Abstract
The ubiquitin system regulates essential cellular processes in eukaryotes. Ubiquitin is ligated to substrate proteins as monomers or chains and the topology of ubiquitin modifications regulates substrate interactions with specific proteins. Thus ubiquitination directs a variety of substrate fates including proteasomal degradation1. Deubiquitinase enzymes cleave ubiquitin from substrates and are implicated in disease2; for example, ubiquitin-specific protease-7 (USP7) regulates stability of the p53 tumour suppressor and other proteins critical for tumour cell survival3. However, developing selective deubiquitinase inhibitors has been challenging4 and no co-crystal structures have been solved with small-molecule inhibitors. Here, using nuclear magnetic resonance-based screening and structure-based design, we describe the development of selective USP7 inhibitors GNE-6640 and GNE-6776. These compounds induce tumour cell death and enhance cytotoxicity with chemotherapeutic agents and targeted compounds, including PIM kinase inhibitors. Structural studies reveal that GNE-6640 and GNE-6776 non-covalently target USP7 12 Å distant from the catalytic cysteine. The compounds attenuate ubiquitin binding and thus inhibit USP7 deubiquitinase activity. GNE-6640 and GNE-6776 interact with acidic residues that mediate hydrogen-bond interactions with the ubiquitin Lys48 side chain5, suggesting that USP7 preferentially interacts with and cleaves ubiquitin moieties that have free Lys48 side chains. We investigated this idea by engineering di-ubiquitin chains containing differential proximal and distal isotopic labels and measuring USP7 binding by nuclear magnetic resonance. This preferential binding protracted the depolymerization kinetics of Lys48-linked ubiquitin chains relative to Lys63-linked chains. In summary, engineering compounds that inhibit USP7 activity by attenuating ubiquitin binding suggests opportunities for developing other deubiquitinase inhibitors and may be a strategy more broadly applicable to inhibiting proteins that require ubiquitin binding for full functional activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012)
Clague, M. J., Heride, C. & Urbé, S. The demographics of the ubiquitin system. Trends Cell Biol. 25, 417–426 (2015)
Nicholson, B. & Suresh Kumar, K. G. The multifaceted roles of USP7: new therapeutic opportunities. Cell Biochem. Biophys. 60, 61–68 (2011)
Ritorto, M. S. et al. Screening of DUB activity and specificity by MALDI-TOF mass spectrometry. Nat. Commun. 5, 4763 (2014)
Hu, M. et al. Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111, 1041–1054 (2002)
Cummins, J. M. et al. Tumour suppression: disruption of HAUSP gene stabilizes p53. Nature 428, 486–487 (2004)
Kon, N . et al. Inactivation of HAUSP in vivo modulates p53 function. Oncogene 29, 1270–1279 (2010)
Pozhidaeva, A. K. et al. Structural characterization of interaction between human ubiquitin-specific protease 7 and immediate-early protein ICP0 of herpes simplex virus-1. J. Biol. Chem. 290, 22907–22918 (2015)
Cheng, J. et al. Molecular mechanism for USP7-mediated DNMT1 stabilization by acetylation. Nat. Commun. 6, 7023 (2015)
Wang, L. et al. Ubiquitin-specific protease-7 inhibition impairs Tip60-dependent Foxp3+ T-regulatory cell function and promotes antitumor immunity. EBioMedicine 13, 99–112 (2016)
Hao, Y. H. et al. USP7 acts as a molecular rheostat to promote WASH-dependent endosomal protein recycling and is mutated in a human neurodevelopmental disorder. Mol. Cell 59, 956–969 (2015)
Clague, M. J. et al. Deubiquitylases from genes to organism. Physiol. Rev. 93, 1289–1315 (2013)
Deshaies, R. J. Proteotoxic crisis, the ubiquitin-proteasome system, and cancer therapy. BMC Biol. 12, 94 (2014)
Heideker, J. & Wertz, I. E. DUBs, the regulation of cell identity and disease. Biochem. J. 467, 191 (2015)
Fåhraeus, R. & Olivares-Illana, V. MDM2’s social network. Oncogene 33, 4365–4376 (2014)
