Abstract
Life on Earth depends on photosynthesis for its conversion of solar energy to chemical energy. Photosynthetic organisms have developed a variety of light-harvesting systems to capture sunlight. The largest light-harvesting complex is the phycobilisome (PBS), the main light-harvesting antenna in cyanobacteria and red algae. It is composed of phycobiliproteins and linker proteins but the assembly mechanisms and energy transfer pathways of the PBS are not well understood. Here we report the structure of a 16.8-megadalton PBS from a red alga at 3.5 Å resolution obtained by single-particle cryo-electron microscopy. We modelled 862 protein subunits, including 4 linkers in the core, 16 rod–core linkers and 52 rod linkers, and located a total of 2,048 chromophores. This structure reveals the mechanisms underlying specific interactions between linkers and phycobiliproteins, and the formation of linker skeletons. These results provide a firm structural basis for our understanding of complex assembly and the mechanisms of energy transfer within the PBS.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sidler, W. A. in The Molecular Biology of Cyanobacteria (ed. Bryant, D. A. ) 139–216 (Kluwer, 1994)
Adir, N. Elucidation of the molecular structures of components of the phycobilisome: reconstructing a giant. Photosynth. Res. 85, 15–32 (2005)
Watanabe, M. & Ikeuchi, M. Phycobilisome: architecture of a light-harvesting supercomplex. Photosynth. Res. 116, 265–276 (2013)
Croce, R. & van Amerongen, H. Natural strategies for photosynthetic light harvesting. Nat. Chem. Biol. 10, 492–501 (2014)
Schirmer, T. et al. Crystal structure analysis and refinement at 2.5 Å of hexameric C-phycocyanin from the cyanobacterium Agmenellum quadruplicatum. The molecular model and its implications for light-harvesting. J. Mol. Biol. 188, 651–676 (1986)
de Marsac, N. T. & Cohen-bazire, G. Molecular composition of cyanobacterial phycobilisomes. Proc. Natl Acad. Sci. USA 74, 1635–1639 (1977)
Arteni, A. A., Ajlani, G. & Boekema, E. J. Structural organisation of phycobilisomes from Synechocystis sp. strain PCC6803 and their interaction with the membrane. Biochim. Biophys. Acta 1787, 272–279 (2009)
Arteni, A. A. et al. Structure and organization of phycobilisomes on membranes of the red alga Porphyridium cruentum. Photosynth. Res. 95, 169–174 (2008)
Gantt, E. & Lipschultz, C. A. Structure and phycobiliprotein composition of phycobilisomes from Griffithsia pacifica (Rhodophyceae). J. Phycol. 16, 394–398 (1980)
Guglielmi, G., Cohen-Bazire, G. & Bryant, D. A. The structure of Gloeobacter violaceus and its phycobilisomes. Arch. Microbiol. 129, 181–189 (1981)
Williams, R. C., Gingrich, J. C. & Glazer, A. N. Cyanobacterial phycobilisomes. Particles from Synechocystis 6701 and two pigment mutants. J. Cell Biol. 85, 558–566 (1980)
Ducret, A., Sidler, W., Wehrli, E., Frank, G. & Zuber, H. Isolation, characterization and electron microscopy analysis of a hemidiscoidal phycobilisome type from the cyanobacterium Anabaena sp. PCC 7120. FEBS 236, 1010–1024 (1996)
MacColl, R. Allophycocyanin and energy transfer. Biochim. Biophys. Acta 1657, 73–81 (2004)
Gao, X. et al. Molecular insights into the terminal energy acceptor in cyanobacterial phycobilisome. Mol. Microbiol. 85, 907–915 (2012)
Lundell, D. J., Yamanaka, G. & Glazer, A. N. A terminal energy acceptor of the phycobilisome: the 75,000-dalton polypeptide of Synechococcus 6301 phycobilisomes—a new biliprotein. J. Cell Biol. 91, 315–319 (1981)
Tang, K. et al. The terminal phycobilisome emitter, LCM: A light-harvesting pigment with a phytochrome chromophore. Proc. Natl Acad. Sci. USA 112, 15880–15885 (2015)
