Abstract
Recent studies on the controllability of complex systems offer a powerful mathematical framework to systematically explore the structure–function relationship in biological, social, and technological networks1,2,3. Despite theoretical advances, we lack direct experimental proof of the validity of these widely used control principles. Here we fill this gap by applying a control framework to the connectome of the nematode Caenorhabditis elegans4,5,6, allowing us to predict the involvement of each C. elegans neuron in locomotor behaviours. We predict that control of the muscles or motor neurons requires 12 neuronal classes, which include neuronal groups previously implicated in locomotion by laser ablation7,8,9,10,11,12,13, as well as one previously uncharacterized neuron, PDB. We validate this prediction experimentally, finding that the ablation of PDB leads to a significant loss of dorsoventral polarity in large body bends. Importantly, control principles also allow us to investigate the involvement of individual neurons within each neuronal class. For example, we predict that, within the class of DD motor neurons, only three (DD04, DD05, or DD06) should affect locomotion when ablated individually. This prediction is also confirmed; single cell ablations of DD04 or DD05 specifically affect posterior body movements, whereas ablations of DD02 or DD03 do not. Our predictions are robust to deletions of weak connections, missing connections, and rewired connections in the current connectome, indicating the potential applicability of this analytical framework to larger and less well-characterized connectomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Caldarelli, G. Scale-Free Networks: Complex Webs in Nature and Technology (Oxford Univ. Press, 2007)
Cohen, R. & Havlin, S. Complex Networks: Structure, Robustness and Function (Cambridge Univ. Press, 2010)
Liu, Y.-Y. & Barabási, A.-L. Control principles of complex networks. Rev. Mod. Phys. 88, 035006 (2016)
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Phil. Trans. R. Soc. Lond. B 314, 1–340 (1986)
Chen, B. L. J. Neuronal Network of C. elegans: From Anatomy to Behavior. PhD thesis, Cold Spring Harbor Laboratory (2007)
Varshney, L. R., Chen, B. L., Paniagua, E., Hall, D. H. & Chklovskii, D. B. Structural properties of the Caenorhabditis elegans neuronal network. PLoS Comput. Biol. 7, e1001066 (2011)
Chalfie, M. et al. The neural circuit for touch sensitivity in Caenorhabditis elegans. J. Neurosci. 5, 956–964 (1985)
Zhen, M. & Samuel, A. D. C. elegans locomotion: small circuits, complex functions. Curr. Opin. Neurobiol. 33, 117–126 (2015)
Bargmann, C. I. & Avery, L. Laser killing of cells in Caenorhabditis elegans. Methods Cell Biol. 48, 225–250 (1995)
Wicks, S. R. & Rankin, C. H. Integration of mechanosensory stimuli in Caenorhabditis elegans. J. Neurosci. 15, 2434–2444 (1995)
Tsalik, E. L. & Hobert, O. Functional mapping of neurons that control locomotory behavior in Caenorhabditis elegans. J. Neurobiol. 56, 178–197 (2003)
Wakabayashi, T., Kitagawa, I. & Shingai, R. Neurons regulating the duration of forward locomotion in Caenorhabditis elegans. Neurosci. Res. 50, 103–111 (2004)
Haspel, G., O’Donovan, M. J. & Hart, A. C. Motoneurons dedicated to either forward or backward locomotion in the nematode Caenorhabditis elegans. J. Neurosci. 30, 11151–11156 (2010)
Gao, J., Liu, Y.-Y., D’Souza, R. M. & Barabási, A.-L. Target control of complex networks. Nat. Commun. 5, 5415 (2014)
Coron, J.-M. Control and Nonlinearity (American Mathematical Society, 2009)
