Abstract
Chromatin is traditionally viewed as a nuclear entity that regulates gene expression and silencing1,2,3. However, we recently discovered the presence of cytoplasmic chromatin fragments that pinch off from intact nuclei of primary cells during senescence4,5, a form of terminal cell-cycle arrest associated with pro-inflammatory responses6. The functional significance of chromatin in the cytoplasm is unclear. Here we show that cytoplasmic chromatin activates the innate immunity cytosolic DNA-sensing cGAS–STING (cyclic GMP–AMP synthase linked to stimulator of interferon genes) pathway, leading both to short-term inflammation to restrain activated oncogenes and to chronic inflammation that associates with tissue destruction and cancer. The cytoplasmic chromatin–cGAS–STING pathway promotes the senescence-associated secretory phenotype in primary human cells and in mice. Mice deficient in STING show impaired immuno-surveillance of oncogenic RAS and reduced tissue inflammation upon ionizing radiation. Furthermore, this pathway is activated in cancer cells, and correlates with pro-inflammatory gene expression in human cancers. Overall, our findings indicate that genomic DNA serves as a reservoir to initiate a pro-inflammatory pathway in the cytoplasm in senescence and cancer. Targeting the cytoplasmic chromatin-mediated pathway may hold promise in treating inflammation-related disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Sen, P., Shah, P. P., Nativio, R. & Berger, S. L. Epigenetic mechanisms of longevity and aging. Cell 166, 822–839 (2016)
Benayoun, B. A., Pollina, E. A. & Brunet, A. Epigenetic regulation of ageing: linking environmental inputs to genomic stability. Nat. Rev. Mol. Cell Biol. 16, 593–610 (2015)
Berger, S. L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007)
Ivanov, A. et al. Lysosome-mediated processing of chromatin in senescence. J. Cell Biol. 202, 129–143 (2013)
Dou, Z. et al. Autophagy mediates degradation of nuclear lamina. Nature 527, 105–109 (2015)
Campisi, J. Aging, cellular senescence, and cancer. Annu. Rev. Physiol. 75, 685–705 (2013)
Shah, P. P. et al. Lamin B1 depletion in senescent cells triggers large-scale changes in gene expression and the chromatin landscape. Genes Dev. 27, 1787–1799 (2013)
Capell, B. C. et al. MLL1 is essential for the senescence-associated secretory phenotype. Genes Dev. 30, 321–336 (2016)
Sadaie, M. et al. Redistribution of the lamin B1 genomic binding profile affects rearrangement of heterochromatic domains and SAHF formation during senescence. Genes Dev. 27, 1800–1808 (2013)
Tasdemir, N. et al. BRD4 connects enhancer remodeling to senescence immune surveillance. Cancer Discov. 6, 612–629 (2016)
Shimi, T. et al. The role of nuclear lamin B1 in cell proliferation and senescence. Genes Dev. 25, 2579–2593 (2011)
Freund, A., Laberge, R. M., Demaria, M. & Campisi, J. Lamin B1 loss is a senescence-associated biomarker. Mol. Biol. Cell 23, 2066–2075 (2012)
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013)
Wu, J. et al. Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013)
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009)
Barber, G. N. STING: infection, inflammation and cancer. Nat. Rev. Immunol. 15, 760–770 (2015)
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008)
Hatch, E. M., Fischer, A. H., Deerinck, T. J. & Hetzer, M. W. Catastrophic nuclear envelope collapse in cancer cell micronuclei. Cell 154, 47–60 (2013)
Zhang, C. Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015)
Freund, A., Patil, C. K. & Campisi, J. p38MAPK is a novel DNA damage response-independent regulator of the senescence-associated secretory phenotype. EMBO J. 30, 1536–1548 (2011)
Chen, Y. et al. p38 inhibition provides anti-DNA virus immunity by regulation of USP21 phosphorylation and STING activation. J. Exp. Med. 214, 991–1010 (2017)
Benci, J. L. et al. Tumor interferon signaling regulates a multigenic resistance program to immune checkpoint blockade. Cell 167, 1540–1554 (2016)
Rodier, F. et al. Persistent DNA damage signalling triggers senescence-associated inflammatory cytokine secretion. Nat. Cell Biol. 11, 973–979 (2009)
Chien, Y. et al. Control of the senescence-associated secretory phenotype by NF-kappaB promotes senescence and enhances chemosensitivity. Genes Dev. 25, 2125–2136 (2011)