Williams, A. B. & Schumacher, B. p53 in the DNA-damage-repair process. Cold Spring Harb. Perspect. Med. 6, a026070 (2016)
Nawijn, M. C., Alendar, A. & Berns, A. For better or for worse: the role of Pim oncogenes in tumorigenesis. Nat. Rev. Cancer 11, 23–34 (2011)
Hodges, A. J. M., Matteucci, M., Sharpe, A., Sun, M., Wang, X. & Tsui, V. H. Pyrazol-4-yl-heterocyclyl-carboxamide compounds as Pim kinase inhibitors and their preparation. US patent 8, 614,206 B2 (2013)
Geurink, P. P., El Oualid, F., Jonker, A., Hameed, D. S. & Ovaa, H. A general chemical ligation approach towards isopeptide-linked ubiquitin and ubiquitin-like assay reagents. ChemBioChem 13, 293–297 (2012)
Faesen, A. C. et al. The differential modulation of USP activity by internal regulatory domains, interactors and eight ubiquitin chain types. Chem. Biol. 18, 1550–1561 (2011)
Huether, R. et al. The landscape of somatic mutations in epigenetic regulators across 1,000 paediatric cancer genomes. Nat. Commun. 5, 3630 (2014)
Schaefer, J. B. & Morgan, D. O. Protein-linked ubiquitin chain structure restricts activity of deubiquitinating enzymes. J. Biol. Chem. 286, 45186–45196 (2011)
Husnjak, K. & Dikic, I. Ubiquitin-binding proteins: decoders of ubiquitin-mediated cellular functions. Annu. Rev. Biochem. 81, 291–322 (2012)
Yu, M. et al. A resource for cell line authentication, annotation and quality control. Nature 520, 307–311 (2015)
Wertz, I. E. et al. Phosphorylation and linear ubiquitin direct A20 inhibition of inflammation. Nature 528, 370–375 (2015)
Haverty, P. M. et al. Reproducible pharmacogenomic profiling of cancer cell line panels. Nature 533, 333–337 (2016)
Chang, M. T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016)
Lehár, J. et al. Synergistic drug combinations tend to improve therapeutically relevant selectivity. Nat. Biotechnol. 27, 659–666 (2009)
Bueno, R. et al. Comprehensive genomic analysis of malignant pleural mesothelioma identifies recurrent mutations, gene fusions and splicing alterations. Nat. Genet. 48, 407–416 (2016)
Reverdy, C. et al. Discovery of specific inhibitors of human USP7/HAUSP deubiquitinating enzyme. Chem. Biol. 19, 467–477 (2012)
Acknowledgements
We thank W. Fairbrother, L. Frick, S. Fong, E. Helgason, T. Hunsaker, C. Lam, B. Liederer, M. Merchant, J. Nonomiya, J. Peng, T. Pham, L. Rangell, R. Rodriguez, U. Segal, R. Tong, L. Wang, R. Gennis, T. Iwasaki, the Genentech Protein Expression, Cell Central, gCSI, and Sequencing groups, and the Boston Biochem team for reagents and collaborations. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under contract number DE-AC02-76SF00515. The SSRl Structural Molecular Biology Program was supported by the DOE Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences (including P41GM103393). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of National Institute of General Medical Sciences or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
L.K., E.L., J.-P.U., S.P., F.P., S.E.M., and I.E.W. performed cell viability and signalling studies, protein turnover and ubiquitination experiments, and independently replicated cellular data. P.D.L. and T.M. performed NMR studies; L.R. and J.M. performed X-ray crystallography and cross-validated data. J.D., T.K., T.W.B., M.C.M.K., Z.S., B.B., Y.T., M.R.A., J.A.E., C.S., and R.A.B. performed in vitro and cellular biochemical studies and independently replicated data. J.H., M.S.R., T.P.M., D.R.A., M.T., and K.Y. performed deubiquitinase selectivity profiling studies, and M.M. and E.B. performed in vivo studies and cross-validated data. K.R.C. and M.H.B. developed assays, and performed and analysed high-throughput screening studies. R.P., P.J., B.R.H., A.R.R., F.C., C.N., X.W., and V.T. performed compound design/synthesis. F.G., M.T.C., C.K., and W.F.F. performed bioinformatics/statistical analysis and replicated data. T.M. and I.E.W. coordinated studies. J.M., T.M., and I.E.W. wrote the manuscript. D.R.A. is supported by the UK Medical Research Council (grant MC_UU_12016/2).