Glazer, A. N. & Bryant, D. A. Allophycocyanin B (λmax 671, 618 nm): a new cyanobacterial phycobiliprotein. Arch. Microbiol. 104, 15–22 (1975)
Peng, P. P. et al. The structure of allophycocyanin B from Synechocystis PCC 6803 reveals the structural basis for the extreme redshift of the terminal emitter in phycobilisomes. Acta Crystallogr. D Biol. Crystallogr. 70, 2558–2569 (2014)
Gantt, E. Structure and function of phycobilisomes: light harvesting pigment complexes in red and blue-green algae. Int. Rev. Cytol. 66, 45–80 (1980)
Adir, N ., Dines, M ., Klartag, M ., McGregor, A. & Melamed-Frank, M. in Complex Intracellular Structures in Prokaryotes (ed. Schwender, J. ) 1–31 (Plant Metabolic Networks, 2009)
Dong, C. et al. ApcD is necessary for efficient energy transfer from phycobilisomes to photosystem I and helps to prevent photoinhibition in the cyanobacterium Synechococcus sp. PCC 7002. Biochim. Biophys. Acta 1787, 1122–1128 (2009)
Ashby, M. K. & Mullineaux, C. W. The role of ApcD and ApcF in energy transfer from phycobilisomes to PSI and PSII in a cyanobacterium. Photosynth. Res. 61, 169–179 (1999)
Chang, L. et al. Structural organization of an intact phycobilisome and its association with photosystem II. Cell Res. 25, 726–737 (2015)
Yamanaka, G., Lundell, D. J. & Glazer, A. N. Molecular architecture of a light-harvesting antenna. Isolation and characterization of phycobilisome subassembly particles. J. Biol. Chem. 257, 4077–4086 (1982)
Scholes, G. D., Mirkovic, T., Turner, D. B., Fassioli, F. & Buchleitner, A. Solar light harvesting by energy transfer: from ecology to coherence. Energy Environ. Sci. 5, 9374–9393 (2012)
Liu, L.-N., Chen, X.-L., Zhang, Y.-Z. & Zhou, B.-C. Characterization, structure and function of linker polypeptides in phycobilisomes of cyanobacteria and red algae: an overview. Biochim. Biophys. Acta 1708, 133–142 (2005)
Reuter, W., Wiegand, G., Huber, R. & Than, M. E. Structural analysis at 2.2 Å of orthorhombic crystals presents the asymmetry of the allophycocyanin-linker complex, AP.LC7.8, from phycobilisomes of Mastigocladus laminosus. Proc. Natl Acad. Sci. USA 96, 1363–1368 (1999)
Gao, X. et al. Crystal structure of the N-terminal domain of linker L(R) and the assembly of cyanobacterial phycobilisome rods. Mol. Microbiol. 82, 698–705 (2011)
Clout, N. J., Tisi, D. & Hohenester, E. Novel fold revealed by the structure of a FAS1 domain pair from the insect cell adhesion molecule fasciclin I. Structure 11, 197–203 (2003)
Klotz, A. V. & Glazer, A. N. Characterization of the bilin attachment sites in R-phycoerythrin. J. Biol. Chem. 260, 4856–4863 (1985)
Nagy, J. O., Bishop, J. E., Klotz, A. V., Glazer, A. N. & Rapoport, H. Bilin attachment sites in the alpha, beta, and gamma subunits of R-phycoerythrin. Structural studies on singly and doubly linked phycourobilins. J. Biol. Chem. 260, 4864–4868 (1985)
Blot, N. et al. Phycourobilin in trichromatic phycocyanin from oceanic cyanobacteria is formed post-translationally by a phycoerythrobilin lyase-isomerase. J. Biol. Chem. 284, 9290–9298 (2009)
Underhaug, J. et al. Mutation in transforming growth factor beta induced protein associated with granular corneal dystrophy type 1 reduces the proteolytic susceptibility through local structural stabilization. Biochim. Biophys. Acta 1834, 2812–2822 (2013)
Gindt, Y. M., Zhou, J., Bryant, D. A. & Sauer, K. Spectroscopic studies of phycobilisome subcore preparations lacking key core chromophores: assignment of excited state energies to the Lcm, beta 18 and alpha AP-B chromophores. Biochim. Biophys. Acta 1186, 153–162 (1994)