Gray, E. E. et al. The AIM2-like receptors are dispensable for the interferon response to intracellular DNA. Immunity 45, 255–266 (2016)
Kang, T. W. et al. Senescence surveillance of pre-malignant hepatocytes limits liver cancer development. Nature 479, 547–551 (2011)
Acosta, J. C. et al. A complex secretory program orchestrated by the inflammasome controls paracrine senescence. Nat. Cell Biol. 15, 978–990 (2013)
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012)
Gluck, S. et al. Innate immune sensing of cytosolic chromatin fragments through cGAS promotes senescence. Nat. Cell Biol. 19, 1061–1070 (2017)
Yang, H., Wang, H., Ren, J., Chen, Q. & Chen, Z. J. cGAS is essential for cellular senescence. Proc. Natl Acad. Sci. USA 114, E4612–E4620 (2017)
Debacq-Chainiaux, F., Erusalimsky, J. D., Campisi, J. & Toussaint, O. Protocols to detect senescence-associated beta-galactosidase (SA-βgal) activity, a biomarker of senescent cells in culture and in vivo. Nat. Protocols 4, 1798–1806 (2009)
Abe, T. et al. STING recognition of cytoplasmic DNA instigates cellular defense. Mol. Cell 50, 5–15 (2013)
Wangensteen, K. J., Zhang, S., Greenbaum, L. E. & Kaestner, K. H. A genetic screen reveals Foxa3 and TNFR1 as key regulators of liver repopulation. Genes Dev. 29, 904–909 (2015)
Freund, A., Orjalo, A. V., Desprez, P. Y. & Campisi, J. Inflammatory networks during cellular senescence: causes and consequences. Trends Mol. Med. 16, 238–246 (2010)
Rai, T. S. & Adams, P. D. ChIP-sequencing to map the epigenome of senescent cells using benzonase endonuclease. Methods Enzymol. 574, 355–364 (2016)
Chen, Q. et al. Carcinoma–astrocyte gap junctions promote brain metastasis by cGAMP transfer. Nature 533, 493–498 (2016)
Acknowledgements
We acknowledge S. Prouty for histology studies, E. Browning for small animal imaging, the Cell & Developmental Biology Microscopy Core, and the high-throughput screening core for technical assistance. We thank A. Brunet, J. Cross, J. Guerriero, I. Harel, E. J. Wherry, and W.-X. Zong for discussions and reading the manuscript. The Penn Skin Biology and Diseases Resource-based Center is supported by 1P30AR069589-01 (S.M.). Z.D. is supported by a fellow award from the Leukemia & Lymphoma Society and by National Institutes of Health (NIH) K99AG053406. S.L.B., P.D.A., and B.A.G. are supported by NIH P01AG031862. S.L.B. is supported by NIH CA078831, and B.A.G. by NIH P01CA196539. S.L.B. acknowledges support by the Glenn Foundation and the Ellison Foundation for research in ageing.
Author information
Authors and Affiliations
Contributions
Z.D., Z.Z., P.D.A., and S.L.B. conceived the project. Z.D. and K.G. performed most of the experiments. M.G.V. contributed Fig. 3a, b and Extended Data Fig. 5b, c. J.Z. contributed Fig. 4g and Extended Data Figs 9 and 10. P.S., B.C., C.X., and Y.Lan contributed Fig. 2d, e and Extended Data Fig. 3e, f. K.W. and K.K. advised on Fig. 3c–h and Extended Data Fig. 6. J.S., Y.Lin, and B.G contributed Fig. 1b, Extended Data Figs 1g and 5a. M.X., J.K., T.J., M.S.-C., J.T.S., and S.M. contributed histology analyses in Fig. 3 and Extended Data Fig. 6. K.M.T. contributed plasmids in Figs 1, 2, 3. G.B. contributed STING mice and advised on in vivo experiments. Z.D., P.D.A., and S.L.B. composed the manuscript. All authors reviewed the manuscript and discussed the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J. van Deursen, K.-P. Hopfner and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 CCF–cGAS–STING activation in senescence.
a, Confocal microscopy analyses of primary mouse embryonic fibroblasts. CCF indicated by arrows. b, Quantification of IMR90 undergoing replicative senescence. PD, population doubling. c, Microscopy-based quantification of parameters as indicated. d–f, Confocal microscopy analyses of BJ (d), IMR90 stained for endogenous cGAS (e), and mitotic IMR90 (f) cells. g, cGAMP detection by nano-LC–MS. MS2 spectra were confirmed for cGAMP. h, IMR90 cells were analysed by immunoblotting. STING blots were performed under non-reducing condition. * STING dimer. i, j, Confocal microscopy images of STING in IMR90 (i) and BJ (j) cells. k, Cells as in Fig. 1c were quantified under microscopy. Bar graphs show mean values of four different fields with over 200 cells and s.d. Scale bars, 10 μm.
