Abstract
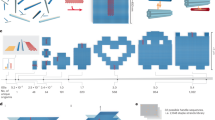
Nucleic acids (DNA and RNA) are widely used to construct nanometre-scale structures with ever increasing complexity1,2,3,4,5,6,7,8,9,10,11,12,13,14, with possible application in fields such as structural biology, biophysics, synthetic biology and photonics. The nanostructures are formed through one-pot self-assembly, with early kilodalton-scale examples containing typically tens of unique DNA strands. The introduction of DNA origami4, which uses many staple strands to fold one long scaffold strand into a desired structure, has provided access to megadalton-scale nanostructures that contain hundreds of unique DNA strands6,7,10,11,12,13,14. Even larger DNA origami structures are possible15,16, but manufacturing and manipulating an increasingly long scaffold strand remains a challenge. An alternative and more readily scalable approach involves the assembly of DNA bricks, which each consist of four short binding domains arranged so that the bricks can interlock8,9. This approach does not require a scaffold; instead, the short DNA brick strands self-assemble according to specific inter-brick interactions. First-generation bricks used to create three-dimensional structures are 32 nucleotides long, consisting of four eight-nucleotide binding domains. Protocols have been designed to direct the assembly of hundreds of distinct bricks into well formed structures, but attempts to create larger structures have encountered practical challenges and had limited success9. Here we show that DNA bricks with longer, 13-nucleotide binding domains make it possible to self-assemble 0.1–1-gigadalton, three-dimensional nanostructures from tens of thousands of unique components, including a 0.5-gigadalton cuboid containing about 30,000 unique bricks and a 1-gigadalton rotationally symmetric tetramer. We also assembled a cuboid that contains around 10,000 bricks and about 20,000 uniquely addressable, 13-base-pair ‘voxels’ that serves as a molecular canvas for three-dimensional sculpting. Complex, user-prescribed, three-dimensional cavities can be produced within this molecular canvas, enabling the creation of shapes such as letters, a helicoid and a teddy bear. We anticipate that with further optimization of structure design, strand synthesis and assembly procedure even larger structures could be accessible, which could be useful for applications such as positioning functional components.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chen, J. & Seeman, N. C. The synthesis from DNA of a molecule with the connectivity of a cube. Nature 350, 631–633 (1991)
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998)
Shih, W. M., Quispe, J. D. & Joyce, G. F. A. 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron. Nature 427, 618–621 (2004)
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006)
Zheng, J. P. et al. From molecular to macroscopic via the rational design of a self-assembled 3D DNA crystal. Nature 461, 74–77 (2009)
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009)
Han, D. et al. DNA origami with complex curvatures in three-dimensional space. Science 332, 342–346 (2011)
Wei, B., Dai, M. & Yin, P. Complex shapes self-assembled from single-stranded DNA tiles. Nature 485, 623–626 (2012)
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012)
Han, D. et al. DNA gridiron nanostructures based on four-arm junctions. Science 339, 1412–1415 (2013)
Iinuma, R. et al. Polyhedra self-assembled from DNA tripods and characterized with 3D DNA-PAINT. Science 344, 65–69 (2014)
Gerling, T., Wagenbauer, K. F., Neuner, A. M. & Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-base pairing 3D components. Science 347, 1446–1452 (2015)
Benson, E. et al. DNA rendering of polyhedral meshes at the nanoscale. Nature 523, 441–444 (2015)
Veneziano, R. et al. Designer nanoscale DNA assemblies programmed from the top down. Science 352, 1534 (2016)
Marchi, A. N., Saaem, I., Vogen, B. N., Brown, S. & Labean, T. H. Towards larger DNA origami. Nano Lett. 14, 5740–5747 (2014)
