Abstract
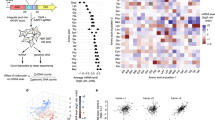
In addition to acting as template for protein synthesis, messenger RNA (mRNA) often contains sensory sequence elements that regulate this process1,2. Here we report a new mechanism that limits the number of complete protein molecules that can be synthesized from a single mRNA molecule of the human AMD1 gene encoding adenosylmethionine decarboxylase 1 (AdoMetDC). A small proportion of ribosomes translating AMD1 mRNA stochastically read through the stop codon of the main coding region. These readthrough ribosomes then stall close to the next in-frame stop codon, eventually forming a ribosome queue, the length of which is proportional to the number of AdoMetDC molecules that were synthesized from the same AMD1 mRNA. Once the entire spacer region between the two stop codons is filled with queueing ribosomes, the queue impinges upon the main AMD1 coding region halting its translation. Phylogenetic analysis suggests that this mechanism is highly conserved in vertebrates and existed in their common ancestor. We propose that this mechanism is used to count and limit the number of protein molecules that can be synthesized from a single mRNA template. It could serve to safeguard from dysregulated translation that may occur owing to errors in transcription or mRNA damage.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Gebauer, F. & Hentze, M. W. Molecular mechanisms of translational control. Nat. Rev. Mol. Cell Biol. 5, 827–835 (2004)
Sonenberg, N. & Hinnebusch, A. G. New modes of translational control in development, behavior, and disease. Mol. Cell 28, 721–729 (2007)
Pegg, A. E. S-Adenosylmethionine decarboxylase. Essays Biochem. 46, 25–46 (2009)
Chiang, P. K. et al. S-Adenosylmethionine and methylation. FASEB J. 10, 471–480 (1996)
Lu, S. C. & Mato, J. M. S-adenosylmethionine in liver health, injury, and cancer. Physiol. Rev. 92, 1515–1542 (2012)
Roje, S. S-Adenosyl-l-methionine: beyond the universal methyl group donor. Phytochemistry 67, 1686–1698 (2006)
Zhang, D. et al. AMD1 is essential for ESC self-renewal and is translationally down-regulated on differentiation to neural precursor cells. Genes Dev. 26, 461–473 (2012)
Scuoppo, C. et al. A tumour suppressor network relying on the polyamine–hypusine axis. Nature 487, 244–248 (2012)
Paasinen-Sohns, A. et al. Chaotic neovascularization induced by aggressive fibrosarcoma cells overexpressing S-adenosylmethionine decarboxylase. Int. J. Biochem. Cell Biol. 43, 441–454 (2011)
Law, G. L., Raney, A., Heusner, C. & Morris, D. R. Polyamine regulation of ribosome pausing at the upstream open reading frame of S-adenosylmethionine decarboxylase. J. Biol. Chem. 276, 38036–38043 (2001)
Michel, A. M. et al. GWIPS-viz: development of a ribo-seq genome browser. Nucleic Acids Res. 42, D859–D864 (2014)
Ji, Z., Song, R., Huang, H., Regev, A. & Struhl, K. Transcriptome-scale RNase-footprinting of RNA-protein complexes. Nat. Biotechnol. 34, 410–413 (2016)
Schueren, F. et al. Peroxisomal lactate dehydrogenase is generated by translational readthrough in mammals. eLife 3, e03640 (2014)
Loughran, G. et al. Evidence of efficient stop codon readthrough in four mammalian genes. Nucleic Acids Res. 42, 8928–8938 (2014)
Stiebler, A. C. et al. Ribosomal readthrough at a short UGA stop codon context triggers dual localization of metabolic enzymes in fungi and animals. PLoS Genet. 10, e1004685 (2014)
Rosenbloom, K. R. et al. The UCSC Genome Browser database: 2015 update. Nucleic Acids Res. 43, D670–D681 (2015)
Gao, X. et al. Quantitative profiling of initiating ribosomes in vivo. Nat. Methods 12, 147–153 (2015)
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011)
Yanagitani, K. et al. Cotranslational targeting of XBP1 protein to the membrane promotes cytoplasmic splicing of its own mRNA. Mol. Cell 34, 191–200 (2009)
Yanagitani, K., Kimata, Y., Kadokura, H. & Kohno, K. Translational pausing ensures membrane targeting and cytoplasmic splicing of XBP1u mRNA. Science 331, 586–589 (2011)
Namy, O., Duchateau-Nguyen, G. & Rousset, J. P. Translational readthrough of the PDE2 stop codon modulates cAMP levels in Saccharomyces cerevisiae. Mol. Microbiol. 43, 641–652 (2002)
Arribere, J. A. et al. Translation readthrough mitigation. Nature 534, 719–723 (2016)
Ryan, M. D. & Drew, J. Foot-and-mouth disease virus 2A oligopeptide mediated cleavage of an artificial polyprotein. EMBO J. 13, 928–933 (1994)
Doronina, V. A. et al. Site-specific release of nascent chains from ribosomes at a sense codon. Mol. Cell. Biol. 28, 4227–4239 (2008)