Whalen, A. J., Brennan, S. N., Sauer, T. D. & Schiff, S. J. Observability and controllability of nonlinear networks: the role of symmetry. Phys. Rev. X 5, 011005 (2015)
Muldoon, S. F. et al. Stimulation-based control of dynamic brain networks. PLoS Comput. Biol. 12, e1005076 (2016)
Kalman, R. E. Mathematical description of linear dynamical systems. J. Soc. Indus. Appl. Math. A 1, 152–192 (1963)
Chew, Y. L. et al. Recordings of Caenorhabditis elegans locomotor behaviour following targeted ablation of single motorneurons. Sci. Data 4, 170156 (2017)
Yemini, E., Jucikas, T., Grundy, L. J., Brown, A. E. & Schafer, W. R. A database of Caenorhabditis elegans behavioral phenotypes. Nat. Methods 10, 877–879 (2013)
Stephens, G. J., Johnson-Kerner, B., Bialek, W. & Ryu, W. S. Dimensionality and dynamics in the behavior of C. elegans. PLoS Comput. Biol. 4, e1000028 (2008)
Brown, A. E., Yemini, E. I., Grundy, L. J., Jucikas, T. & Schafer, W. R. A dictionary of behavioral motifs reveals clusters of genes affecting Caenorhabditis elegans locomotion. Proc. Natl Acad. Sci. USA 110, 791–796 (2013)
Gray, J. M., Hill, J. J. & Bargmann, C. I. A circuit for navigation in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 102, 3184–3191 (2005)
Huang, K. M., Cosman, P. & Schafer, W. R. Machine vision based detection of omega bends and reversals in C. elegans. J. Neurosci. Methods 158, 323–336 (2006)
Yan, G. et al. Spectrum of controlling and observing complex networks. Nat. Phys. 11, 779–786 (2015)
Gu, S. et al. Controllability of structural brain networks. Nat. Commun. 6, 8414 (2015)
Pósfai, M. & Hövel, P. Structural controllability of temporal networks. New J. Phys. 16, 123055 (2014)
Pan, Y. & Li, X. Structural controllability and controlling centrality of temporal networks. PLoS ONE 9, e94998 (2014)
Li, A., Cornelius, S. P., Liu, Y. Y., Wang, L. & Barabási, A. L. The fundamental advantages of temporal networks. arXiv, 1607.06168 (2016)
Driscoll, M. & Kaplan, J. in C. elegans II (Cold Spring Harbor monograph series 33) (eds Riddle, D. L. et al.) Ch. 23 (Cold Spring Harbor Laboratory Press, 1997)
Acknowledgements
We thank M. Angulo, J. Gao, Y.-Y. Liu, J.-J. Slotine, K. Albrecht, S. P. Cornelius, and A. Li for discussions, and L. Grundy, A. Brown, and E. Yemini for help with analysis of tracking data. We are grateful to V. Butler and the Caenorhabditis Genetics Center, which is funded by National Institutes of Health Office of Research Infrastructure Programs (P40 OD010440), for C. elegans strains. This work is supported by the John Templeton Foundation: Mathematical and Physical Sciences grant number PFI-777; European Commission grant number 641191 (CIMPLEX); Medical Research Council grant number MC-A023-5PB91; Wellcome Trust grant number WT103784MA. P.E.V. is supported by the Medical Research Council grant number MR/K020706/1. Y.L.C. is supported by an EMBO Long Term Fellowship.
Author information
Authors and Affiliations
Contributions
A.-L.B., G.Y., and P.E.V. conceived the project. G.Y. did the control analysis. P.E.V. and E.K.T. analysed the results. W.R.S. conceived the experimental validation. Y.L.C. and D.S.W. planned and performed the new experiments. Y.L.C. and W.R.S. analysed the experimental data, and W.R.S. and A.-L.B. discussed the results. A.-L.B., E.K.T., W.R.S., P.E.V., and G.Y. wrote the manuscript, Y.L.C. and D.S.W. edited it. G.Y., Y.L.C., and D.S.W. wrote the Supplementary Information, and W.R.S., E.K.T., and P.E.V. edited it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks C. Bargmann, E. Izquierdo and M. Zhen for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 C. elegans connectome.
