Abstract
Type 2 innate lymphoid cells (ILC2s) both contribute to mucosal homeostasis and initiate pathologic inflammation in allergic asthma. However, the signals that direct ILC2s to promote homeostasis versus inflammation are unclear. To identify such molecular cues, we profiled mouse lung-resident ILCs using single-cell RNA sequencing at steady state and after in vivo stimulation with the alarmin cytokines IL-25 and IL-33. ILC2s were transcriptionally heterogeneous after activation, with subpopulations distinguished by expression of proliferative, homeostatic and effector genes. The neuropeptide receptor Nmur1 was preferentially expressed by ILC2s at steady state and after IL-25 stimulation. Neuromedin U (NMU), the ligand of NMUR1, activated ILC2s in vitro, and in vivo co-administration of NMU with IL-25 strongly amplified allergic inflammation. Loss of NMU–NMUR1 signalling reduced ILC2 frequency and effector function, and altered transcriptional programs following allergen challenge in vivo. Thus, NMUR1 signalling promotes inflammatory ILC2 responses, highlighting the importance of neuro-immune crosstalk in allergic inflammation at mucosal surfaces.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Change history
15 November 2017
Please see accompanying Erratum (http://doi.org/10.1038/nature24480). Author Pankaj Baral was listed incorrectly as a corresponding author in the HTML rather than author Patrick R. Burkett. This has been corrected online.
References
Cheng, D. et al. Epithelial interleukin-25 is a key mediator in Th2-high, corticosteroid-responsive asthma. Am. J. Respir. Crit. Care Med. 190, 639–648 (2014)
Huang, Y. et al. IL-25-responsive, lineage-negative KLRG1hi cells are multipotential ‘inflammatory’ type 2 innate lymphoid cells. Nat. Immunol. 16, 161–169 (2015)
Gudbjartsson, D. F. et al. Sequence variants affecting eosinophil numbers associate with asthma and myocardial infarction. Nat. Genet. 41, 342–347 (2009)
Halim, T. Y. et al. Group 2 innate lymphoid cells are critical for the initiation of adaptive T helper 2 cell-mediated allergic lung inflammation. Immunity 40, 425–435 (2014)
Salimi, M. et al. A role for IL-25 and IL-33-driven type-2 innate lymphoid cells in atopic dermatitis. J. Exp. Med. 210, 2939–2950 (2013)
Moro, K. et al. Innate production of T(H)2 cytokines by adipose tissue-associated c-Kit+Sca−1+ lymphoid cells. Nature 463, 540–544 (2010)
Neill, D. R. et al. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464, 1367–1370 (2010)
Chang, Y. J. et al. Innate lymphoid cells mediate influenza-induced airway hyper-reactivity independently of adaptive immunity. Nat. Immunol. 12, 631–638 (2011)
Monticelli, L. A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 12, 1045–1054 (2011)
Tanay, A. & Regev, A. Scaling single-cell genomics from phenomenology to mechanism. Nature 541, 331–338 (2017)
Wagner, A., Regev, A. & Yosef, N. Revealing the vectors of cellular identity with single-cell genomics. Nat. Biotechnol. 34, 1145–1160 (2016)
Gaublomme, J. T. et al. Single-cell genomics unveils critical regulators of Th17 cell pathogenicity. Cell 163, 1400–1412 (2015)
Habib, N. et al. Div-Seq: single-nucleus RNA-seq reveals dynamics of rare adult newborn neurons. Science 353, 925–928 (2016)
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323.e30 (2016)
Gury-BenAri, M. et al. The spectrum and regulatory landscape of intestinal innate lymphoid cells are shaped by the microbiome. Cell 166, 1231–1246.e13 (2016)
Monticelli, L. A. et al. Arginase 1 is an innate lymphoid-cell-intrinsic metabolic checkpoint controlling type 2 inflammation. Nat. Immunol. 17, 656–665 (2016)