Miller-Fleming, L., Olin-Sandoval, V., Campbell, K. & Ralser, M. Remaining mysteries of molecular biology: the role of polyamines in the cell. J. Mol. Biol. 427, 3389–3406 (2015)
Kurian, L., Palanimurugan, R., Gödderz, D. & Dohmen, R. J. Polyamine sensing by nascent ornithine decarboxylase antizyme stimulates decoding of its mRNA. Nature 477, 490–494 (2011)
Fixsen, S. M. & Howard, M. T. Processive selenocysteine incorporation during synthesis of eukaryotic selenoproteins. J. Mol. Biol. 399, 385–396 (2010)
Grentzmann, G., Ingram, J. A., Kelly, P. J., Gesteland, R. F. & Atkins, J. F. A dual-luciferase reporter system for studying recoding signals. RNA 4, 479–486 (1998)
Loughran, G., Howard, M. T., Firth, A. E. & Atkins, J. F. Avoidance of reporter assay distortions from fused dual reporters. RNA 23, 1285–1289 (2017)
Dyer, B. W., Ferrer, F. A., Klinedinst, D. K. & Rodriguez, R. A noncommercial dual luciferase enzyme assay system for reporter gene analysis. Anal. Biochem. 282, 158–161 (2000)
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 ( 2001)
Terenin, I. M., Andreev, D. E., Dmitriev, S. E. & Shatsky, I. N. A novel mechanism of eukaryotic translation initiation that is neither m7G-cap-, nor IRES-dependent. Nucleic Acids Res. 41, 1807–1816 (2013)
Andreev, D. E. et al. Differential contribution of the m7G-cap to the 5′ end-dependent translation initiation of mammalian mRNAs. Nucleic Acids Res. 37, 6135–6147 (2009)
Degnin, C. R., Schleiss, M. R., Cao, J. & Geballe, A. P. Translational inhibition mediated by a short upstream open reading frame in the human cytomegalovirus gpUL4 (gp48) transcript. J. Virol. 67, 5514–5521 (1993)
Bhushan, S. et al. Structural basis for translational stalling by human cytomegalovirus and fungal arginine attenuator peptide. Mol. Cell 40, 138–146 (2010)
Mariotti, M. & Guigó, R. Selenoprofiles: profile-based scanning of eukaryotic genome sequences for selenoprotein genes. Bioinformatics 26, 2656–2663 (2010)
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 37, D5–D15 (2009)
Huerta-Cepas, J., Serra, F. & Bork, P. ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol. Biol. Evol. 33, 1635–1638 (2016)
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007)
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012)
Acknowledgements
We are grateful to I. Jungreis and M. Kellis for allowing us to use CodAlignView and to P.R. Bhatt for technical advice on stable peptidyl–tRNA complex formation. We acknowledge financial support from Science Foundation Ireland ((12/IA/1335) to P.V.B., (13/IA/1853) to J.F.A. and (12/RC/2276) to D.B.P.); National Institute of Health (CA080946) and (GM065204) to V.N.G.; Health Research Board (PhD/2007/04) to I.T.; and Russian Science Foundation (16-14-10065) to D.E.A.
Author information
Authors and Affiliations
Contributions
P.B.F.O. made the initial observation of unusual ribosome footprint density. M.M. and P.V.B. performed phylogenetic analysis. M.M.Y., G.L., A.V.Z., I.T., P.S. and D.E.A. designed and performed biochemical experiments. S.J.K. and A.M.M. analysed publicly available ribosome profiling data. P.V.B. proposed the model. M.M.Y. and P.V.B. drafted the manuscript. V.N.G., D.B.P. and J.F.A. contributed to interpretation of the experimental data and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.M.M., G.L. and P.V.B. are founders and shareholders of RiboMaps Ltd, a company providing ribosome profiling as a service. As the finding reported here stemmed from the analysis of ribosome profiling data, its publication may increase the attractiveness of this technology and indirectly benefit the company.
Additional information
Reviewer Information Nature thanks A. Geballe, P. Van Damme and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Cross-species examination of AMD1 tail using publicly available ribosome profiling data in the GWIPS-viz browser.
Available ribosome footprints aligned to the genomes of (a) mouse, (b) rat, (c) frog and (d) zebrafish are shown along with gene annotation tracks and ORF plots in which ATG codons are shown in green and stop codons are shown in red. Note that, even under very low coverage, peaks of density are consistently present at the stop codon of AMD1 tail. The low number of footprints in mouse is due to ambiguous mapping caused by the presence of a retrotransposed single-exon AMD1 copy (AMD2). Only uniquely aligned footprints are currently displayed in GWIPS-viz.