Corresponding authors
Ethics declarations
Competing interests
L.K., P.D.L., L.R., R.P., K.R.C., J.D., T.K., E.L., J.-P.U., S.P., J.H., M.M., T.W.B., M.C.M.K., T.P.M., F.C., K.Y., F.P., F.G., M.T.C., C.K., E.B., S.E.M., W.F.F., J.A.E., C.N., X.W., M.H.B., V.T., R.A.B., J.M., T.M., and I.E.W. are or were Genentech employees. M.S.R. is a Pfizer employee.
Additional information
Reviewer Information Nature thanks M. Rolfe, S. Scherer and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Screening cascades and hit-to-lead assay data for USP7 inhibitors.
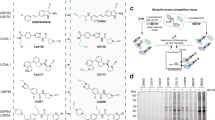
a, High-throughput activity-based screening cascade to identify USP7 inhibitors. Screening stages are identified in bold print. Numbers of compounds at each stage are provided to the right of each box. Criteria for progression to the next stage are highlighted in italics to the left of each arrow. b, Fragment NMR screen diagram. Screening stages are identified in bold print. Numbers of compounds at each stage are provided to the right of each box. Criteria for progression to the next stage are highlighted in italics to the left of each arrow. Protein saturation transfer difference (STD) experiments were performed at 284 K. Primary USP7 catalytic domain binders were selected on the basis of the signal-to-noise ratio of the respective compound with a cut-off of greater than 5. All primary binders were re-measured as single compounds under otherwise identical conditions and confirmed binders selected having a signal-to-noise ratio of greater than 10. Hits were further tested for specific binding to USP7 catalytic domain by measurement of 1H/15N TROSY spectra. Positive hits were defined as compounds that induced chemical shift perturbations. Perturbations were classified by the chemical shift patterns and selected compounds passed onto X-ray soaking experiments. c, Table summarizing the hit-to-lead assay results of the lead compounds identified by the high-throughput screening and NMR fragment screening campaigns. Compound series are grouped in columns and hit-to-lead selection assay data are listed in the indicated rows. At least two experimental replicates were performed; averages are shown ± s.e.m. See Fig. 1b and text for more details. d, Endogenous MDM2 immunofluorescence studies. HCT-116 human colorectal carcinoma cells were treated with a range of concentrations of the indicated USP7 inhibitors or DMSO vehicle for 24 h and endogenous MDM2 protein levels were detected by immunofluorescence imaging. Representative images show cells treated with 10 μM of the indicated compounds or DMSO vehicle control. Scale bar, 20 μm. The graph shows data from experimental duplicates of the quantified mean nuclear MDM2 protein levels per cell over a range of concentrations of GNE-8735 and GNE-2916. The half-maximal effective concentration (EC50) for the elevation in MDM2 caused by GNE-8735 was 2.9 μM. Image source data: Supplementary Fig. 1. e, Quantification of total- and ubiquitinated-MDM2 (Ub-MDM2) in USP7 inhibitor-treated cells. SJSA-1 human osteosarcoma cells were treated with a range of concentrations of the indicated USP7 inhibitors or DMSO vehicle control for 24 h and the level of ubiquitinated-MDM2 and total MDM2 were measured using a multiplexed mesoscale immunoassay. Graphs show experimental duplicates of the quantified level of either total MDM2 (right column, red), ubiquitinated MDM2 (central column, blue), or the percentage change in the ratio of the ubiquitinated-MDM2 signal and the total MDM2 signal (left column, orange). All data are shown as percentage change in each value, relative to DMSO vehicle-treated samples. The maximal extent of the increase in the ubiquitinated-MDM2 to total MDM2 ratio varied between compounds; therefore to compare the potency of the increase in the ratio of ubiquitinated-MDM2 to total MDM2 between compounds, the top level was universally set to 100%. This normalization allowed calculation of the EC50 of the percentage change in this ratio relative to DMSO (left column). Indole tricyclic compounds including GNE-8735 increased total MDM2 levels and inhibited cathepsin-B, indicating poor selectivity and induction of general cell stress by this chemical series (d, e). Indole tricyclics also precipitated caspase-3, although they passed dynamic light scattering analysis (see also Extended Data Fig. 2a). The peptidomimetic compounds had weaker biochemical potency, poor selectivity, and covalently modified USP7 cysteine (Cys) residues other than the catalytic Cys (data not shown). Given these data, and because optimization of indole tricyclic and peptidomimetic compounds proved challenging, these series were discontinued. The tetrahydroacridine and fragment compounds were relatively potent, selective, and enhanced cellular MDM2 ubiquitination without significantly increasing total MDM2 (d, e). Tetrahydroacridine compounds passed cathepsin-B inhibition assays, demonstrating additional protease selectivity. Neither tetrahydroacridine nor fragment compounds showed evidence of USP7 aggregation in dynamic light scattering or in NMR studies (data not shown; see Extended Data Fig. 2a). Tetrahydroacridine compounds, including GNE-6831, covalently modified USP7, consistent with a previous report describing a similar series30 (see also Extended Data Fig. 2b).