Bonaventura, C. & Myers, J. Fluorescence and oxygen evolution from Chlorella pyrenoidosa. Biochim. Biophys. Acta 189, 366–383 (1969)
Murata, N. Control of excitation transfer in photosynthesis. I. Light-induced change of chlorophyll a fluorescence in Porphyridium cruentum. Biochim. Biophys. Acta 172, 242–251 (1969)
Kanˇa, R. et al. Phycobilisome mobility and its role in the regulation of light harvesting in red algae. Plant Physiol. 165, 1618–1631 (2014)
Liu, H. et al. Phycobilisomes supply excitations to both photosystems in a megacomplex in cyanobacteria. Science 342, 1104–1107 (2013)
Ren, Y. et al. Clarifying and illustrating the electronic energy transfer pathways in trimeric and hexameric aggregation state of cyanobacteria allophycocyanin within the framework of Förster theory. J. Comput. Chem. 36, 137–145 (2015)
Contreras-Martel, C. et al. The structure at 2 Å resolution of Phycocyanin from Gracilaria chilensis and the energy transfer network in a PC–PC complex. Biophys. Chem. 125, 388–396 (2007)
Knox, R. S. Ultrashort processes and biology. J. Photochem. Photobiol. B 49, 81–88 (1999)
Glazer, A. N. Light guides. Directional energy transfer in a photosynthetic antenna. J. Biol. Chem. 264, 1–4 (1989)
Nganou, C., David, L., Adir, N. & Mkandawire, M. Linker proteins enable ultrafast excitation energy transfer in the phycobilisome antenna system of Thermosynechococcus vulcanus. Photochem. Photobiol. Sci. 15, 31–44 (2016)
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protocols 8, 1494–1512 (2013)
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005)
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res. 33, 6494–6506 (2005)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Scheres, S. H. W. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014)
Nield, J., Rizkallah, P. J., Barber, J. & Chayen, N. E. The 1.45 Å three-dimensional structure of C-phycocyanin from the thermophilic cyanobacterium Synechococcus elongatus. J. Struct. Biol. 141, 149–155 (2003)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997)
Brown, A. et al. Tools for macromolecular model building and refinement into electron cryo-microscopy reconstructions. Acta Crystallogr. D 71, 136–153 (2015)
Nicholls, R. A., Fischer, M., McNicholas, S. & Murshudov, G. N. Conformation-independent structural comparison of macromolecules with ProSMART. Acta Crystallogr. D 70, 2487–2499 (2014)
Acknowledgements
We thank J. Lei, L. Cheng and the staff at the Tsinghua University Branch of the National Center for Protein Sciences Beijing for providing facility support; J. Wang for model building, structure refinement and model validation; J. Dai for recommending RNA sequencing; L. Li and T. Zhu for transcriptome annotation; P. Meng and F. Chen for genomic DNA sequencing and data mining; V. Ragunath, O. Kovalevskiy and G. Murshudov (MRC Laboratory of Molecular Biology) for structure refinement; N. Gao for model building; H. Lin, L. Chang and Y. Liu for discussion; H.-W. Wang and J. Wu for commenting on the manuscript; and the members of the Tsinghua National Laboratory for Information Science and Technology for providing the “Explorer 100” cluster system as a computational resource. S.-F.S. and J.Zhao were supported by the National Basic Research Program (2016YFA0501101, 2017YFA0504600 and 2015CB150100). S.-F.S. and S.S. were supported by the National Natural Science Foundation of China (31230016, 31370717 and 31670745 and 31400632 and 31670746). J. Zhao also received support from the CAS fund (QYZDY-SSW-SMC004).