Robinette, M. L. et al. Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets. Nat. Immunol. 16, 306–317 (2015)
Björklund, A. K. et al. The heterogeneity of human CD127+ innate lymphoid cells revealed by single-cell RNA sequencing. Nat. Immunol. 17, 451–460 (2016)
Kowalczyk, M. S. et al. Single-cell RNA-seq reveals changes in cell cycle and differentiation programs upon aging of hematopoietic stem cells. Genome Res. 25, 1860–1872 (2015)
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016)
Dixit, A. et al. Perturb-Seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866.e17 (2016)
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017)
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013)
Moriyama, M. et al. The neuropeptide neuromedin U activates eosinophils and is involved in allergen-induced eosinophilia. Am. J. Physiol. Lung Cell. Mol. Physiol. 290, L971–L977 (2006)
Hedrick, J. A. et al. Identification of a human gastrointestinal tract and immune system receptor for the peptide neuromedin U. Mol. Pharmacol. 58, 870–875 (2000)
Szekeres, P. G. et al. Neuromedin U is a potent agonist at the orphan G protein-coupled receptor FM3. J. Biol. Chem. 275, 20247–20250 (2000)
Shan, L. et al. Identification of a novel neuromedin U receptor subtype expressed in the central nervous system. J. Biol. Chem. 275, 39482–39486 (2000)
Talbot, S. et al. Silencing nociceptor neurons reduces allergic airway inflammation. Neuron 87, 341–354 (2015)
Nussbaum, J. C. et al. Type 2 innate lymphoid cells control eosinophil homeostasis. Nature 502, 245–248 (2013)
Audrit, K. J., Delventhal, L., Aydin, Ö. & Nassenstein, C. The nervous system of airways and its remodeling in inflammatory lung diseases. Cell Tissue Res. 367, 571–590 (2017)
Polte, T., Behrendt, A. K. & Hansen, G. Direct evidence for a critical role of CD30 in the development of allergic asthma. J. Allergy Clin. Immunol. 118, 942–948 (2006)
Schuliga, M. et al. Plasminogen-stimulated inflammatory cytokine production by airway smooth muscle cells is regulated by annexin A2. Am. J. Respir. Cell Mol. Biol. 49, 751–758 (2013)
Heshmat, N. M. & El-Hadidi, E. S. Soluble CD30 serum levels in atopic dermatitis and bronchial asthma and its relationship with disease severity in pediatric age. Pediatr. Allergy Immunol. 17, 297–303 (2006)
Katsunuma, T. et al. Analysis of gene expressions of T cells from children with acute exacerbations of asthma. Int. Arch. Allergy Immunol. 134, 29–33 (2004)
Sekigawa, T. et al. Gene-expression profiles in human nasal polyp tissues and identification of genetic susceptibility in aspirin-intolerant asthma. Clin. Exp. Allergy 39, 972–981 (2009)
Kurakula, K. et al. Nuclear receptor Nur77 attenuates airway inflammation in mice by suppressing NF-κB activity in lung epithelial cells. J. Immunol. 195, 1388–1398 (2015)
Modena, B. D. et al. Gene expression correlated with severe asthma characteristics reveals heterogeneous mechanisms of severe disease. Am. J. Respir. Crit. Care Med. 195, 1449–1463 (2017)
Beale, J. et al. Rhinovirus-induced IL-25 in asthma exacerbation drives type 2 immunity and allergic pulmonary inflammation. Sci. Transl. Med. 6, 256ra134 (2014)
von Moltke, J., Ji, M., Liang, H. E. & Locksley, R. M. Tuft-cell-derived IL-25 regulates an intestinal ILC2-epithelial response circuit. Nature 529, 221–225 (2016)
Gerbe, F. et al. Intestinal epithelial tuft cells initiate type 2 mucosal immunity to helminth parasites. Nature 529, 226–230 (2016)
Howitt, M. R. et al. Tuft cells, taste-chemosensory cells, orchestrate parasite type 2 immunity in the gut. Science 351, 1329–1333 (2016)
Gu, X. et al. Chemosensory functions for pulmonary neuroendocrine cells. Am. J. Respir. Cell Mol. Biol. 50, 637–646 (2014)