The filled nodes are the previously known neurons involved in the worm’s response to gentle touch.
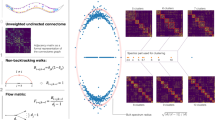
Extended Data Figure 2 Structural controllability, the construction of the linking graph, and the derivation of the controllability criterion.
a, b, The blue nodes receive external signals and the pink nodes are those we aim to control. Thus, S = 1 and M = 2 for both networks. c, The construction of the linking graph for the network in a. d, The calculation of the linking size can be mapped into a multi-source-multi-sink max-flow problem, with the constraint that the capacity of each node and each edge is one. The red edges show the two disjoint paths that achieve the maximum flow. e, f, Schematic picture for the derivation of the lower bound z*.
Extended Data Figure 3 Control theoretic mechanisms of the loss of muscular control.
Loss of control induced by the ablation of the AVA (a) or AS (b) neuronal class in C. elegans.
Extended Data Figure 4 Control theoretic mechanisms of the loss of muscular control.
Loss of control induced by the ablation of the DA (a) or DB (b) neuronal class in C. elegans.
Extended Data Figure 5 Control theoretic mechanisms of the loss of muscular control.
Loss of control induced by the ablation of the VA (a), VB (b), or VD (c) neuronal class in C. elegans.
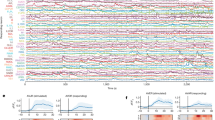
Extended Data Figure 6 Illustrative examples of the behavioural phenotypes observed for PDB- and DD-ablated animals.
Time series plots of sample videos, and still images from these videos, illustrating the locomotion abnormalities resulting from ablation. The green dot indicates the animal’s head, and the red dot in the mid-body indicates the ventral side. a, For PDB-ablated animals compared with mock-ablated controls, we observed differences in Eigen projection 1, which describes the large wavelength body bends that occur during turning. The lower negative values observed in PDB ablations indicate a loss of the ventral bias to these turns. Still images show PDB-ablated animals making a large dorsal turn, whereas turns in control animals are usually ventral. The videos used here (from left to right) are ‘mockPDB_onfood_L_2016_ 11_03__14_16_37___7___1’, ‘ablPDB_onfood_L_2016_11_03__14_40_04___4__2’, and ‘ablPDB_onfood_L_2016_11_04__14_28_26___5___1’. b, DD5-ablated animals showed lower values for Eigen projection 4, which captures the small wavelength oscillations in the head and tail. The lower values indicate a reduction in amplitude of tail oscillations compared with controls, that is, a characteristic stiff tail appearance. The videos shown here (from left to right) are ‘mockDD_onfood_L_2016_10_29__13_13_35 ___7___6’, ‘DD2_onfood_R_2016_10_30__12_13_57___7___4’, and ‘DD5_onfood_ L_2016_10_29__13_13_25___5___6’.
Extended Data Figure 7 Predictive robustness against random deletions, additions, and rewiring of links.
The vertical axis represents the probability that each of the predicted neuron classes remains significant in the controllability of muscles (a–c) or motor neurons (d, e) after the network is altered. The horizontal axis denotes the number of deleted weak links, added links, or rewired links between neurons in C. elegans connectome. Each probability is calculated from 200 independent runs.
Extended Data Figure 8 The role of individual neurons within the DB, PDB, AVA, and AS neuronal classes.
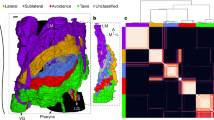
Role of neurons within the DB (a), PDB (b), AVA (c), and AS (d) neuronal classes in the loss of muscular controllability in C. elegans. The shade of green represents the probability with which control is lost over each muscle following the ablation of individual neurons. Each cross indicates a direct connection between a neuron and a muscle cell. Note that there are other muscles directly connected to the neurons but not shown here, because of zero probability for reduced control over these muscles following ablation of these neuronal classes.
Extended Data Figure 9 The role of individual neurons within the DA and DD neuronal classes.