Author information
Authors and Affiliations
Contributions
S.-F.S. initiated and supervised the project; J.Zhang and J.M. prepared the samples; J.Zhang collected the EM data and performed the EM analysis and the model building; J.M. performed the biochemical and biophysical analyses; J.Zhang and D.L. refined the structure; S.S. and S.-F.S. analysed the structure; S.Q. contributed to the sample selection; J.Zhao participated in the analysis of biochemical data; and S.S., J.Zhao and S.-F.S. wrote the manuscript with help from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks H. van Amerongen, Z. Liu and C. Mullineaux for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Preparation and characterization of the PBS from Griffithsia pacifica and cryo-EM analysis using Tecnai Arctica microscope and FEI Falcon II direct electron detector.
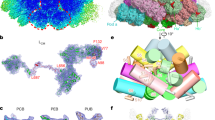
a, Isolation of PBSs using sucrose density gradient centrifugation. Band 1 is the sample of PBSs used for single particle analysis in this study. Band 2 is disassembled PBSs as evidenced by the fluorescence spectra shown in c. b, Absorption spectrum of band 1. The peaks at 498 nm, 620 nm and 650 nm are from phycourobilins of the LRγ proteins, phycocyanobilins of phycocyanins and phycocyanobilins of allophycocyanins, respectively. The peaks at 540 nm and 565 nm are from phycoerythrobilins of phycoerythrin. c, Fluorescence emission spectra of band 1 and band 2. Band 1 has an emission peak at 674 nm and band 2 has an emission peak at 580 nm, indicating that band 1 contains intact PBSs and band 2 contains disassembled PBSs. d, SDS–PAGE analysis of protein components in PBS from band 1. The gel was stained with Coomassie brilliant blue. The bands of LCM, LR and LRC proteins and PBPs identified by MS analysis are indicated. For gel source data, see Supplementary Fig. 1. e, Protein composition analysis of band 1 by SDS–PAGE stained with ZnSO4 to detect bilin-containing proteins with ultraviolet light by Zn-enhanced fluorescence. The bands of LRγ4–8 and PBPs were identified by MS analysis and are indicated. For gel source data, see Supplementary Fig. 1. f, MS analysis of purified PBSs. Two batches of sample were analysed. The similar results confirmed the consistency of our purification method. g, A representative motion-corrected electron micrograph of PBSs. Scale bar, 100 nm. h, Typical good, reference-free 2D class averages from single-particle images of PBSs. Scale bar, 20 nm. i, Gold-standard Fourier shell correlation (FSC) curves of the cryo-EM map of PBS with or without C2 symmetry imposed. j, Euler angle distribution of particles contributing to the final reconstruction of PBS. Each cylinder represents one view and the size of the cylinder is proportional to the number of particles for that view. k, The workflow for 3D classification. l, The crystal structure of allophycocyanin (PDB 1B33) can be fitted well with the density map of PBS in different view.
Extended Data Figure 2 Cryo-EM analysis of PBS using Tecnai Krios microscope and Gatan K2 Summit direct electron detector.
a, A representative motion-corrected electron micrograph of PBSs. Scale bar, 100 nm. b, Fourier power spectrum of the micrograph showing the Thon ring extending to 3 Å. c, Typical good, reference-free 2D class averages from single particle PBS images. Scale bar, 100 nm. d, Gold-standard FSC curve of the final cryo-EM map with a value of 0.143 at a resolution of 3.5 Å and the FSC curve between the refined model and the final map with a value of 0.5 at a resolution of 4.0 Å. e, Colour-coded final 3D reconstruction of a PBS showing local resolution in different views. f, The workflow for the 2D and 3D classifications for cryo-EM data processing. g, The masking strategy for dealing with flexible regions of PBS. Several local masks for different parts were applied and further, reduced-size masks were applied to some specific areas, resulting in improved resolution and local map qualities.