Branchfield, K. et al. Pulmonary neuroendocrine cells function as airway sensors to control lung immune response. Science 351, 707–710 (2016)
Prendergast, C. E., Morton, M. F., Figueroa, K. W., Wu, X. & Shankley, N. P. Species-dependent smooth muscle contraction to Neuromedin U and determination of the receptor subtypes mediating contraction using NMU1 receptor knockout mice. Br. J. Pharmacol. 147, 886–896 (2006)
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 166, 1500–1511.e9 (2016)
Jäger, A., Dardalhon, V., Sobel, R. A., Bettelli, E. & Kuchroo, V. K. Th1, Th17, and Th9 effector cells induce experimental autoimmune encephalomyelitis with different pathological phenotypes. J. Immunol. 183, 7169–7177 (2009)
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015)
Zeileis, A., Kleiber, C. & Jackman, S. Regression models for count data in R. J. Stat. Softw. 27, (2008)
Jackman, S. pscl: Classes and Methods for R Developed in the Political Science Computational Laboratory, Stanford University. Department of Political Science, Stanford University. Stanford, California. R package version 1.4.9. http://pscl.stanford.edu/
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016)
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000 Res. 4, 1521 (2015)
Acknowledgements
We thank J. Xia, G. Zhu, D. Kozoriz, the Dana Farber Cancer Institute Rodent Histopathology Core, and the Harvard Medical School Transgenic Core for technical expertise. The KOMP repository, CSD Consortium, and Velocigene at Regeneron Inc. generated Nmur1-LacZ mice with the support of the NIH (U01HG004085, U01HG004080). M. Kojima (Kurume University) generated Nmu knockout mice. L. Gaffney assisted with figure preparation. R. Herbst and A. Haber provided statistical advice. A.W. is jointly supervised by V.K.K. and H.-M. Jäck (Friedrich-Alexander University of Erlangen-Nürnberg, Erlangen, Germany) and is supported by a Boehringer Ingelheim Fonds PhD fellowship. P.R.B. (1K08AI123516), R.E.A. (1K08HL130540), B.D.L. (RO1-HL122531) and V.K.K. (PO1 AI056299, AI039671) are supported by the N.I.H. M.S.K. was supported by Charles A. King Trust Postdoctoral Research Fellowship Program and the Simeon J. Fortin Charitable Foundation. C.S.N.K. is supported by the German Research Foundation (DFG; KL 2963/1-1). T.M. is supported by the Crohn’s and Colitis Foundation of America (CCFA). D.A. is supported by the NIH (AI061570, AI087990, AI074878, AI083480, AI095466, AI095608, AI102942 and AI097333), the Burroughs Wellcome Fund, and the CCFA. A.R. is an Investigator of the Howard Hughes Medical Institute. We acknowledge the support of the Food Allergy Scientific Initiative and the Klarman Cell Observatory at the Broad Institute.
Author information
Authors and Affiliations
Contributions
A.W., S.J.R. and P.R.B. contributed equally to this study. P.R.B., A.R. and V.K.K. co-conceived the study. P.R.B., A.W., S.J.R., M.S.K., V.K.K. and A.R. designed the experiments and interpreted results. A.W. and P.R.B. performed and analysed the functional biological experiments, including preparation of cells for scRNA-seq, except immunofluorescence microscopy. S.J.R. designed and performed the computational analysis, with assistance from M.H., M.S.C, B.J.H. and T.L.T. M.S.K. directed scRNA-seq efforts, in conjunction with O.R. M.S.K., J.N., D.D., C.R., D.F. and J.J.T. performed scRNA-seq. R.A.E. and B.D.L. assisted with measuring airway resistance. C.S.N.K, T.M. and D.A. performed and analysed immunofluorescence staining of the lung. P.B. and I.C. assisted with analysis of neurons. The manuscript was written by P.R.B., S.J.R. and A.W., and was edited by A.R. and V.K.K., with input from all the authors.