Nickels, P. C. et al. DNA origami structures directly assembled from intact bacteriophages. Small 10, 1765–1769 (2014)
Liu, Y., Ke, Y. & Yan, H. Self-assembly of symmetric finite-size DNA nanoarrays. J. Am. Chem. Soc. 127, 17140–17141 (2005)
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014)
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009)
Midgley, P. A. & Weyland, M. 3D electron microscopy in the physical sciences: the development of Z-contrast and EFTEM tomography. Ultramicroscopy 96, 413–431 (2003)
Myhrvold, C. et al. Barcode extension for analysis and reconstruction of structures (BEARS). Nat. Commun. 8, 14698 (2017)
Jacobs, W. M., Reinhardt, A. & Frenkel, D. Rational design of self-assembled pathways for complex multicomponent structures. Proc. Natl Acad. Sci. USA 112, 6313–6318 (2015)
Nickels, P. C. et al. Molecular force spectroscopy with a DNA origami-based nanoscopic force clamp. Science 354, 305–307 (2016)
Douglas, S. M., Chou, J. J. & Shih, W. M. DNA-nanotube-induced alignment of membrane proteins for NMR structure determination. Proc. Natl Acad. Sci. USA 104, 6644–6648 (2007)
Fu, J. et al. Multi-enzyme complexes on DNA scaffolds capable of substrate channeling with an artificial swinging arm. Nat. Nanotechnol. 9, 531–536 (2014)
Sun, W. et al. Casting inorganic structures with DNA molds. Science 346, 1258361 (2014)
Knudsen, J. B. et al. Routing of individual polymers in designed patterns. Nat. Nanotechnol. 10, 892–898 (2015)
Acuna, G. P. et al. Fluorescence enhancement at docking sites of DNA-directed self-assembled nanoantennas. Science 338, 506–510 (2012)
Schmidt, T. L. et al. Scalable amplification of strand subsets from chip-synthesized oligonucleotide libraries. Nat. Commun. 6, 8634 (2015)
Rajendran, A., Endo, M., Katsuda, Y., Hidaka, K. & Sugiyama, H. Programmed two-dimensional self-assembly of multiple DNA origami jigsaw pieces. ACS Nano 5, 665–671 (2011)
Sobczak, J.-P. J., Martin, T. G., Gerling, T. & Dietz, H. Rapid folding of DNA into nanoscale shapes at constant temperature. Science 338, 1458–1461 (2012)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012)
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008)
Lin, C. et al. Sub-micrometre geometrically encoded fluorescent barcodes self-assembled from DNA. Nat. Chem. 4, 832–839 (2012)
El Beheiry, M. & Dahan, M. ViSP: representing single-particle localizations in three dimensions. Nat. Methods 10, 689–690 (2013)
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012)
Quail, M. A. et al. A large genome centre’s improvements to the Illumina sequencing system. Nat. Methods 5, 1005–1010 (2008)
Acknowledgements
We thank N. Ponnuswamy, R. Sørensen, J. Hahn, J. Lara, L. Chou, N. Garreau, S. Saka, H. Sasaki, J. B. Woehrstein and C. B. Marks for experimental help. We also thank B. Wei, W. Sun and W.M. Shih for discussions, M. Beatty and J. Cheng for help in developing the Nanobricks platform, and C. Chen for assistance with draft preparation. The work was funded by Office of Naval Research grants N000141010827, N000141310593, N000141410610, N000141612182 and N000141612410, an Army Research Office grant W911NF1210238, National Science Foundation grants CCF-1054898, CCF-1162459, CCF-1317291, CMMI-1333215, CMMI-1334109 and CMMI-1344915, an Air Force Office of Scientific Research grant AFA9550-15-1-0514, and National Institute of Health grants 1DP2OD007292 and 1R01EB018659, 167814 (P.Y.); an Emory Biomedical Engineering Department Startup Fund, an Emory Winship Cancer Institute Billi and Bernie Marcus Research Award, a Winship Cancer Institute grant number IRG-14-188-0 from the American Cancer Society, and a National Science Foundation CAREER Award DMR–1654485 (Y.K.); French National Research Agency grants ANR-16-CE09-0004-01 and ANR-15-CE09-0003-02 (G.B.); and a French National Research Agency grant ANR-10-INBS-05 (P.B.). L.L.O. was funded by an NSF graduate research fellowship. N.H. was funded by the German National Academic Foundation and German Academic Exchange Service. M.T.S. acknowledges support from the International Max Planck Research School for Molecular and Cellular Life Sciences (IMPRS-LS).
