Abstract
The peptide-loading complex (PLC) is a transient, multisubunit membrane complex in the endoplasmic reticulum that is essential for establishing a hierarchical immune response. The PLC coordinates peptide translocation into the endoplasmic reticulum with loading and editing of major histocompatibility complex class I (MHC-I) molecules. After final proofreading in the PLC, stable peptide–MHC-I complexes are released to the cell surface to evoke a T-cell response against infected or malignant cells1,2. Sampling of different MHC-I allomorphs requires the precise coordination of seven different subunits in a single macromolecular assembly, including the transporter associated with antigen processing (TAP1 and TAP2, jointly referred to as TAP), the oxidoreductase ERp57, the MHC-I heterodimer, and the chaperones tapasin and calreticulin3,4. The molecular organization of and mechanistic events that take place in the PLC are unknown owing to the heterogeneous composition and intrinsically dynamic nature of the complex. Here, we isolate human PLC from Burkitt’s lymphoma cells using an engineered viral inhibitor as bait and determine the structure of native PLC by electron cryo-microscopy. Two endoplasmic reticulum-resident editing modules composed of tapasin, calreticulin, ERp57, and MHC-I are centred around TAP in a pseudo-symmetric orientation. A multivalent chaperone network within and across the editing modules establishes the proofreading function at two lateral binding platforms for MHC-I molecules. The lectin-like domain of calreticulin senses the MHC-I glycan, whereas the P domain reaches over the MHC-I peptide-binding pocket towards ERp57. This arrangement allows tapasin to facilitate peptide editing by clamping MHC-I. The translocation pathway of TAP opens out into a large endoplasmic reticulum lumenal cavity, confined by the membrane entry points of tapasin and MHC-I. Two lateral windows channel the antigenic peptides to MHC-I. Structures of PLC captured at distinct assembly states provide mechanistic insight into the recruitment and release of MHC-I. Our work defines the molecular symbiosis of an ABC transporter and an endoplasmic reticulum chaperone network in MHC-I assembly and provides insight into the onset of the adaptive immune response.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Neefjes, J., Jongsma, M. L., Paul, P. & Bakke, O. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat. Rev. Immunol. 11, 823–836 (2011)
Blum, J. S., Wearsch, P. A. & Cresswell, P. Pathways of antigen processing. Annu. Rev. Immunol. 31, 443–473 (2013)
Hulpke, S. & Tampé, R. The MHC I loading complex: a multitasking machinery in adaptive immunity. Trends Biochem. Sci. 38, 412–420 (2013)
Ortmann, B. et al. A critical role for tapasin in the assembly and function of multimeric MHC class I-TAP complexes. Science 277, 1306–1309 (1997)
Ortmann, B., Androlewicz, M. J. & Cresswell, P. MHC I/β2-microglobulin complexes associate with TAP transporters before peptide binding. Nature 368, 864–867 (1994)
Sadasivan, B., Lehner, P. J., Ortmann, B., Spies, T. & Cresswell, P. Roles for calreticulin and a novel glycoprotein, tapasin, in the interaction of MHC class I molecules with TAP. Immunity 5, 103–114 (1996)
Herbring, V., Bäucker, A., Trowitzsch, S. & Tampé, R. A dual inhibition mechanism of herpesviral ICP47 arresting a conformationally thermostable TAP complex. Sci. Rep. 6, 36907 (2016)
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008)
Oldham, M. L. et al. A mechanism of viral immune evasion revealed by cryo-EM analysis of the TAP transporter. Nature 529, 537–540 (2016)
Zhang, S. et al. Structural basis of cross-allele presentation by HLA-A*0301 and HLA-A*1101 revealed by two HIV-derived peptide complexes. Mol. Immunol. 49, 395–401 (2011)
Kozlov, G. et al. Structural basis of cyclophilin B binding by the calnexin/calreticulin P-domain. J. Biol. Chem. 285, 35551–35557 (2010)
Dong, G., Wearsch, P. A., Peaper, D. R., Cresswell, P. & Reinisch, K. M. Insights into MHC class I peptide loading from the structure of the tapasin–ERp57 thiol oxidoreductase heterodimer. Immunity 30, 21–32 (2009)
Wearsch, P. A. & Cresswell, P. Selective loading of high-affinity peptides onto major histocompatibility complex class I molecules by the tapasin–ERp57 heterodimer. Nat. Immunol. 8, 873–881 (2007)
Wearsch, P. A., Peaper, D. R. & Cresswell, P. Essential glycan-dependent interactions optimize MHC class I peptide loading. Proc. Natl Acad. Sci. USA 108, 4950–4955 (2011)