Extended Data Figure 3 Overall structure of PBS.
a, 3D cryo-EM density map of PBS. b, Top view of the atomic model of PBS. c, Schematic diagram showing the organization of the rods and core cylinders from three perpendicular views. d, The organization of the rods and core displayed in surface representation. e, The organization of the key structural elements (rods a–c and the core) of PBS displayed in surface representation. The protein components are colour-coded as shown. f, Structures of the 20 well-resolved linker proteins. Atomic models of these linker proteins superimposed with their segmented cryo-EM densities (transparent grey). Domains of αLCM, Pfam00427 (00427), Pfam01383 (01383), CBDγ, and FAS1 are labelled. g, Schematic drawing of structural elements of the PBS subunits. PBS subunits are classified into four groups according to their positions and functions: phycobiliproteins (PBPs), linkers in the core, rod linkers and rod–core linkers. Different structural elements are separately coloured.
Extended Data Figure 4 Structures and binding partners of rod linker proteins.
a, Comparison of the Pfam00427 domain of LR1 with the crystal structure of the Pfam00427 domain (PDB 3NPH) from linker LR30 from Synechocystis sp. PCC 6803. The two structures are almost identical. b, Comparison of the Pfam01383 domain of LR1 with the crystal structure of the Pfam01383 domain (PDB 1B33) from the PBS of Mastigocladus laminosus. The long α-helix and three β-strands are labelled. c, LR2 is located in the inner hexamer of rod Rg (Rg1). Its C-terminal extended loop and a small β-sheet that is composed of two antiparallel β-strands at the very C terminus contact the outer surface of hexamers Rb1 and Rb2. d, LR3 is located in the inner hexamer of rod Rf (Rf1) and its C terminus ends with a long α-helix. The N-terminal part of the long helix inserts into the area between hexamers Rc1 and Rf1 and the C-terminal part of the long helix forms a coiled-coil structure (red oval) with its symmetric counterpart LR3′ located in hexamer Rf1′. These structures stabilize the rod pair Rf and Rf′ by association with each other and anchor them to rods Rc and Rc′. e, Structural superimposition of LRγ4–8 proteins, showing the almost identical CBDγ structures and different conformations outside the CBDγ. f–h, LRγ6–8 with their CBDγ domains located in hexamers Hd, Ha, and Hb, respectively. The C-terminal loop of LRγ6 anchors on the outer side of the innermost hexamer of rod Rc (Rc1) (f); the C terminus of LRγ7 contains several short α-helices and interacts with the surface of hexamers Rd2 and Rg1 (g); and an α-helix of the N terminus of LRγ8 inserts into the space between the two hexamers Ra2 and Ra3 (h). i, Structural superimposition of the N-terminal domain of LR9 and FAS1 domain (PDB 2LTB). j, LR9 sits between rods Ra, Rb, Rd and Rg and interacts with them. The FAS1 domain is circled by a red dashed line.
Extended Data Figure 5 Extensive interactions between rod linker proteins and PBPs in rod Rb.
a, b, Expanded views of interactions between the CTR of LR1 and trimer Rb1II, corresponding to Fig. 2c. c–e, Expanded views of interactions between the intertwined NTE and CTE of the N-terminal region of LR1 and trimer Rb2I, corresponding to Fig. 2d. f, g, Expanded views of interactions between the Pfam00427 domain of LR1 and trimer Rb2I, corresponding to Fig. 2f. h, Expanded view of interactions between the Pfam00427 domain of LR1 and trimer Rb2II, corresponding to Fig. 2g. i–k, Expanded views of interactions between the CBDγ of LRγ4 and trimer Rb3I, corresponding to Fig. 2i. l, Expanded view of interactions between the CBDγ of LRγ4 and trimer Rb3II, corresponding to Fig. 2j. Helix X is labelled and highlighted. Residues involved in atomic interaction sites are shown in the stick representation. α, β and H denote α-subunit, β-subunit and helix, respectively. m, CBDγ contacts the inner face of the rod hexamer in a symmetrical fashion. When the first repeat of CBDγ of LRγ4 (LRγ4-CBDγ-1Rep) is aligned with the second repeat of CBDγ of LRγ4 (LRγ4-CBDγ-2Rep), the trimers (Rb3I and Rb3II) that they interact with are also aligned well.