Author information
Authors and Affiliations
Contributions
L.L.O. conceived the project, designed and performed the experiments, analysed the data and wrote the paper. N.H. designed and performed the experiments, analysed the data and wrote the paper. O.K.Y., B.W. and P.W. performed the experiments and analysed the data. M.T.S. and F.S. performed the 3D DNA-PAINT experiments, analysed the data and wrote the paper. C.G. and J.Y.K. developed the Nanobricks software and wrote the paper. P.B. and J.L.-K.-H. performed the electron tomography experiments. C.M. designed and analysed the sequencing experiments and wrote the paper. A.Z. performed the experiments. R.J. supervised the DNA-PAINT experiments, interpreted data and wrote the paper. G.B. designed and supervised the electron tomography study, interpreted data and wrote the paper. Y.K. and P.Y. conceived, designed and supervised the study, interpreted the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
A patent has been filed based on this work. P.Y. is co-founder of Ultivue Inc. and NuProbe Global.
Additional information
Reviewer Information Nature thanks C. Lin and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Gel electrophoresis analysis of DNA brick cuboids.
a–h, Structures of varying size (see schematics on the left) were assembled isothermally for 5–7 days at the temperatures indicated above each gel lane, with strand concentrations of 30 nM (a–d), 5 nM (e, g), 3 nM (f) and 20 nM (h). The number below each lane indicates the formation yield of the target structure. Lane ‘M’ contains a 1-kilobase ladder.
Extended Data Figure 2 Characterization of 30H × 30H × 260B cavity shapes.
a, Schematic of the 30H × 30H × 260B molecular canvas (grey) compared with a DNA-origami-sized structure (blue). b, For each structure (numbered 1–7), the top panels show 3D models of the designed structure, the bottom left panels show expected TEM projections and the bottom right panels show the TEM averages from at least six particles. c, The structures were folded with 5 nM per strand by isothermal annealing or by using a narrow ramp from 52.5 °C to 51 °C. Products were analysed on a 0.5% agarose gel in the presence of 10 mM MgCl2. The percentage listed below a target band indicates the gel yield; labels correspond to those in b or in Fig. 3.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Data, Supplementary Tables 1-3, Supplementary Figures 1-113 and Supplementary References – see contents pages for details. (PDF 30485 kb)
Supplementary Data 1
This file contains the sequences used for each structure on separate tabs. (XLSX 4706 kb)
Bear structure
Tomography video of the bear structure (AVI 11680 kb)
GEB structure
Tomography video of the GEB structure (AVI 12801 kb)
Helix structure
Tomography video of the helix structure (AVI 7636 kb)
Rights and permissions
About this article
Cite this article
Ong, L., Hanikel, N., Yaghi, O. et al. Programmable self-assembly of three-dimensional nanostructures from 10,000 unique components. Nature 552, 72–77 (2017). https://doi.org/10.1038/nature24648
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24648
This article is cited by
-
Pattern recognition in the nucleation kinetics of non-equilibrium self-assembly
Nature (2024)
-
The harmony of form and function in DNA nanotechnology
Nature Nanotechnology (2023)
-
Isothermal self-assembly of multicomponent and evolutive DNA nanostructures
Nature Nanotechnology (2023)
-
DNA T-shaped crossover tiles for 2D tessellation and nanoring reconfiguration
Nature Communications (2023)
-
Biotemplated precise assembly approach toward ultra-scaled high-performance electronics
Nature Protocols (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.