated on the bottom were treated with either water (NT) or flg22 (T) and changes in FRK1 transcript levels were quantified by qPCR analyses. Dots represent individual observations from three independent experiments. n denotes numbers of biologically independent mRNA samples: WT (n = 9 (NT), n = 9 (T)), fir (n = 9, n = 9) and fls2 (n = 6, n = 6). Statistical significance was determined using linear mixed effect modelling followed by comparison of each genotype to the wild-type control using unpaired two-sided t-test followed by multiple testing correction using the Holm method. fir *P = 1.42 × 10−7, fls2 *P = 4 × 10−16. b, Growth of Pto DC3000 on the genetic backgrounds indicated at the bottom of the chart. Four-week-old plants were infiltrated with 105 cfu ml−1 in the absence (black bars) or presence (grey bars) of 1 μM flg22. The number of bacteria per area of leaf (cfu ml−1) was plotted on a log10 scale for day 0 (open bars) and day 3 (closed bars). Dots represent individual observations from two independent experiments. n denotes numbers of samples, each including 4 biologically independent leaf discs. For day 0, WT (n = 6), fir (n = 6), fls2 (n = 6); for day 3, WT (n = 6), fir (n = 6), fls2 (n = 6); for day 3 + flg22, WT (n = 6), fir (n = 6), fls2 (n = 6). Statistical significance for bacterial growth was estimated by two-way ANOVA. A third experiment performed at an inoculum of 106 cfu ml−1 corroborated these results. c, Morphology of 7-day-old seedlings grown in the absence (−) or presence (+) of 1 μM flg22. Genotypes are indicated. The experiment was conducted twice with similar results. d, Primary root length (cm) from seedlings grown in the presence (T) or absence (NT) of 1 μM flg22. Fold changes are T/NT ratios. Dots represent individual observations from two independent experiments. n denotes the following numbers of biologically independent roots: WT (n = 32 (NT), n = 36 (T)), fir (n = 34 (NT), n = 32 (T)), fls2 (n = 27 (NT), n = 26 (T)). Statistical significance for two biological replicates was determined using linear mixed effect modelling followed by comparison of each genotype to the wild-type control using unpaired two-sided t-test followed by multiple testing correction using the Holm method. fir *P = 2.02 × 10−6, fls2 *P = 2.02 × 10−6. All box plots display the first and third quartiles, split by the median; whiskers extend to include the maximum and minimum values.
Extended Data Figure 7 CSILRR network representation and table of nodes with their corresponding identification numbers or acronyms.
The network construction and other features are the same as shown in Fig. 2b. The nodes surrounded by white halos are articulation points. The numbers in each node corresponding to the ECD of each LRR-RK are shown in the table.
Extended Data Figure 8 Characterization of independent apex mutant and 35S::APEX transgenic lines.
a, Top, rosette morphology of 4-week-old wild-type, apex-1 and apex-2, and apex-3 knockdown lines grown under long-day photoperiod at 22 °C. Genetic backgrounds are indicated. No obvious changes in rosette morphology are observed. The experiment was conducted three times with similar results. Bottom, qPCR analyses showing fold reduction of APEX transcripts in the independent mutant lines. Relative expression levels were calculated and ACTIN was used as reference control gene. Dots represent individual observations from three independent experiments. n = 9 biologically independent mRNA samples for each genotype. Statistical significance was determined using linear mixed effect modelling followed by comparison of each genotype to the wild-type control using unpaired two-sided t-test followed by multiple testing correction using the Holm method. apex-1 *P = 6 × 10−16, apex-2 *P = 5.33 × 10−15, apex-3 *P = 6 × 10−16. b, Top, rosette morphology of 3-week-old wild type and 35S::APEX lines 1 and 2 grown under long-day photoperiod at 22 °C. Genetic backgrounds are indicated on the top. Rosettes of 35S::APEX lines are slightly larger than WT under long-day photoperiod at 22 °C. The experiment was conducted three times with similar results. Middle: Quantitative real-time PCR analyses showing fold induction of the APEX transgene in the overexpression lines used in this study. Relative expression levels were calculated and ACTIN was used as reference gene to control for cDNA amount in each reaction. Dots represent individual observations from two independent experiments. n = 6 biologically independent mRNA samples for each genotype. Statistical significance was determined using linear mixed effect modelling followed by comparison of each genotype to the WT control using an unpaired