Corresponding authors
Ethics declarations
Competing interests
A.R. is a member of the scientific advisory board of ThermoFisher, Syros Pharmaceuticals, and Driver Group. All other authors have no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Massively parallel scRNA-seq of lung ILCs.
a, Schematic of experimental method for in vivo activation of ILCs. b, c, Quality measures. b, Scatter plots show for each cell the relation between the number of UMIs (nUMI, x axis) and the number of genes (nGene, y axis). Cells are coloured according to whether they are included (grey) or excluded (red) from further analysis by these measures. c, Box plots show the distribution of the number of unique genes (y axis, left), UMIs (y axis, middle) and per cent of normalized expression from mitochondrial genes (y axis, right) in each treatment and replicate (rep; x axis). White diamond indicates the mean. d–g, ILC classification by signatures. d, Density plots show the distributions of ILC subset signature scores (x axis) for ILC1 (left), ILC2 (middle) and ILC3 (right) in each of three treatments (control, grey; IL-25, yellow; IL-33, blue). Dashed lines mark the cut-offs used in the ILC classification for each signature. e, Bar plots of the proportion of 24,187 cells (y axis) classified in each treatment (x axis) based on transcriptional signatures of known ILC subsets as ILC1 (green), ILC2 (pink), ILC3 (blue), mixed (yellow, scoring for two or more signatures), or none (grey, scoring for none of the three signatures). f, Bar plots show the number of ILCs (y axis) classified to each subset (colour code) in the PBS, IL-25 or IL-33 conditions (x axis). g, Expression of key ILC marker genes. Violin plots show the distribution of expression levels (logTPX, y axis) of each of 12 ILC marker genes (marked on top) in cells classified in each subtype (rows) in each treatment (x axis). h, Cell groupings are independent of batch. tSNE plot of cells (as in Fig. 1a) from replicate 1 (left plot) and replicate 2 (right plot), coloured by condition. i, Expression of key genes. tSNE is coloured by the relative expression of the indicated genes. j, Gene expression for marker genes of ILCs and other immune cell types. For each gene (columns) in each cluster (rows), the proportion of cells in the cluster expressing the gene (dot size) and the relative mean log nUMI of expressing cells (colour) is plotted. k, l, Cluster composition. Bar plots show the number of cells (y axis) from each ILC type (k) and condition (l) in each cluster (x axis). m, Validation of expression patterns identified by scRNA-seq using flow cytometry. In each histogram, the expression of proteins encoded by genes identified with scRNA-seq on ILCs from mice treated with PBS (grey, closed) versus one of the alarmins IL-25 or IL-33 (blue, closed) is shown, as well as a fluorescence minus one (FMO) control (blue, dashed). Numbers indicate the mean fluorescence intensity of the marker in PBS (grey) or condition (blue) or, for CTLA4 and MHC class II, the frequency of cells positive for the marker in PBS (grey) or condition (blue). Data are representative of at least two individual experiments.
Extended Data Figure 2 Alarmin-induced ILC proliferation in vitro.
ILCs were labelled with CellTrace Violet and cultured in vitro under the indicated conditions. After 3 days, proliferation was analysed by flow cytometric analysis of CellTrace Violet dilution. The frequency of non-proliferating ILCs is indicated. Data are representative of two individual experiments.
Extended Data Figure 3 Plate-based scRNA-seq of lung-resident ILCs.
a, Quality measures. Scatter plots for each plate showing the total number of read counts per cell (x axis) and number of unique detected genes (y axis) in each cell. Dashed lines indicate cut-offs for passing quality control. Only cells in the central quadrant were retained for analysis. b–e, ILC subset classification. b, ILC signature score distributions. Density plots show the distributions of ILC subset signature scores (x axis) for ILC1 (top), ILC2 (middle) and ILC3 (bottom) in each of three treatments (control, grey; IL-25, yellow; IL-33, blue). Dashed lines mark the minimum score cut-off used in the ILC classification for each signature. c, Expression of key ILC marker genes. Violin plots show the distribution of expression levels (logTPM, y axis) of each of 12 ILC marker genes (marked on top) in cells classified in each subtype (rows) in each treatment (x axis). d, Distinct cell subsets by ILC expression profile. tSNE plot shows individual cells (dots) in a nonlinear reduced representation of the top 9 PCs, with cells coloured by ILC subset. e, Composition by treatment condition. Bar plots show the proportion of each ILC subtype (y axis) in each treatment condition (x axis).