Dick, T. P., Bangia, N., Peaper, D. R. & Cresswell, P. Disulfide bond isomerization and the assembly of MHC class I-peptide complexes. Immunity 16, 87–98 (2002)
Peaper, D. R., Wearsch, P. A. & Cresswell, P. Tapasin and ERp57 form a stable disulfide-linked dimer within the MHC class I peptide-loading complex. EMBO J. 24, 3613–3623 (2005)
Wang, R., Natarajan, K. & Margulies, D. H. Structural basis of the CD8αβ/MHC class I interaction: focused recognition orients CD8β to a T cell proximal position. J. Immunol. 183, 2554–2564 (2009)
Simone, L. C., Georgesen, C. J., Simone, P. D., Wang, X. & Solheim, J. C. Productive association between MHC class I and tapasin requires the tapasin transmembrane/cytosolic region and the tapasin C-terminal Ig-like domain. Mol. Immunol. 49, 628–639 (2012)
Fisette, O., Wingbermühle, S., Tampé, R. & Schäfer, L. V. Molecular mechanism of peptide editing in the tapasin-MHC I complex. Sci. Rep. 6, 19085 (2016)
Lewis, J. W., Neisig, A., Neefjes, J. & Elliott, T. Point mutations in the α2 domain of HLA-A2.1 define a functionally relevant interaction with TAP. Curr. Biol. 6, 873–883 (1996)
Thomas, C. & Tampé, R. Proofreading of peptide–MHC complexes through dynamic multivalent interactions. Front. Immunol. 8, 65 (2017)
Gao, B. et al. Assembly and antigen-presenting function of MHC class I molecules in cells lacking the ER chaperone calreticulin. Immunity 16, 99–109 (2002)
Wijeyesakere, S. J., Bedi, S. K., Huynh, D. & Raghavan, M. The C-terminal acidic region of calreticulin mediates phosphatidylserine binding and apoptotic cell phagocytosis. J. Immunol. 196, 3896–3909 (2016)
Zhang, Y. et al. ERp57 does not require interactions with calnexin and calreticulin to promote assembly of class I histocompatibility molecules, and it enhances peptide loading independently of its redox activity. J. Biol. Chem. 284, 10160–10173 (2009)
Frickel, E. M. et al. TROSY-NMR reveals interaction between ERp57 and the tip of the calreticulin P-domain. Proc. Natl Acad. Sci. USA 99, 1954–1959 (2002)
Garbi, N. et al. Impaired immune responses and altered peptide repertoire in tapasin-deficient mice. Nat. Immunol. 1, 234–238 (2000)
Garbi, N., Tanaka, S., Momburg, F. & Hämmerling, G. J. Impaired assembly of the major histocompatibility complex class I peptide-loading complex in mice deficient in the oxidoreductase ERp57. Nat. Immunol. 7, 93–102 (2006)
Kyritsis, C. et al. Molecular mechanism and structural aspects of transporter associated with antigen processing inhibition by the cytomegalovirus protein US6. J. Biol. Chem. 276, 48031–48039 (2001)
Panter, M. S., Jain, A., Leonhardt, R. M., Ha, T. & Cresswell, P. Dynamics of major histocompatibility complex class I association with the human peptide-loading complex. J. Biol. Chem. 287, 31172–31184 (2012)
Reits, E. A., Vos, J. C., Grommé, M. & Neefjes, J. The major substrates for TAP in vivo are derived from newly synthesized proteins. Nature 404, 774–778 (2000)
Fischbach, H. et al. Ultrasensitive quantification of TAP-dependent antigen compartmentalization in scarce primary immune cell subsets. Nat. Commun. 6, 6199 (2015)
Meyer, T. H., van Endert, P. M., Uebel, S., Ehring, B. & Tampé, R. Functional expression and purification of the ABC transporter complex associated with antigen processing (TAP) in insect cells. FEBS Lett. 351, 443–447 (1994)
van Endert, P. M. et al. A sequential model for peptide binding and transport by the transporters associated with antigen processing. Immunity 1, 491–500 (1994)
Hulpke, S., Baldauf, C. & Tampé, R. Molecular architecture of the MHC I peptide-loading complex: one tapasin molecule is essential and sufficient for antigen processing. FASEB J. 26, 5071–5080 (2012)
Rufer, E., Leonhardt, R. M. & Knittler, M. R. Molecular architecture of the TAP-associated MHC class I peptide-loading complex. J. Immunol. 179, 5717–5727 (2007)
Kim, J. Y., Kwak, P. B., Gebert, M., Duong, H. A. & Weitz, C. J. Purification and analysis of PERIOD protein complexes of the mammalian circadian clock. Methods Enzymol. 551, 197–210 (2015)
Lin, J. et al. A negative feedback modulator of antigen processing evolved from a frameshift in the cowpox virus genome. PLoS Pathog. 10, e1004554 (2014)
Shevchenko, A. et al. A strategy for identifying gel-separated proteins in sequence databases by MS alone. Biochem. Soc. Trans. 24, 893–896 (1996)
Olsen, J. V. et al. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics 4, 2010–2021 (2005)
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nat. Methods 9, 904–906 (2012)
Tao, H. et al. Engineered nanostructured β-sheet peptides protect membrane proteins. Nat. Methods 10, 759–761 (2013)
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005)