two-sided t-test followed by multiple testing correction using the Holm method and is indicated on top of the boxes: 35S::APEX line 1 *P = 3.38 × 10−14, 35S::APEX line 2 *P = 7.77 × 10−14. Bottom, detection of APEX–YFP in stable transgenic T3 lines by western blot using an anti-GFP antibody. c, Modulation of BRI1 signalling by APEX gene dosage. Morphology of representative seedlings corresponding to Fig. 4a. Genotypes are indicated. The experiment was conducted over three times with similar results. d, Hypocotyl length ratios of seedlings grown in the presence (T) or absence (NT) of 500 nM brassinolide (BL). Genotypes are indicated. Dots represent individual observations from three independent experiments. n denotes numbers of biologically independent hypocotyls. WT (n = 43 (NT), n = 33 (T)), apex-1 (n = 31, n = 35), apex-2 (n = 32, n = 33), apex-3 (n = 39, n = 38), bri1 (n = 28, n = 32). Statistical significance was determined using linear mixed effect modelling followed by comparison of each genotype to the wild-type control using unpaired two-sided t-test followed by multiple testing correction using the Holm method. apex-1 *P = 2.53 × 10−14, apex-2 *P = 1.10 × 10−5, apex-3 *P = 1.55 × 10−12, bri1 *P = 8 × 10−16. e, flg22-induced oxidative bursts represented as total photon counts over 40 min. Genetic backgrounds are indicated. Dots represent individual observations from three independent experiments. n denotes numbers of biologically independent leaf discs: WT (n = 31), apex-1 (n = 19), apex-2 (n = 23), apex-3 (n = 25), fls2 (n = 15). Statistical significance was determined using linear mixed effect modelling followed by comparison of each genotype to the wild-type control using an unpaired two-sided t-test followed by multiple testing correction using the Holm method. apex-1 *P = 2.99 × 10−3, apex-2 *P = 2.84 × 10−2, apex-3 *P = 2.84 × 10−2, fls2 *P = 8 × 10−16. All box plots display the first and third quartiles, split by the median (red line); whiskers extend to include the maximum and minimum values.
Extended Data Figure 9 Modulation of brassinosteroid signalling by AT5G51560.
a, Morphology of representative seedlings grown for 7 days in the absence (NT) or presence (BL) of 500 nM brassinolide. Genotypes are indicated. The experiment was conducted twice with similar results. b, Hypocotyl length fold changes corresponding to a. Genotypes are indicated. Dots represent individual observations from two independent experiments. n denotes numbers of biologically independent hypocotyl: WT (n = 39 (NT), n = 29 (T)), at5g51560 line 1 (n = 36 (NT), n = 26 (T)), at5g51560 line 2 (n = 39 (NT), n = 34 (T)), bri1 (n = 25 (NT), n = 27 (T)). Box plots display the first and third quartiles, split by the median; whiskers extend to include the maximum and minimum values. Statistical significance was determined using linear mixed effect modelling followed by comparison of each genotype to the wild-type control using an unpaired two-sided t-test followed by multiple testing correction using the Holm method. at5g51560 line 1 *P = 3.75 × 10−6, at5g51560 line 2 *P = 2.26 × 10−12, bri1 *P = 6 × 10−16.
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion, full legends for Supplementary Tables 1-11, Supplementary Methods and Supplementary Figures 1-2. (PDF 4572 kb)
Supplementary Data
This zipped file contains Supplementary Tables 1-11 – see Supplementary Information document for full table descriptions. (ZIP 1079 kb)
Source data
Rights and permissions
About this article
Cite this article
Smakowska-Luzan, E., Mott, G., Parys, K. et al. An extracellular network of Arabidopsis leucine-rich repeat receptor kinases. Nature 553, 342–346 (2018). https://doi.org/10.1038/nature25184
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25184
This article is cited by
-
Marker and readout genes for defense priming in Pseudomonas cannabina pv. alisalensis interaction aid understanding systemic immunity in Arabidopsis
Scientific Reports (2024)
-
Phylogeny of the plant receptor-like kinase (RLK) gene family and expression analysis of wheat RLK genes in response to biotic and abiotic stresses
BMC Genomics (2023)
-
Changing turn-over rates regulate abundance of tryptophan, GS biosynthesis, IAA transport and photosynthesis proteins in Arabidopsis growth defense transitions
BMC Biology (2023)
-
A Phytophthora receptor-like kinase regulates oospore development and can activate pattern-triggered plant immunity
Nature Communications (2023)
-
Whole-Genome Resequencing Deciphers New Insight Into Genetic Diversity and Signatures of Resistance in Cultivated Cotton Gossypium hirsutum
Molecular Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.