Extended Data Figure 2 Biophysical and cellular characterization of selected USP7 inhibitors.
a, The dynamic light scattering autocorrelation functions are shown for 100 μM rottlerin (red), full-length USP7 with 100 μM GNE-8735 (blue), full-length USP7 with 100 μM GNE-2090 (brown), and full-length USP7 with 0.1% DMSO vehicle control (black). The percentage aggregate by mass is shown in the legend. Two experimental replicates were performed. b, USP7 full-length protein was incubated overnight with excess of GNE-6831 and analysed by liquid chromatography–mass spectrometry. Unmodified and covalently modified USP7 are represented in the top and bottom panels, respectively. Three experimental replicates were performed. c, Table of active and inactive tetrahydroacridine compounds with structures. Hit-to-lead selection assay data are listed in the indicated rows. Averages are shown ± s.e.m., where applicable, with the number of biologically independent experiments indicated below. n = 1 experimental replicate, if not specified. See Fig. 1b and text for more details. d, Table of active and inactive fragment compounds with structures. Hit-to-lead selection assay data are listed in the indicated rows. Averages are shown ± s.e.m., where applicable, with the number of biologically independent experiments indicated below. n = 1 experimental replicate, if not specified. See Fig. 1b and text for more details. e, Cell viability assays in AMO-1 cells treated as indicated with the tetrahydroacridine compounds (top graph: purple lines, inactive controls; green line, active compound) and fragment compounds (bottom graph: red lines, inactive controls; blue line, active compound). Experimental triplicate data normalized to vehicle control are plotted as a function of compound concentration. f, Cell viability assays in KMS12-PE cells treated as indicated with the tetrahydroacridine compounds (top graph: purple line, inactive control; green lines, active compounds) and fragment compounds (bottom graph: red line, inactive control; blue line, active compound). Experimental triplicate data normalized to vehicle control are plotted as a function of compound concentration. Tetrahydroacridine compounds GNE-6831 and GNE-2090 decreased viability of KMS12-PE and AMO1 multiple myeloma cell lines but this activity was not differentiated from control compounds GNE-0956, GNE-2143, and GNE-2148. By contrast, the fragment compound GNE-2916 decreased multiple myeloma cell viability significantly more than control compounds GNE-2917, GNE-2931, and GNE-9603. Thus work on tetrahydroacridine series was discontinued and the fragment series was further optimized.
Extended Data Figure 3 Deubiquitinase inhibition and cellular activity of optimized fragment compounds.
