Abstract
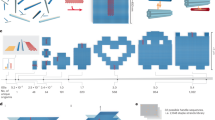
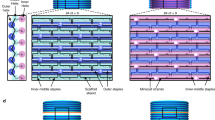
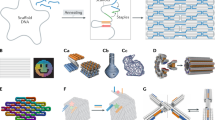
Self-assembled DNA nanostructures1 enable nanometre-precise patterning that can be used to create programmable molecular machines2,3,4,5,6 and arrays of functional materials7,8,9. DNA origami10 is particularly versatile in this context because each DNA strand in the origami nanostructure occupies a unique position and can serve as a uniquely addressable pixel. However, the scale of such structures11,12,13,14 has been limited to about 0.05 square micrometres, hindering applications that demand a larger layout15 and integration with more conventional patterning methods. Hierarchical multistage assembly of simple sets of tiles16,17 can in principle overcome this limitation, but so far has not been sufficiently robust to enable successful implementation of larger structures using DNA origami tiles. Here we show that by using simple local assembly rules18 that are modified and applied recursively throughout a hierarchical, multistage assembly process, a small and constant set of unique DNA strands can be used to create DNA origami arrays of increasing size and with arbitrary patterns. We illustrate this method, which we term ‘fractal assembly’, by producing DNA origami arrays with sizes of up to 0.5 square micrometres and with up to 8,704 pixels, allowing us to render images such as the Mona Lisa and a rooster. We find that self-assembly of the tiles into arrays is unaffected by changes in surface patterns on the tiles, and that the yield of the fractal assembly process corresponds to about 0.95m − 1 for arrays containing m tiles. When used in conjunction with a software tool that we developed that converts an arbitrary pattern into DNA sequences and experimental protocols, our assembly method is readily accessible and will facilitate the construction of sophisticated materials and devices with sizes similar to that of a bacterium using DNA nanostructures.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Seeman, N. C. Nucleic acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982)
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2010)
Gu, H., Chao, J., Xiao, S.-J. & Seeman, N. C. A proximity-based programmable DNA nanoscale assembly line. Nature 465, 202–205 (2010)
Wickham, S. F. et al. A DNA-based molecular motor that can navigate a network of tracks. Nat. Nanotechnol. 7, 169–173 (2012)
Thubagere, A. J. et al. A cargo-sorting DNA robot. Science 357, eaan6558 (2017)
Chatterjee, G., Dalchau, N., Muscat, R. A., Phillips, A. & Seelig, G. A spatially localized architecture for fast and modular DNA computing. Nat. Nanotechnol. 12, 920–927 (2017)
Maune, H. T. et al. Self-assembly of carbon nanotubes into two-dimensional geometries using DNA origami templates. Nat. Nanotechnol. 5, 61–66 (2010)
Knudsen, J. B. et al. Routing of individual polymers in designed patterns. Nat. Nanotechnol. 10, 892–898 (2015)
Gopinath, A., Miyazono, E., Faraon, A. & Rothemund, P. W. Engineering and mapping nanocavity emission via precision placement of DNA origami. Nature 535, 401–405 (2016)
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006)
Woo, S. & Rothemund, P. W. Programmable molecular recognition based on the geometry of DNA nanostructures. Nat. Chem. 3, 620–627 (2011)
Rajendran, A., Endo, M., Katsuda, Y., Hidaka, K. & Sugiyama, H. Programmed two-dimensional self-assembly of multiple DNA origami jigsaw pieces. ACS Nano 5, 665–671 (2011)
Zhao, Z., Liu, Y. & Yan, H. Organizing DNA origami tiles into larger structures using preformed scaffold frames. Nano Lett. 11, 2997–3002 (2011)
Marchi, A. N., Saaem, I., Vogen, B. N., Brown, S. & LaBean, T. H. Toward larger DNA origami. Nano Lett. 14, 5740–5747 (2014)
Pinheiro, A. V., Han, D., Shih, W. M. & Yan, H. Challenges and opportunities for structural DNA nanotechnology. Nat. Nanotechnol. 6, 763–772 (2011)
Doty, D. Theory of algorithmic self-assembly. Commun. ACM 55, 78–88 (2012)
Park, S. H. et al. Finite-size, fully addressable DNA tile lattices formed by hierarchical assembly procedures. Angew. Chem. 118, 749–753 (2006)