Extended Data Figure 4 Accurate detection of Nmur1 from massively parallel 3′end scRNA-seq requires cell-type specific annotation.
a, Comparison of gene expression estimates from plate-based and droplet-based scRNA-seq. Scatter plots show for each gene (dot) the fraction of cells expressing it according to plate-based scRNA-seq (x axis) and droplet-based scRNA-seq (y axis) in the PBS (top) and IL-25 (bottom) conditions. Of the differentially expressed genes in the plate-based data that are expressed in a substantially different proportion of cells compared to droplet-based data (red), Nmur1 and Rapgef4 are among the highest ranked and are only detected in the plate-based data. b, Re-annotation of the Nmur1 locus by RNA-seq and assembly. Shown is a window of approximately 7 kb around the annotated Nmur1 locus on chromosome 1 (top) along with read alignments to that region from either droplet-based 3′scRNA-seq (top, 55-nt reads) or from population (bulk) RNA-seq of ILCs (bottom, 75-nt reads). The Refseq annotation of Nmur1 (top blue track) does not extend to the 3′ end of the transcript, as defined by either the scRNA-seq reads (10× Nmur1 extended; middle blue track) or by a transcriptome reassembly from bulk RNA-seq (StringTie reassembled; bottom blue track). c, Corrected annotation recovers Nmur1. Histogram shows the distribution of expression levels of Nmur1 in single cells based on droplet-based scRNA-seq data when expression was calculated by aligning reads with the original RefSeq annotation (left), the scRNA-seq read-based extended annotation (middle) or the reassembled transcript annotation from bulk RNA-seq (right). d, Rapgef4 locus correctly annotated. Shown is a plot for Rapgef4, arranged as in b. The 3′ annotation in the RefSeq annotation agrees with the observed end from bulk RNA-seq.
Extended Data Figure 5 Expression of Nmur1 and Nmur2 in lung-resident immune cells.
a, Minimal expression of Nmur1 on in vitro differentiated TH2 cells. Bar chart shows Nmur1 mRNA levels by qPCR in both TH2 cells differentiated under the indicated conditions in vitro, and in freshly isolated ILCs. Data points represent technical replicates (n = 2). b, Nmur2 is not expressed by lung-resident immune cells, including ILCs. Bar chart shows Nmur2 mRNA levels by qPCR for the indicated cell types, which were sorted from mice either at steady state (PBS) or after HDM challenge. The mean of technical replicates is shown (n = 2). c, Summary graph of the flow cytometry data, of which a representative is shown in Fig. 2e. Plot shows the frequency of NMUR1+ cells in different cell populations from Nmur1-LacZ reporter mice as assayed by flow cytometry. Data points represent individual mice (n = 3). d, Nmur1-LacZ β-galactosidase activity reveals ILC-specific NMUR1 expression. Representative histograms show the distribution of expression of NMUR1 in the indicated cell types as determined by flow cytometry in cells isolated from wild-type (red) and Nmur1-LacZ heterozygous (blue) mice. The frequency of Nmur1-LacZ-positive cells is indicated. e, NMUR1 is preferentially expressed by ST2+ ILCs. The expression of ST2, IL-17RB and NMUR1 was analysed by flow cytometry, and the frequency of NMUR1+ ILCs within the indicated populations is shown. Each data point reflects an individual mouse (n = 4 for PBS and IL-33; n = 3 for IL-25). All panels represent one of two individual experiments, mean is indicated, error bars represent s.e.m.; *P < 0.05 by two-tailed t-test.