a, Table summarizing deubiquitinase biochemical assay data and ubiquitin–MDM2 assay data from optimized fragment compounds and inactive controls. Averages are shown ± s.e.m, where applicable, with the number of biologically independent experiments indicated below. n = 1 experimental replicate, if not specified. Fragment compound structures are shown to the right. b, Representative western blot analysis of USP7, p53, and p21 levels from the cycloheximide (CHX)-chase study of MDM2 turnover shown in Fig. 1c, performed in experimental triplicate. c. Analysis of endogenous MDM2 polyubiquitinated with K48-linked chains. Top: denatured lysates from MCF7 cells treated for 8 h with the indicated compounds were immunoprecipitated with a K48 polyubiquitin linkage-specific antibody and immunocomplexes were western blotted with an anti-MDM2 antibody. Western blot analysis of whole-cell lysates for the indicated proteins are shown below. Representative of two experimental replicates. d, Cell viability of wild-type and USP7-null HCT116 colon adenocarcinoma cells, treated as indicated, and analysed with a CellTiter-Glo assay. Experimental triplicates normalized to vehicle control are plotted as a function of compound concentration. Top: averages ± s.e.m. Two-sided t-tests were used to calculate P values between wild-type HCT-116 and USP7-null cells treated with GNE-6640: 7.5 μM P = 0.01, 10 μM P = 0.041, 12.5 μM P = 0.009, 15 μM P = 0.011, 20 μM P = 0.017. Bottom: experimental triplicate data are shown. e. Cell viability of wild-type and USP7-null HCT116 colon adenocarcinoma cells, treated and analysed as in d with the indicated doses of GNE-6776. Top: averages are shown ± s.e.m. Two-sided t-tests were used to calculate P values between wild-type HCT-116 and USP7-null cells: 1 μM P = 0.023, 2.5 μM P = 0.003, 5 μM P = 0.001, 7.5 μM P = 0.003, 10 μM P = 0.007, 12.5 μM P = 0.001, 15 μM P = 0.007, 17.5 μM P = 0.001, 20 μM P = 0.008. Bottom: experimental triplicate data are shown. Gel source data: Supplementary Fig. 1.
Extended Data Figure 4 Selectivity of USP7 inhibitors, in vitro and in vivo drug metabolism and pharmacokinetic (DMPK) profiling, and xenograft growth inhibition studies with GNE-6776.
a, GNE-6776 dose-dependent cleavage inhibition by USP7 of the indicated di-ubiquitin chains as measured by MALDI–TOF. Data from experimental triplicates are shown. b, Percentage inhibition of the indicated deubiquitinases for cleaving di-ubiquitins after treatment with 100 μM of the indicated USP7 inhibitor compounds, representative of one experimental duplicate. Deubiquitinase concentrations and di-ubiquitin substrates are as in Fig. 2a. c. Supporting western blots for Fig. 2b, left. HEK293T cell lysates were treated with the indicated USP7 inhibitors (0.1% DMSO control = 0 μM compound) and endogenous deubiquitinases were reacted with the HA–ubiquitin–vinylsulfone activity-based probe (HA–Ub–VS). Reacted cell lysates were immunoblotted with the indicated antibodies. *Unreacted deubiquitinases; **probe-reacted deubiquitinases; arrowhead points to a band identified by anti-HA immunoblotting that runs at the expected molecular mass of USP7 and is diminished in lysates treated with GNE-6640. Representative of one experimental duplicate. d, Supporting western blots for Fig. 2b, right. HEK293T cell lysates were treated with the indicated USP7 inhibitors, endogenous deubiquitinases were reacted with the HA–Ub–VS activity-based probe, and reacted cell lysates were immunoblotted with the indicated antibodies. *Unreacted deubiquitinases; **probe-reacted deubiquitinases. Representative of one experimental duplicate. e, In vitro pharmacokinetic assessment of USP7 inhibitors. Calculated drug properties are indicated: molecular mass (MW), lipophilicity at pH 7.4 (logD7.4), total polar surface area (tPSA), stability in hepatic microsomes (LM CLhep), or hepatocytes (Hep CLhep) from human/rat/mouse/dog/cynomolgus monkey (h/r/m/d/c) species, percentage plasma protein binding (PPB %), and permeability across an MDCK cell monolayer from basolateral to apical (B to A) or apical to basolateral (A to B) directions. f, EOL-1 cell line viability in response to GNE-6776 as measured in a 5-day CellTiter-Glo assay performed in experimental triplicates. g, In vivo pharmacokinetic analysis of GNE-6776. Mice (three per group) were dosed PO with 100 mg kg-1 (body weight) or 200 mg kg-1 (body weight) of GNE-6776. Plasma concentrations of GNE-6776 were measured at the indicated time points and the mean ± s.d. (left) or individual data points (right) plotted as a function of time. Exposure metrics relating to the free fraction EC50 for EOL-1 cells are