Extended Data Figure 2 Interferon genes are repressed in senescent human fibroblasts.
a, b, ER:HRasV12 IMR90 cells were induced by OHT and quantified for CCF (a) or analysed by RT–qPCR (b). c, d, IMR90 cells were treated with etoposide and analysed similarly as above. Results shown in b, d are from triplicate technical replicates, and were normalized to the untreated sample. Bar graphs (a, c) show mean values of four different fields with over 200 cells and s.d. e, RNA-seq values of indicated genes. n = 3; error bars, s.d. f, IMR90 cells were treated with a p38 inhibitor. *P < 0.005, **P < 0.0001, compared with dimethylsulfoxide (DMSO). g, Cultured media from proliferating or senescent IMR90 cells were administered to proliferating cells, followed by dsDNA90 transfection. *P < 0.0001, compared with control media. h, IMR90 cells were incubated with recombinant IL1α and transfected with dsDNA90. *P < 0.01, **P < 0.0001, compared with no-IL1α transfected groups. f–h, RT–qPCR analyses with mean values and s.d.; n = 3; unpaired two-tailed Student’s t-test. i, Schematic illustration of interferon repression in senescence.
Extended Data Figure 3 CCF–cGAS–STING pathway activates the SASP.
a, Cells transfected with dsDNA90 were analysed by RT–qPCR. b, Cells as in Fig. 2c were stained for SA-β-gal and quantified. c, IMR90 cells were analysed by RT–qPCR. *P < 0.0001, compared with sh-NTC etoposide. d, IMR90 cells were analysed by immunoblotting. e, Track views of indicated genes from RNA-seq. f, Heat map representation of SASP genes. g, Cultured media were analysed by IL8 immunoblotting. h, Related to Fig. 2f, quantification of secreted cytokines. *P < 0.001, **P < 0.0001, compared with sh-NTC. i, j, RT–qPCR analyses of established senescent cells. *P < 0.005, **P < 0.0001, compared with +OHT sh-NTC. j, IFI16 does not regulate the SASP. k, IFI16 plays a regulatory but not essential role upon dsDNA90 transfection. Bar graphs show mean values with s.d.; n = 3; one-way ANOVA coupled with Tukey’s post hoc test for c, h; unpaired two-tailed Student’s t-test for i.
Extended Data Figure 4 Role of CCF–cGAS–STING in SASP activation.
a–c, IMR90 cells were analysed by confocal microscopy. *P < 0.005, compared with sh-NTC HRasV12. d, p65 Chromatin immunoprecipitation–qPCR analyses. *P < 0.05, **P < 0.01, compared with sh-NTC. e–h, IMR90 cells overexpressed with lamin B1 were analysed by immunofluorescence (e), immunoblotting (f, g), or RT–qPCR (h). *P < 0.01, **P < 0.001. i, IMR90 cells were transfected with dsDNA90 and analysed 4 days later by RT–qPCR. *P < 0.0001. j, k, IMR90 cells were transfected with chromatin fragments, stained for H3, quantified for CCF (j), and analysed by RT–qPCR (k). *P < 0.005, **P < 0.0001, compared with sh-NTC transfected. Bar graphs for a–c, e, j are the average values of four different fields with over 200 cells. Error bars, s.d.; n = 3 unless noted; one-way ANOVA coupled with Tukey’s post hoc test (a–d); unpaired two-tailed Student’s t-test (e, g–k). Scale bars, 10 μm.
Extended Data Figure 5 Characterization of ionizing irradiation in mouse liver.
a, Detection of cGAMP in ionizing irradiation hepatocytes by nano-LC–MS. b, Control or ionizing irradiation hepatocytes of WT mice were isolated and stained as indicated. Representative confocal images are shown. CCF are indicated by arrows. Scale bar, 5 μm. c, Related to Fig. 3a, immunohistochemistry staining in no ionizing irradiation control liver.
Extended Data Figure 6 STING promotes Ras-induced SASP in the liver.
a, Immunohistochemistry of WT liver injected with NRasV12/D38A mutant. b, Hepatocytes of injected WT mice were isolated on day 6 and stained. CCF-positive hepatocytes were quantified. Results are average values of four different fields with over 200 cells; *P < 0.001, compared with control and NRasV12/D38A. c, Liver was analysed on day 6 for p21. n = 4 mice. d, e, SA-β-gal analyses of liver on day 6. n = 3 mice, mean with s.e.m. for e. f, Liver was analysed by immunohistochemistry on day 6 and quantified. n = 8 mice; *P < 0.005, **P < 0.001, ***P < 0.0005. g, Liver tumour stained for NRas. One-way ANOVA coupled with Tukey’s post hoc test (b) and unpaired two-tailed Student’s t-test for all others. Scale bars, 10 μm (b); 100 μm for all others. Error bars, s.e.m.
Extended Data Figure 7 Re-expression of STING in the null liver rescues the SASP.
a, Illustration of constructs used for hydrodynamic injection. b, Liver was harvested on day 6 and analysed by immunoblotting. c, Liver was harvested on day 6 and analysed by RT–qPCR. n = 8 mice. d, Immunohistochemistry analyses of liver. Regions with clusters of immune cells are indicated with red arrows, and a representative region is shown in inset. Scale bar, 100 μm. e, Quantification of immune cell clusters and NRas hepatocytes per field. n = 4 mice, *P < 0.05, **P < 0.0005. Unpaired two-tailed Student’s t-test. Error bars, s.e.m.