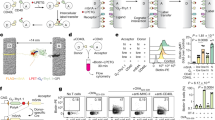
Extended Data Figure 6 ILCs and neurons are in proximity in the lung, and NMU is expressed by dorsal root ganglion neurons.
a, Representative image (left) of lung sections stained for the neuronal marker SNAP-25 (green), KLRG1 (red), and CD3ε (blue). Arrows indicate KLRG1+ CD3ε– cells in close proximity to SNAP-25+ nerve fibres. Scale bar, 100 μm. Bar plot (right) shows the frequency of ILCs (x axis) within indicated distances (y axis) of SNAP-25+ nerve fibres. All KLRG1+CD3ε− cells in the field of view were counted, and the distances between ILCs and the closest SNAP-25+ nerve fibre were measured (3–5 lung sections per mouse, n = 3 mice). Data are representative of three independent experiments with similar results. b, c, NMU is expressed in thoracic dorsal root ganglion (DRG) neurons. NMU expression was examined by qPCR. b, NMU expression in the nodose/jugular ganglion and DRG ex vivo. Data points represent technical replicates (n = 2). c, NMU expression in DRG neurons stimulated in vitro with IL-13. Data points represent two technical replicates from each of six biologic replicates. All panels are representative of at least two individual experiments, mean is indicated, error bars represent s.e.m.; *P < 0.05 by two-tailed t-test.
Extended Data Figure 7 NMU induces minimal lung inflammation in the absence of alarmins.
a, NMU does not alter in vitro expression of type 2 cytokines in T cells. Expression of Il5 and Il13 is similar in TH2 cells cultured with or without NMU, as assessed by qPCR. b, c, ILCs were isolated from naive mice and cultured in vitro with the indicated stimuli. b, ILCs cultured in vitro have markedly enhanced Il5 and Il13 mRNA expression in response to combination of NMU and IL-25, as measured by qPCR. c, IL-33-induced IL-5 and IL-13 secretion is enhanced by NMU, as determined by LegendPlex. Data points represent technical replicates, and the mean is indicated. d–h, NMU alone induces minimal allergic lung inflammation. d, Schematic of experimental method for activation of ILCs with NMU in vivo. e, Il5 and Il13 expression was determined in lung mononuclear cells by qPCR. f, The concentration of IL-5 and IL-13 protein in BALF was analysed by LegendPlex. g, h, Eosinophil frequencies in lung parenchyma (g) and BALF (h, left), and BALF eosinophil numbers (h, right), as determined by flow cytometry. Data points correspond to individual mice (n = 3). i, The combination of IL-25 combined with NMU markedly enhanced Il5 (top) and Il13 (bottom) mRNA expression in lung mononuclear cells, as measured by qPCR. j, NMU and IL-25 synergize to promote tissue eosinophilia. Eosinophil frequency (top) and number (bottom) in lung parenchyma was assessed by flow cytometry. Data points correspond to individual mice (PBS, n = 3; IL-25, n = 9; IL-25 combined with NMU, n = 7). For all panels, data are representative of at least two individual experiments, mean is indicated, error bars represent s.e.m.; *P < 0.05 by two-tailed t-test.
Extended Data Figure 8 Phenotype of Nmu-knockout mice.