Extended Data Figure 2 Human ribosome profiling data obtained with approaches that enrich ribosomes at translation initiation sites.
a, AMD1 locus; b, XBP1 locus. Three tracks are shown as indicated in the figure. Ribosome footprint density corresponds to aggregated data obtained with drug treatments that preferentially arrest initiating ribosomes. Under these treatments, actively elongating ribosomes run off. However, stalled ribosomes remain bound to mRNA and produce footprints along with initiating ribosomes blocked by these inhibitors. The peaks corresponding to ribosome stalling at the end of the AMD1 tail and at the end of the XBP1 coding region (in unprocessed mRNA) are indicated with arrows.
Extended Data Figure 3 Assessment of dual luciferase mRNA stability.
a, Scheme of constructs. b, RT–qPCR analysis with primers targeting R Luc sequence; n = 9. c, Absolute values of R Luc and F Luc; n = 12. Biological replicates that belong to the same independent experiment are indicated with the same symbols in b and c.
Extended Data Figure 4 Expression of GFP constructs.
Western blotting analysis of GFP fusions with a fragment of the actin 3′ trailer of the same length as full-length AMD1 tail; n = 2 (a) and truncated from the 5′ end tail; n = 2 (b), separated by either stop or sense codons as indicated.
Extended Data Figure 5 In vitro translation of AMD1 tail fusion reporters.
Western blotting analysis of luciferase-expressing mRNAs in the HEK293T cell-free translation system; n = 1.
Extended Data Figure 6 Potential trans effect of AMD1 tail translation.
HEK293T cells were transfected in triplicate wells of half-area 96-well plates with the indicated expression constructs (left) for 24 h. Cells were lysed in 15 μl PLB and incubated with shaking for 15 min at room temperature. Five microlitres of each were removed for immunoblotting with both anti-GFP and anti-β-actin (upper right; n = 3); the remaining lysate was assayed for both R Luc and F Luc activities (lower right), n = 3.
Extended Data Figure 7 Expression of StopGo constructs.
a, Scheme of constructs. b, Western blots with antibodies against Renilla and firefly; n = 1. c, RT–qPCR analysis; n = 4. Line represents mean.
Extended Data Figure 8 EEF1A2 readthrough extension.
GWIPS-viz screenshot of ribosome footprint density at the last 3′ exon of EEF1A2 (hg38).
Supplementary information
Supplementary Information
This file contains the sequences of DNA primers used in this study, the Python code for identifying peaks of ribosome density in extended ORFs and a guide to the supplementary files.
Supplementary Figure 1
Gel Source Data. Original source Images of the gels that have been used for making figures with weight markers. Cropped parts are indicated.
Supplementary Data 1
Genomic alignment of tetrapods from UCSC Genome browser 100 species alignment. Codon alignment obtained with CodAlignView, positions of AMD1 stop and AMD1 tail stop are annotated (second row).
Supplementary Data 2
Alignment of AMD1 coding region and surrounding areas from 146 vertebrate species. Synonymous and nonsynonymous substitutions are indicated by blue and red colours, respectively, and gaps are in grey. Ka/Ks ratio and sequence identity (see Methods) are shown at the bottom.
Supplementary Data 3
Human transcripts with ribosome density profiles similar to AMD1. List of GENCODE transcripts containing peaks of ribosome density downstream and in-frame of protein coding regions. For each transcript information on the chromosome, coordinates, locus, GENCODE ID and the number of footprints are provided in comma delimited format.
Supplementary Data 4
Vectors and plasmids. Sequences of vectors and plasmids used in this study in fasta format.
Supplementary Data 5
Genomic sequences of AMD1 coding regions. Genomic sequence of AMD1 coding regions for 146 vertebrate species used in this study in fasta format. Genbank IDs for the source sequences are provided in the comment line for each sequence.
Supplementary Data 6
Ribosome profiling datasets used for GWIPS-viz global aggregate tracks. Datasets are listed on separate sheets for each genome, first column indicates the publication in which the datasets are described (first author name followed by the year, full reference can be found in GWIPS-viz), second column provides GEO or SRA IDs for each individual dataset from the corresponding study.
Rights and permissions
About this article
Cite this article
Yordanova, M., Loughran, G., Zhdanov, A. et al. AMD1 mRNA employs ribosome stalling as a mechanism for molecular memory formation. Nature 553, 356–360 (2018). https://doi.org/10.1038/nature25174
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25174
This article is cited by
-
Genome-wide DNA methylation profiling of CD4+ T lymphocytes identifies differentially methylated loci associated with adult primary refractory immune thrombocytopenia
BMC Medical Genomics (2023)
-
Noncoding translation mitigation
Nature (2023)
-
Comparative ribosome profiling reveals distinct translational landscapes of salt-sensitive and -tolerant rice
BMC Genomics (2021)
-
Humans and other commonly used model organisms are resistant to cycloheximide-mediated biases in ribosome profiling experiments
Nature Communications (2021)
-
Selection Shapes Synonymous Stop Codon Use in Mammals
Journal of Molecular Evolution (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.