Extended Data Figure 8 Cytoplasmic chromatin promotes pro-inflammatory responses in OIS-evaded and cancer cells.
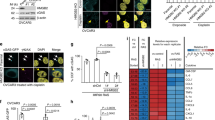
a, OIS-evaded IMR90 cells were analysed by confocal microscopy. b, c, OIS-evaded IMR90 cells were analysed by RT–qPCR. n = 3, *P < 0.05, **P < 0.0001, compared with sh-NTC. d, e, Cancer cells were imaged under confocal microscopy; cytoplasmic chromatin indicated by arrows. f, Cytoplasmic chromatins were quantified and presented as normalized values from four different fields with over 200 cells. *P < 0.05, **P < 0.01, ***P < 0.005, ****P < 0.0001, compared with control. g, The four cell lines were stably infected as indicated, analysed by RT–qPCR, and are presented as a heat map. h, Ten breast cancer cell lines were analysed for cytoplasmic chromatin and pro-inflammatory genes. Cell lines with the lowest and highest 50% of cytoplasmic chromatin were grouped and the cytokine expression levels compared. Error bars, s.e.m. for h and s.d. for all others; one-way ANOVA coupled with Tukey’s post hoc test (c, f); unpaired two-tailed Student’s t-test (h). Scale bars, 10 μm.
Extended Data Figure 9 CCLE analyses of pro-inflammatory gene expression.
a, Related to Fig. 4g, additional genes associated with STING or lamin B1. b, Analyses of cGAS with pro-inflammatory gene expression profiles. Samples with the highest 25% and the lowest 25% of cGAS expression were selected and grouped; the numbers of samples are indicated. c, Lamin A/C does not show negative correlation with inflammatory genes. d, MAVS does not correlate with pro-inflammatory gene expression. Statistical significance was judged by one-sided Wilcoxon rank-sum test. P values are shown for each comparison. NS, non-significant (P > 0.05). See Methods for additional details.
Extended Data Figure 10 STING associates with pro-inflammatory gene expression in human cancers.
Box plots of TCGA RNA expression profiles in pancreatic adenocarcinoma (a), cutaneous melanoma (b), prostate adenocarcinoma (c), and breast adenocarcinoma (d). In each cancer type, samples with the highest 25% and the lowest 25% of STING expression were selected and grouped; the numbers of samples are indicated. Pro-inflammatory gene expression levels were then analysed between STING-high and STING-low groups. Statistical significance was judged by one-sided Wilcoxon rank-sum test. P values are shown for each comparison. NS, non-significant (P > 0.05). See Methods for additional details.
Supplementary information
Supplementary Figure
This file contains the original western blot images. (PDF 750 kb)
Supplementary Table 1
Complete GO analysis results from RNA-seq. Related to Figure 2, differentially expressed genes comparing sh-cGAS vs sh-NTC were subjected to GO analysis. Complete GO terms and genes are listed in the table. (XLSX 51 kb)
Supplementary Table 2
Cancer cell line pro-inflammatory cytokine results. Related to Extended Data Figure 8g, the raw data and p-values for each condition of the four cancer cell lines are included. (XLSX 13 kb)
Supplementary Table 3
Analyses of ten breast cancer cell lines. Related to Extended Data Figure 8h, the raw data of cytoplasmic chromatin and the values of cytokines are included. (XLSX 11 kb)
Supplementary Table 4
TCGA analyses of human cancer transcriptome. Related to Extended Data Figure 10, the expression levels of pro-inflammatory genes and interferon genes were analyzed, together with STING and MAVS, in the four types of human cancers. (XLSX 13 kb)
Rights and permissions
About this article
Cite this article
Dou, Z., Ghosh, K., Vizioli, M. et al. Cytoplasmic chromatin triggers inflammation in senescence and cancer. Nature 550, 402–406 (2017). https://doi.org/10.1038/nature24050
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24050
This article is cited by
-
Dormancy of cutaneous melanoma
Cancer Cell International (2024)
-
The senescence journey in cancer immunoediting
Molecular Cancer (2024)
-
Second messenger 2'3'-cyclic GMP-AMP (2'3'-cGAMP): the cell autonomous and non-autonomous roles in cancer progression
Acta Pharmacologica Sinica (2024)
-
Mitochondrial injury induced by a Salmonella genotoxin triggers the proinflammatory senescence-associated secretory phenotype
Nature Communications (2024)
-
TXNRD1 drives the innate immune response in senescent cells with implications for age-associated inflammation
Nature Aging (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.