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017)
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016)
Voss, N. R., Yoshioka, C. K., Radermacher, M., Potter, C. S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009)
Lander, G. C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013)
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017)
Fernández, J. J., Luque, D., Castón, J. R. & Carrascosa, J. L. Sharpening high resolution information in single particle electron cryomicroscopy. J. Struct. Biol. 164, 170–175 (2008)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010)
Kozlov, G. et al. Structural basis of carbohydrate recognition by calreticulin. J. Biol. Chem. 285, 38612–38620 (2010)
Ellgaard, L. et al. NMR structure of the calreticulin P-domain. Proc. Natl Acad. Sci. USA 98, 3133–3138 (2001)
Schrag, J. D. et al. The structure of calnexin, an ER chaperone involved in quality control of protein folding. Mol. Cell 8, 633–644 (2001)
Topf, M. et al. Protein structure fitting and refinement guided by cryo-EM density. Structure 16, 295–307 (2008)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
DeLano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, 2002)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Grimm, M., Zimniak, T., Kahraman, A. & Herzog, F. xVis: a web server for the schematic visualization and interpretation of crosslink-derived spatial restraints. Nucleic Acids Res. 43, W362–W369 (2015)
Acknowledgements
This research was supported by the German Research Foundation (SFB 807 and GRK 1986 to R.T.). C.S. acknowledges funding from the Federal Ministry for Education and Research (BMBF, ZIK program, 03Z22HN22), the European Regional Development Funds (EFRE, ZS/2016/04/78115) and the MLU Halle-Wittenberg. We thank K. Zehl for technical support; all members of the Institute of Biochemistry (Goethe University Frankfurt) for comments on the manuscript; and especially W. Kühlbrandt, D. Mills, M. Wilkes, and the staff at the Department of Structural Biology (MPI of Biophysics, Frankfurt/Main) for discussions, cryo-EM infrastructure and support. E. D’Imprima and R. Sanchez provided the code for RecenterParticles.
Author information
Authors and Affiliations
Contributions
A.B. isolated the PLC and performed all biochemical experiments. D.J. and A.M. carried out all EM imaging and single-particle analyses. T.H. and C.S. performed the mass spectrometry experiments. N.K. implemented the single-cell-based transport analyses. S.T. and A.B. designed the purification strategy for the PLC. S.T. and D.J. built the PLC model. A.B., D.J., S.T., A.M., and R.T. interpreted the data and wrote the manuscript with contributions from all authors. A.M., S.T., and R.T. conceived the study, designed the research, and planned the experiments. R.T. initiated and planned the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks P. Cresswell, G. Skiniotis and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Functional arrest of endogenous PLC by ICP47–SBP and biochemical analyses of purified human PLC.
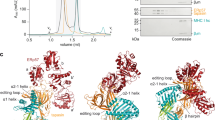
a, Design of ICP47–SBP. Full-length ICP47 was C-terminally fused to a glycine/serine (G/S) linker and a streptavidin-binding peptide (SBP). b, Single-cell-based peptide translocation assay. Burkitt’s lymphoma cells were semi-permeabilized with streptolysin O and incubated with fluorescent peptide RRYQNSTC(AF647)L (NST-AF647, 30 nM). Peptide translocation was carried out at 37 °C for 15 min in the presence of ATP or ADP (10 mM each) and with or without ICP47–SBP (5 μM). Peptide translocation was stopped by EDTA (20 mM) and analysed by flow cytometry. c, Histograms of the single-cell events (660-nm channel) demonstrate the inhibition of ATP-dependent translocation of antigenic peptides by ICP47–SBP (solid line, ATP; grey filled, ADP). Data are representative of two independent experiments. d, Summary of single-cell events indicated by mean fluorescence intensity (MFI), mean ± s.d. (n = 4). e, PLC subunits were analysed by SDS–PAGE and subsequent immunoblotting of the indicated proteins. f, PLC was treated by EndoH or PNGaseF and analysed by SDS–PAGE (Coomassie). Band shifts are indicated by short lines. Asterisk, nonspecific band. g, PLC was incubated with increasing amounts of the respective antibodies and antibody shifts were visualized by native PAGE and immunoblotting. The asterisk indicates formation of higher oligomeric states of the PLC observed only with the anti-tapasin antibody. Images shown are representative of five independent experiments.