Extended Data Figure 6 Sequence alignment of LRγ4–8 and the γ-subunits from different red algal species and the bilins in rod b.
a, Amino acid sequence alignment. The predicted transit peptides deduced from the sequence alignment and the CBDγ domains based on the structures are labelled. Conserved cysteine residues covalently attached to chromophores are highlighted in blue. Note that one of the cysteine residues in the CBDγ of LRγ7 is replaced by serine (highlighted in yellow), consistent with the fact that only four chromophores are found in LRγ7 as compared with other CBDγ domains, which have five chromophores. P34784, P92928, γ-subunits from Aglaothamnion neglectum; R7Q9W8, R7QDA3, γ-subunits from Chondrus crispus; Q7XZS8, P83592, γ-subunits from Corallina officinalis. b, Distances between some bilins in Rb. Magenta, bilins of LRγ; orange, bilins of β-subunits in phycoerythrin hexamers; yellow, bilins of α-subunits in phycoerythrin hexamers; dark green, bilins of β-subunits in phycocyanin hexamer; cyan, bilins of α-subunits in phycocyanin hexamer; pink, bilins nearest to the core in phycocyanin hexamer. The numbers near the dashed lines indicate the distances (Å) between bilins on linker proteins and adjacent bilins on PBPs. c–f, Cryo-EM densities (mesh) of bilins (stick) in the phycoerythrin trimer Rb2II (c), the phycocyanin trimer Rb1II (d), the LRγ4 protein (e) and the LRγ5a protein (f).
Extended Data Figure 7 Interactions between LRC1–3 and the core.
a, Superimposition of the αAPC subunits and the spatial positions of their interacting LRC proteins. The alignments were performed using the αAPC subunit as a reference. Note that the contacting helices of all LRC proteins (black arrow) are also aligned very well. Inset shows the groove formed by helices B and E of the αAPC subunit. b, c, Interactions between the αAPC subunit and the helices of LRC2 (b) and LRC3 (c). The residues involved in the interaction of LRC proteins are coloured green and shown in stick representation. The αAPC are shown in surface representation, and the residues involved in the interaction are red. d, e, Sequence alignment of LRC1–3 (d) and αAPC (e) from different red algal and cyanobacterial species. The residues involved in the interaction are indicated by asterisks and highlighted yellow.
Extended Data Figure 8 Structures and assembling roles of LRC4, LRC5 and LRC6.
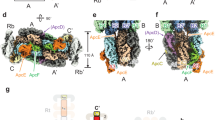
a, Overview of LRC4 and LRC5 wrapping their target protein components. S1′ and S2′ are single α- and β-subunits, respectively. b, The long α-helix at the middle of LRC4′ spans across one α-subunit of core trimer B1. c, The long α-helix at the middle of LRC5 spans across one α-subunit of the trimer B2. d, The N-terminal extension of LRC4′ meanders at the surface of trimer Rb1′I and LRC1b′. e, The C-terminal extension of LRC5 interacts with core layer B2, rod trimer Rb1′I, and LRC1b′. f, The N-terminal extension of LRC5 wraps around the surfaces of trimers Rc1I and B1, and also contacts LRC1c. g, The structure of LRC6. h, Structural superimposition of the FAS1 domain of LRC6 and the known FAS1 domain (PDB 2LTB). α-helices and β-strands are labelled. i, LRC6′, like a wedge, is inserted into the area between trimers Rd1′I and Re1I and layer A′3 of the core, and makes extensive contacts with the surrounding protein components, including A′3, LCM′, LC′, LRC2′, Rd1′I and Re1I with its extended loops on both sides of the FAS1 domain.