also indicated, where target exposure = (EOL-1 IC50)/(1 − percentage plasma protein binding) = 1.54 μM/0.066 = 23.33 μM. h, Western blot analysis of MCF7-Ser xenografted tumours. Mice bearing MCF7-Ser xenograft tumours were were dosed by mouth with vehicle or 200 mg kg-1 (body weight) GNE-6776 at 0 and 4 h; 8 h after the initial treatment, tumours were excised and the indicated proteins were examined by immunoblotting tumour lysates. Data show three biological replicates. i, Western blot analysis of EOL-1 xenografted tumours. Mice bearing EOL-1 xenograft tumours were dosed by mouth with vehicle or 200 mg kg-1 (body weight) GNE-6776 at 0 and 4 h; 8 h after the initial treatment, tumours were excised and the indicated proteins were examined by immunoblotting tumour lysates. Data show three biological replicates. j, EOL-1 xenograft growth inhibition study of mice administered vehicle or the indicated doses of GNE-6776 by mouth with n = 7 mice per group. The mean ± s.e.m. (top) or individual data points (bottom) are plotted as a function of time. The P values were calculated using Dunnett’s multiple comparison test. Asterisks indicate significant growth inhibition relative to vehicle-treated mice. Day 4 100 mg kg-1 (body weight) P = 0.0163, day 4 200 mg kg-1 (body weight) P = 0.0138, day 6 200 mg kg-1 (body weight) P = 0.0344. Graph source data for g and j can be accessed in the Supplementary Information. Gel source data: Supplementary Fig. 1.
Extended Data Figure 5 Bioinformatics analysis of USP7 inhibitor cell viability screens.
a, Schematic of the cellular viability assay workflow and bioinformatics analysis. The six tumour cell line indications included leukaemias, lymphomas, lung carcinomas, and breast, colon, and prostate adenocarcinomas. b, Histogram of IC50 values of GNE-6640, GNE-6446, and GNE-6641 in 181 cell lines. Mean viability is calculated as the arithmetic average of the fitted viabilities at each tested dose of GNE-6640 or GNE-6446 normalized to the mean viability of GNE-6641. See also Supplementary Table 1. c, Univariate analysis of features associated with viability differences. The x axis represents the fold change (log2) in normalized mean viability between cell lines present or absent for a given feature. The y axis represents the nominal P value (−log10 scale). Features with q values less than 0.05 and absolute log2(fold change) greater than 0.1 are coloured in red. Features with only absolute log2(fold change) greater than 0.1 are coloured in gold. Other analysed features that did not reach significance are indicated in grey. P values were determined using the two-sided Student’s t-test, and q values were determined by correcting resulting P values for multiple hypothesis testing using the Benjamini–Hochberg approach. The size of each point corresponds to the number of cell lines present with the feature. d, e, Boxplots of selected features and their respective associations with normalized mean viability. The respective P and q values are indicated below and were calculated as outlined in the Methods. ‘TP53 LOF’ indicates loss-of-function mutations and previously identified hotspot mutations in TP53 (ref. 27). The number of samples, minima, 25th percentile, centre, 75th percentile, and maxima for each boxplot are as follows: GNE-6640 TP53 LOF TRUE (20, −0.261, −0.119, −0.071, 0.017, 0.304), GNE-6640 TP53 LOF FALSE (156, −0.623,-0.186, −0.084, −0.008, 0.294), GNE-6640 TP53 WT TRUE(80, −0.623, −0.182, −0.085, −0.014, 0.293), GNE-6640 TP53 WT FALSE (96, −0.425, −0.190, −0.083, 0.012, 0.304), GNE-6776 TP53 LOF TRUE (20, −0.166, −0.082, −0.003, 0.029, 0.105), GNE-6776 TP53 LOF FALSE(156, −0.434, −0.087, −0.032, 0.015, 0.225), GNE-6776 TP53 WT TRUE (80, −0.434, −0.098, −0.036, 0.008, 0.225), GNE-6776 TP53 WT FALSE(96, −0.304, −0.077, −0.021, 0.023, 0.113), GNE-6640 AMl TRUE(8, −0.623, −0.394, −0.340, −0.271, −0.222), GNE-6640 AMl FALSE(168, −0.488, −0.173, −0.079, 0.001, 0.304), GNE-6640 TP53 Y220 TRUE(3, 0.070, 0.091, 0.112, 0.142, 0.171), GNE-6640 TP53 Y220 FALSE(173, −0.623, −0.184, −0.085, −0.010, 0.304), GNE-6640 TP53 R175 TRUE(5, −0.425, −0.261, −0.155, −0.119, 0.028), GNE-6640 TP53 R175 FALSE(171, −0.623, −0.183, −0.082, −0.004, 0.304), GNE-6776 AMl TRUE(8, −0.434, −0.216, −0.061, −0.036, −0.009), GNE-6776 AMl FALSE(168, −0.363, −0.085, −0.030, 0.021, 0.225), GNE-6776 TP53 Y220 TRUE(3, −0.033, 0.010, 0.052, 0.070, 0.088), GNE-6776 TP53 Y220 FALSE(173, −0.434, −0.087, −0.031, 0.018, 0.225), GNE-6776 TP53 R175 TRUE(5, −0.304, −0.081, −0.040, −0.034, −0.012), GNE-6776 TP53 R175 FALSE(171, −0.434, −0.087, −0.031, 0.019, 0.225). See Supplementary Table 1 for additional information.