a–c, ILC frequency and function are not altered in the absence of NMU in steady state. Lung-resident ILCs isolated from wild-type and NMU-deficient mice were analysed by flow cytometry. a, b, The frequency of ILCs (among CD90+ cells, a) and expression of KLRG1 and ST2 on ILCs (b) are unchanged in Nmu-knockout mice compared to control mice. c, Frequency of IL-5- and IL-13-producing ILCs in Nmu-knockout mice is comparable to that of control mice. Each data point corresponds to an individual mouse (wild-type, n = 4; Nmu-knockout, n = 5). d–f, Analysis of BALF and T-cell responses in Nmu-knockout mice after HDM challenge. d, BALF cell numbers (left) and the frequency of eosinophils (right) were analysed after HDM challenge in Nmu-knockout mice. Total cell numbers (wild-type, n = 15; Nmu-knockout, n = 13) and eosinophil frequency are unchanged in Nmu-knockout mice (n = 17) compared to wild-type mice (n = 21). e, Increased lung-infiltrating CD4 T cells in the absence of NMU. The frequency of lung-infiltrating T cells was determined by flow cytometry after HDM challenge (wild-type, n = 15; Nmu-knockout, n = 13). f, Intact TH2 response in the absence of NMU. Expression of IL-5 and IL-13 by CD4 T cells was analysed by flow cytometry following HDM challenge. Each data point represents an individual mouse (wild-type, n = 5; Nmu-knockout, n = 5). All panels are representative of at least two individual experiments. For all panels, mean is indicated, error bars represent s.e.m.; *P < 0.05 by two-tailed t-test (a–d, f) or one-way ANOVA (e).
Extended Data Figure 9 Massively parallel, droplet-based scRNA-seq of ILCs from mice treated with NMU or IL-25 combined with NMU.
a, Quality measures. Bar plots show the distribution of the number of unique genes (y axis, top), per cent of reads from mitochondrial genes (y axis, middle) and number of UMIs (y axis, bottom) in each treatment and replicate (x axis). White diamond indicates the mean. b, Grouping of ILCs is independent of batch. tSNE plots (as in Fig. 4a) of cells from replicate 1 (top) and replicate 2 (bottom) coloured by condition. c, Faceted tSNE plot of ILCs from all alarmin and NMU treatments, coloured by condition, shows that IL-25 combined with NMU and IL-33 induce partially overlapping transcriptional responses. d, e, Cluster composition. d, Bar plot shows the proportion of ILCs from each condition in each cluster. e, Bar plot shows the number of cells from each condition in each cluster. f, g, IL-25 combined with NMU promotes ILC proliferation. f, The frequency of Ki-67+ ILCs was determined by flow cytometry in mice treated with IL-25 or IL-25 combined with NMU. Data points represent individual mice (IL-25, n = 8; IL-25 combined with NMU, n = 6). g, Violin plots show the distributions of the proliferation signature scores (y axis) for the cells in each cluster (x axis) (white diamond, mean; lines, first and third quartiles). Proliferation scores in cluster 11 are significantly higher than those in the other clusters dominated by IL-25 combined with NMU (8, 9 and 10) (t-test, P < 2.2 × 10−16). h, IL-25 combined with NMU enhances expression of Il17rb in mononuclear lung cells, as shown by qPCR. Data points represent individual mice (PBS, n = 3; IL-25, n = 9; IL-25 combined with NMU, n = 7). i, Validation of expression patterns identified by scRNA-seq using flow cytometry. Expression levels for proteins encoded by genes identified by scRNA-seq on ILCs from mice treated with PBS (grey, closed histogram) versus one of the treatments (blue, closed histogram), as well as an FMO control (dashed open histogram) is shown. Numbers indicate the mean fluorescence intensity of the respective marker or, if a gate is indicated, the frequency of positive ILCs in PBS (grey) and in the indicated condition (blue). j, Distinct patterns of expression of key differentially expressed genes across clusters. Violin plots show the expression levels in logTPX (y axis) for the indicated genes across the cells in each cluster (x axis) (white diamond, mean; line, median). k, NMUR1 is downregulated after activation with IL-25 combined with NMU. The frequency of NMUR1+ ILCs (y axis) in each condition among ILCs isolated from mice treated with the indicated stimuli is shown. Dots represent individual mice (n = 4, except for IL-25, where n = 3). l, m, Increased frequency of KLRG1hiST2− inflammatory ILC2s (iILC2s) after treatment with IL-25 combined with NMU. l, Frequency of iILC2s (of total ILCs) isolated from mice after different treatments. Dots represent individual mice (n = 4, except for IL-25, where n = 3). m, Representative flow cytometry plots of ST2 and KLRG1 expression on ILCs for the indicated conditions. For panels f, h k, l, mean is indicated and error bars represent s.e.m.; *P < 0.05 by two-tailed t-test (f) or one-way ANOVA (h, k, l).