Extended Data Figure 9 Structure of the core.
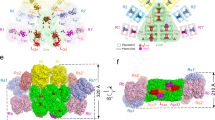
a, Structure of the core in four perpendicular views shows the assembly and arrangement of the core layers. The bottom view (the same as the view of Fig. 4c) shows the surface electrostatic potential distributions of the core. The surfaces of αLCM molecules are circled. The surface electrostatic potentials were calculated in PyMol. Red, negative electrostatic potential; blue, positive electrostatic potential; grey, non-polar. b, A cartoon model of the core of a hemidiscoidal PBS (adapted from ref. 1). Note that the basal cylinder has four trimers. c, The inter-cylinder interaction in the core is symmetrical: trimers B1, A2 and A′1 interact with one another and trimers B2, A′2 and A1 interact with one another. The association is mediated by the interaction between three α-subunits (red circle) and two pairs of α−β subunits (black circle). d, Comparison of the three regions of LCM with the crystal structure of the Pfam00427 domain. The structures of the three regions of LCM (LCM-Reg1–3) are displayed as indicated and they plus the crystal structure of the Pfam00427 domain (PDB 3NPH) are aligned with one another. e–g, Close-up views of three PCB pockets of the β-subunits boxed by dashed lines in Fig. 4d: β1 (a), β2 (b) and ApcF (c). The bilins and their surrounding residues are shown in ball-stick representation and stick representation, respectively. Their cryo-EM densities (mesh) superimposed with the respective atomic models (stick) are shown in the lower panels. h, Structural superimposition of ApcF and βAPC shows that ApcF has an extension at the tip of the G–H helix hairpin. i, j, Cryo-EM densities (mesh) of the bilins in αLCM and the α3 subunit of core layer A2, and their surrounding residues, corresponding to Fig. 4f, superimposed with their respective atomic models (stick; coloured as in Fig. 4).
Extended Data Figure 10 Identification of the APC subunits.
Cryo-EM densities (mesh) of representative regions of the APC subunits, superimposed with respective atomic models (stick). Note the different densities of the residues labelled in red.
Extended Data Figure 11 Plausible energy transfer between the individual hexamer and rods; between rods Rd and Re and the core; and between rods.
a, Distances (Å) between bilins at the interfaces of the individual hexamer Ha and rod Rd (left), and Hb and Ra (right). b, Plausible energy transfer paths from bilins on Rd and Re′ to the core. Inset panels indicate the distances (Å) between two adjacent bilins. c, Plausible energy transfer paths between bilins on Rd and Rb. Inset panels indicate the distances (Å) between two adjacent bilins. d, Plausible energy transfer from bilins on Re to Ra or Rc. Inset panels indicate the distances (Å) between two adjacent bilins.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 (summary of model building for the phycobilisome), Supplementary Table 2 (25 PBS protein sequences), and Supplementary Figure 1 (the uncropped SDS PAGE gels for Extended Data Figure 1d, e.). (PDF 2285 kb)
Rights and permissions
About this article
Cite this article
Zhang, J., Ma, J., Liu, D. et al. Structure of phycobilisome from the red alga Griffithsia pacifica. Nature 551, 57–63 (2017). https://doi.org/10.1038/nature24278
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24278
This article is cited by
-
ApcE plays an important role in light-induced excitation energy dissipation in the Synechocystis PCC6803 phycobilisomes
Photosynthesis Research (2024)
-
Crystallographic and biochemical analyses of a far-red allophycocyanin to address the mechanism of the super-red-shift
Photosynthesis Research (2024)
-
Relationship between non-photochemical quenching efficiency and the energy transfer rate from phycobilisomes to photosystem II
Photosynthesis Research (2024)
-
In situ structure of the red algal phycobilisome–PSII–PSI–LHC megacomplex
Nature (2023)
-
Cryo-EM and femtosecond spectroscopic studies provide mechanistic insight into the energy transfer in CpcL-phycobilisomes
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.