Extended Data Figure 6 Live cell imaging of USP7 inhibitor-treated cells and combination studies with chemotherapeutic and targeted agents.
a–e, Graphs showing cell confluence as a function of time (top rows) and normalized caspase activity (bottom rows) in cells treated with the indicated USP7 inhibitors. Data are representative of three biologically independent experiments and each data point represents the average ± s.e.m. of 12 technical replicates. a, TP53 wild-type or TP53-null HCT-116 colon adenocarcinoma cells. b, TP53 wild-type MCF7 or TP53-null MDA-MB157 breast adenocarcinoma cells. c, TP53 wild-type U2OS cells or TP53-null SaOS osteosarcoma cells. At least three experimental replicates were performed for all experiments. d, Graphs showing cell confluence as a function of time (top rows) and normalized caspase activity (bottom rows) in MCF7 breast adenocarcinoma cells treated with GNE-6640 or doxorubicin alone or in combination. e, Graphs showing cell confluence as a function of time (top rows) and normalized caspase activity (bottom rows) in U2OS osteosarcoma cells treated with GNE-6640 or cisplatin alone or in combination. f, Pie chart illustrating the distribution of compound classes in the Genentech Chemical Genomics Compound library, comprising 589 compounds. NHR, nuclear hormone receptor; GEF, guanine nucleotide exchange factor; DNA, DNA-damaging agent. Graph source data for a–e can be accessed in the Supplementary Information. g, Bar plot visualizing the −log10(transformed P value) from the Wilcoxon rank-sum test evaluating the enrichment of a given compound target over all concentrations of USP7 inhibitors versus DMSO experiments in EOL-1 cells. Only compound targets with three or more compounds in the screen were visualized; the distribution results are indicated by the individual data points from each compound. Higher values indicate synergy with USP7 inhibitors and were further evaluated by Bliss analysis (Fig. 2 and Extended Data Fig. 7). Compounds common to certain signalling pathways including PI3K/PIM, RTK/MAPK, epigenetic regulation, and DNA damage are colour-coded as indicated. See Supplementary Table 2 for additional information.
Extended Data Figure 7 Mechanism of action studies with USP7 inhibitor and PIM inhibitor combinations.
a, Schematic of the PI3K signalling pathway and regulation by PIM kinases. PIM and AKT kinases regulate Bad and TSC1/2 phosphorylation status. Phospho-proteins highlighted in yellow were profiled in cellular studies shown in Fig. 2d. b, Compound structure of GDC-0339. c, PIM inhibitor viability curves at the indicated fixed doses of GNE-6676 in EOL-1 cells (technical duplicates). d, Bliss analysis of 9 × 9 dose–response matrix with PIM inhibitor GDC-0339 and GNE-6776 in EOL-1 cells, representative of technical duplicates. Left: curve-fitted viability values at each dose across the matrix. Zero represents no effect whereas 100 indicates complete loss of viability. Right: the difference in observed versus predicted values using the Bliss independence model. Positive values indicate a greater than predicted decrease in viability. Calculated synergy score = 4.84. e, Immunoblot analysis of cell lysates from the indicated cell lines treated with GNE-6776 (2 μM for 18 h), either alone or in combination with a UAE1 inhibitor MLN-7243 (5 μM for 45 min) or the proteasome inhibitor bortezomib (5 μM for 45 min) as indicated. f, Immunoblot analysis of cell lysates from the indicated cell lines, either untreated or treated with the proteasome inhibitor bortezomib (5 μM for 45 min). g, Immunoprecipitation of cellular lysates using an anti-USP7 antibody or an isotype-matched control antibody followed by western blot analysis of lysates or immunoprecipitates with the indicated antibodies. h, Wild-type recombinant USP7, but not catalytically inactive USP7 C223S, deubiquitinates endogenous polyubiquitinated PIM2 that was immunoprecipitated from proteasome inhibitor-treated MV-4-11 cells. Data in e–h are representative of at least two experimental replicates. Gel source data: Supplementary Fig. 1.