Role of neurons within the DA (a) and DD (b) neuronal classes in the loss of muscular controllability in C. elegans. The shade of green represents the probability with which the control is lost over each muscle induced by the ablation of individual neurons. Each cross indicates a direct connection between a neuron and a muscle cell. Note that there are other muscles directly connected to the neurons but not shown here, because of zero probability for reduced control over these muscles following ablation of these neuronal classes.
Extended Data Figure 10 The role of individual neurons within the VA, VB, and VD neuronal classes.
Role of neurons within the VA (a), VB (b) and VD (c) neuronal classes in the loss of muscular controllability in C. elegans. The shade of green represents the probability with which the control is lost over each muscle induced by the ablation of individual neurons. Each cross indicates a direct connection between a neuron and a muscle cell. Note that there are other muscles directly connected to the neurons but not shown here, because of zero probability for reduced control over these muscles following ablation of these neuronal classes.
Supplementary information
Supplementary Information
This file contains the Supplementary Methods and Discussion. It provides details of the mathematical framework, experimental set-up and detailed results, and additional analyses. It also contains Supplementary Tables 1-3, which provide the statistical descriptions of the experimental observations. (PDF 742 kb)
Mock ablation (DD) sample video
Shown is a short clip from the video mockDD_onfood_L_2016_10_29__13_13_35___7___6_seg.avi. The entire clip is available at https://figshare.com/s/72716a92be1ab0f1e1d4#/articles/5087020 (MP4 614 kb)
DD5 ablation sample video
Shown is a short clip from the video DD5_onfood_L_2016_10_29__13_13_25___5___6_seg. The entire clip is available at https://figshare.com/s/72716a92be1ab0f1e1d4#/articles/5087020.Shown is a short clip from the video DD5_onfood_L_2016_10_29__13_13_25___5___6_seg. The entire clip is available at https://figshare.com/s/72716a92be1ab0f1e1d4#/articles/5087020. (MP4 336 kb)
DD2 ablation sample video
Shown is a short clip from the video DD2_onfood_R_2016_10_30__12_13_57___7___4_seg. The entire clip is available at https://figshare.com/s/72716a92be1ab0f1e1d4#/articles/5087020. (MP4 382 kb)
Mock ablation (PDB) sample video
Shown is a short clip from the video mockPDB_on_food_L_2016_11_03__14_16_37___7___1_seg. The animal is executing a ventral omega turn. The entire clip is available at https://figshare.com/s/72716a92be1ab0f1e1d4#/articles/5087020 (MP4 392 kb)
PDB ablation sample video 1
Shown is a short clip from the video ablPDB_on_food_L_2016_11_03__14_40_04___4___2_seg. The animal is executing a dorsal omega turn. The entire clip is available at https://figshare.com/s/72716a92be1ab0f1e1d4#/articles/5087020. (MP4 376 kb)
PDB ablation sample video 2
Shown is a short clip from the video ablPDB_on_food_L_2016_10_22__13_23_05___5___3_seg. The animal is executing a dorsal omega turn. The entire clip is available at https://figshare.com/s/72716a92be1ab0f1e1d4#/articles/5087020 (MP4 0 kb)
Source data
Rights and permissions
About this article
Cite this article
Yan, G., Vértes, P., Towlson, E. et al. Network control principles predict neuron function in the Caenorhabditis elegans connectome. Nature 550, 519–523 (2017). https://doi.org/10.1038/nature24056
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24056
This article is cited by
-
Measuring criticality in control of complex biological networks
npj Systems Biology and Applications (2024)
-
The low-rank hypothesis of complex systems
Nature Physics (2024)
-
Controllability in attention deficit hyperactivity disorder brains
Cognitive Neurodynamics (2024)
-
Network models to enhance the translational impact of cross-species studies
Nature Reviews Neuroscience (2023)
-
Alprazolam modulates persistence energy during emotion processing in first-degree relatives of individuals with schizophrenia: a network control study
Molecular Psychiatry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.