Extended Data Figure 10 Transcriptional analysis by massively parallel scRNA-seq of ILCs from wild-type and Nmur1-knockout mice during airway inflammation.
a, Quality measures. The top row shows the distributions of the number of genes (log scale, x axis, left histogram) and UMIs (log scale, x axis, centre histogram) per cell and their relation (scatter plot, right) in the aggregated data. The middle row shows the distributions of the estimated saturation for UMIs (x axis, left histogram) and genes (x axis, middle histogram) per cell, as well as the ratio of the number of unique UMIs to total number of UMIs per cell (x axis, right histogram), for a representative sample. The bottom row shows the distribution of the relative difference of the total number of UMIs and number of unique UMIs (x axis, left histogram), and the relation between the number of UMIs and number of genes (scatter plot, right), for the same representative sample. Dashed lines show lower and upper cut-offs used. Outlier cells (red dots, scatter plot) were also removed. b, Minor batch effects. tSNE plots show cells from replicate 1 (left) and replicate 2 (right) coloured by condition and genotype (PBS-treated wild-type mice (light grey), HDM-treated wild-type mice (red), PBS-treated Nmur1-knockout mice (dark grey), HDM-treated Nmur1-knockout mice (salmon)). c, d, Signature-based scoring of ILCs. Density plots show the distribution of scores for ILC1, 2 and 3 signatures in the cells from indicated treatment conditions. Dashed line indicates the cut-off for each signature used in assigning ILC type. e, Composition by ILC type. Bar plot shows proportion of each ILC type in the conditions indicated. f, ILC proliferation is strongly increased by IL-25 combined with NMU. Violin plots show the distribution of proliferation scores (y axis) across the cells in indicated conditions (x axis) (white diamond, mean; lines, first and third quartiles). g, Similar clusters are induced by HDM and by IL-25 combined with NMU treatment. tSNE plot shows ILCs from PBS- and HDM-treated wild-type and Nmur1-knockout mice (as in Fig. 5a), coloured by cluster and labelled by post hoc annotations. Note similarity to clustering in Fig. 4b. h, Cluster-specific ILC proliferation in cells from HDM-treated mice. Violin plots show the distribution of proliferation scores (y axis) across the cells in each cluster (x axis), as in g.
Supplementary information
Supplementary Table 1
Differential expression. This file contains lists of differentially expressed genes for dataset groups A, B, C, and plate-based (defined in Methods) and the genes defining the inflammatory ILC2 signature. Each list is found in a separate tab. The condition/cluster in which the gene is differentially expressed and direction of expression (Methods) are indicated. (XLSX 50 kb)
Supplementary Table 2
ILC type and proliferation signatures. This table contains lists of genes defining the ILC subset signatures and the cell proliferation signature. Each list is found in a separate tab. (XLSX 39 kb)
Rights and permissions
About this article
Cite this article
Wallrapp, A., Riesenfeld, S., Burkett, P. et al. The neuropeptide NMU amplifies ILC2-driven allergic lung inflammation. Nature 549, 351–356 (2017). https://doi.org/10.1038/nature24029
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24029
This article is cited by
-
Cooperation of ILC2s and TH2 cells in the expulsion of intestinal helminth parasites
Nature Reviews Immunology (2024)
-
Physiological and immunological barriers in the lung
Seminars in Immunopathology (2024)
-
Group 2 innate lymphoid cells and their surrounding environment
Inflammation and Regeneration (2023)
-
Nr4a1 marks a distinctive ILC2 activation subset in the mouse inflammatory lung
BMC Biology (2023)
-
HTR2A agonists play a therapeutic role by restricting ILC2 activation in papain-induced lung inflammation
Cellular & Molecular Immunology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.