Tikhomirov, G., Petersen, P. & Qian, L. Programmable disorder in random DNA tilings. Nat. Nanotechnol. 12, 251–259 (2017)
Mandelbrot, B. B. The Fractal Geometry of Nature (W. H. Freeman and Co., 1982)
Nangreave, J., Yan, H. & Liu, Y. Studies of thermal stability of multivalent DNA hybridization in a nanostructured system. Biophys. J. 97, 563–571 (2009)
Rinker, S., Ke, Y., Liu, Y., Chhabra, R. & Yan, H. Self-assembled DNA nanostructures for distance-dependent multivalent ligand–protein binding. Nat. Nanotechnol. 3, 418–422 (2008)
Schreiber, R. et al. Hierarchical assembly of metal nanoparticles, quantum dots and organic dyes using DNA origami scaffolds. Nat. Nanotechnol. 9, 74–78 (2014)
Castro, C. E. et al. A primer to scaffolded DNA origami. Nat. Methods 8, 221–229 (2011)
Chemical synthesis and purification of oligonucleotideshttps://www.idtdna.com/pages/docs/technical-reports/chemical-synthesis-of-oligonucleotides.pdf (Integrated DNA Technologies, 2005)
Petersen, P. FracTile Compilerhttp://qianlab.caltech.edu/FracTileCompiler (2017)
Zenk, J., Tuntivate, C. & Schulman, R. Kinetics and thermodynamics of Watson–Crick base pairing driven DNA origami dimerization. J. Am. Chem. Soc. 138, 3346–3354 (2016)
Lin, C., Perrault, S. D., Kwak, M., Graf, F. & Shih, W. M. Purification of DNA-origami nanostructures by rate-zonal centrifugation. Nucleic Acids Res. 41, e40 (2013)
Xia, Y. & Whitesides, G. M. Soft lithography. Annu. Rev. Mater. Sci. 28, 153–184 (1998)
Chandran, H., Gopalkrishnan, N., Phillips, A. & Reif, J. Localized hybridization circuits. Lect. Notes Comput. Sci. 6937, 64–83 (2011)
Qian, L. & Winfree, E. Parallel and scalable computation and spatial dynamics with DNA-based chemical reaction networks on a surface. Lect. Notes Comput. Sci. 8727, 114–131 (2014)
Delbrück, T. & Mead, C. in Vision Chips: Implementing Vision Algorithms with Analog VLSI Circuits (eds Koch, C. & Li, H. ) 139–161 (IEEE, 1995)
Acknowledgements
We thank R. M. Murray for sharing an acoustic liquid-handling robot and P. W. K. Rothemund for sharing a qPCR machine. We thank E. Winfree and P. W. K. Rothemund for critiques on the manuscript. G.T. was supported by a BWF grant (1010684). P.P. was supported by a NIH/NRSA training grant (5 T32 GM07616). L.Q. was supported by a Career Award at the Scientific Interface from the Burroughs Wellcome Fund (1010684) and a Faculty Early Career Development Award from NSF (1351081).
Author information
Authors and Affiliations
Contributions
G.T. and P.P. initiated the project, designed and performed the experiments, and analysed the data; P.P. developed the software tool; G.T., P.P. and L.Q. wrote the manuscript; and L.Q. guided the project.
Corresponding author
Ethics declarations
Competing interests
A provisional patent application has been filed for this work, submitted to the Office of Technology Transfer at the California Institute of Technology.
Additional information
Reviewer Information Nature thanks T. LaBean, H. Yan and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
This file contains Supplementary Materials and Methods, Supplementary Notes, Supplementary Data and Analysis (Supplementary Figures 1-23), DNA sequences (Supplementary Tables S1-S6), and additional references – see contents page for details. (PDF 17127 kb)
Rights and permissions
About this article
Cite this article
Tikhomirov, G., Petersen, P. & Qian, L. Fractal assembly of micrometre-scale DNA origami arrays with arbitrary patterns. Nature 552, 67–71 (2017). https://doi.org/10.1038/nature24655
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24655
This article is cited by
-
DNA as a universal chemical substrate for computing and data storage
Nature Reviews Chemistry (2024)
-
Self-similar chiral organic molecular cages
Nature Communications (2024)
-
The Need for Seed (in the Abstract Tile Assembly Model)
Algorithmica (2024)
-
A temporally resolved DNA framework state machine in living cells
Nature Machine Intelligence (2023)
-
Biotemplated precise assembly approach toward ultra-scaled high-performance electronics
Nature Protocols (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.