Extended Data Figure 2 Direct comparison of negatively stained PLC particles.
The individual particles (expanded views of the representative micrographs) and 2D class averages of native PLC without any treatment (a) and prepared by GraFix (b) display the same architecture, confirming that the cross-linking procedure did not affect the organization of the whole particle. The scale bar is 100 nm in the micrograph and 25 nm in the 2D averages; inset is magnified 4×.
Extended Data Figure 3 Cryo-EM analysis of the human PLC.
a, Representative micrograph and 2D class averages. Scale bars, 50 nm in the micrograph and 10 nm in 2D class averages. Various views of the individual particles are readily discernable in the raw image. In the 2D class averages, note the clear densities for the ER-lumenal domain and the blurred signal for TAP. b, Angular assignment of the final dataset. The occurrence of multiple individual views, which are well spread over the entire sphere, displays an almost random orientation of the PLC in the sample. Individual cylinder bars are proportional in height to the number of particles in each view. The most frequent views are coloured red and the least common ones blue. c, Fourier shell correlation (FSC) curve of unfiltered reconstructions from two independently refined half datasets of the full PLC, generated by post-processing in Relion, displays 9.9 Å resolution as judged from the 0.143 threshold. d, FSC curves for the pseudo-C2 symmetric editing modules and the single module generated in Frealign show resolutions of 7.2 Å and 5.8 Å, respectively.
Extended Data Figure 4 Processing work flow.
The full dataset was directly submitted to multimodel classification in 3D. Particles from the two best classes were merged into a single stack and refined together as a consensus map. This dataset was further subjected to multimodel refinement (left branch), which led to the identification of different PLC assemblies. For high-resolution structure determination, a single editing module was extracted computationally from the consensus map and further refined individually (dotted lines indicate the use of the Frealign software package, whereas the solid lines represent processing in Relion).
Extended Data Figure 5 Individual subunits of the PLC editing module.
a–e, Individual segments of the single PLC editing module highlight the quality of the fit in the EM density. For each segment, the experimental map, the corresponding low-pass filtered version of the atomic model, the actual atomic model, and its fit into the experimental map are shown side by side in two different views to emphasize the consistency of the domain-based docking. f, The multi-model 3D classification, focused on a single editing module, emphasizes the flexibility of calreticulin (yellow). Whereas tapasin, ERp57, and the MHC-I heterodimer display the same relative position in all classes, calreticulin shows a substantial shift in its position, indicating high flexibility.
Extended Data Figure 6 Two opposing tapasin molecules shape the central scaffold.
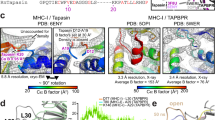
a, E225 of the N-terminal immunoglobulin-like domain of one molecule and R60 located in the short helical motif of the seven-stranded N-terminal β barrel of the second molecule are in salt-bridge distance. b, Multiple sequence alignment (MAFFT; https://mafft.cbrc.jp/alignment/software/) of tapasin orthologues from human, bovine, rat, mouse, fish, and chicken. Conserved Arg and Glu residues forming the salt bridges are indicated (asterisks). c, Multiple sequence alignment (MAFFT) of different human HLA-A/B/C allomorphs. Conserved Thr and Gln residues in the α3 domain are indicated (asterisks). Numbering is according to UniProt, including signal sequences.
Extended Data Figure 7 Cross-linking network.
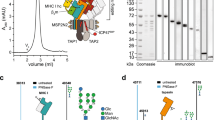
a, ICP47–SBP-purified PLC was cross-linked with BS3 and applied to in-gel or in-solution digestion before LC–MS/MS analysis (duplicates for in-gel digestion and single in-solution). Bands used for in-gel digestion are indicated. A representative gel is shown. b, Intra- and inter-cross link network (xVis60).
Supplementary information
Supplementary Information
This file contains the uncropped scans with size marker indications. (PDF 1329 kb)
Rights and permissions
About this article
Cite this article
Blees, A., Januliene, D., Hofmann, T. et al. Structure of the human MHC-I peptide-loading complex. Nature 551, 525–528 (2017). https://doi.org/10.1038/nature24627
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24627
This article is cited by
-
MR1 antigen presentation to MAIT cells and other MR1-restricted T cells
Nature Reviews Immunology (2024)
-
iRhom2 regulates ectodomain shedding and surface expression of the major histocompatibility complex (MHC) class I
Cellular and Molecular Life Sciences (2024)
-
Targeting MHC-I molecules for cancer: function, mechanism, and therapeutic prospects
Molecular Cancer (2023)
-
Emerging phagocytosis checkpoints in cancer immunotherapy
Signal Transduction and Targeted Therapy (2023)
-
Crystal structures of MHC class I complexes reveal the elusive intermediate conformations explored during peptide editing
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.