Extended Data Figure 8 Enzymatic analysis and supporting structural biology data for USP7 inhibitors and USP7.
a, Michaelis–Menten kinetic analysis of USP7 and a series of ubiquitin-AMC substrate titrations with the indicated USP7 inhibitors. Initial rate of substrate hydrolysis was determined using the Magellan software on a Tecan Safire2 plate reader and kinetic parameters were modelled using nonlinear regression analysis with GraphPad Prism software. Averages with standard errors were calculated from three technical replicates and are representative of two (GNE-6776) or three (GNE-6640, GNE-6641, and no inhibitor) biologically independent experiments. b, Affinity values of ubiquitin binding to USP7 catalytic domain in the absence and presence of USP7 inhibitors. The values were determined by titration of unlabelled ubiquitin to labelled USP7 catalytic domain and the NMR chemical shift changes were fitted as described in the Methods. The mean values and standard deviations are representative of three experimental replicates and were calculated by averaging the values obtained for eight independent chemical shift perturbations of the well-resolved cross peaks from Y339, S341, D342, G375, A381, G382, D412, and I419. c, Overlay of a region of the two-dimensional 1H/15N TROSY spectrum of the USP7 catalytic domain (orange) highlighting changes induced by binding of ubiquitin in the absence (left, blue) and presence of GNE-6640 (right, black). Individual peaks stemming from residues E371 and Q387 are highlighted. Data are representative of three experimental replicates. d, Comparison of the crystal structure of USP7 catalytic domain in complex with GNE-6640 (cyan) and GNE-6776 (yellow). The catalytic domain is shown as an orange cartoon and the side chains of the residues in proximity to the inhibitor binding sites are shown as orange sticks. GNE-6640 and GNE-6776 compound structures are indicated above. e, PDB accession numbers, data collection, and refinement statistics for GNE-6640 and GNE-6776 crystal structures with the USP7 catalytic domain.
Extended Data Figure 9 Analysis of the functional significance of the interactions between ubiquitin-K48 and USP7-D305, USP7-E308 residues.
a, Titration curves showing the effect of unlabelled wild-type ubiquitin (left) and ubiquitin K48A (right) addition to [2H/15N/13C]USP7 catalytic domain. The weighted combined 1H/15N chemical shift change is plotted against the ubiquitin concentration for five well-resolved peaks stemming from E371, Q387, A381, D342, and Y339 residues in the 1H/15N TROSY spectrum and fitted as described in the Supplementary Information. The standard error was calculated on the basis of the average values over the five fits. Single data points are representative of three experimental replicates. b, Time-course analysis of peptide-conjugated tetra-ubiquitin chains reacted with the USP7 catalytic domain D305A/E308A mutant. Data are representative of three experimental replicates. c, Michaelis–Menten analysis of USP7 catalytic domain D305A/E308A mutant showing the results of three experimental replicates. d, Evaluation of endogenous MDM2 ubiquitination status upon expression of wild-type, C223A, or D305A/E308A full-length USP7. Top: denatured lysates from MCF7 cells transfected with the indicated USP7 expression constructs were immunoprecipitated with a K48 linkage-specific antibody and immunocomplexes were blotted with an MDM2 antibody. Western blot analysis of whole-cell lysates for the indicated proteins are shown below. Data represent three experimental replicates. Gel source data: Supplementary Fig. 1.
Extended Data Figure 10 Analysis of differential USP7 binding to K48- and K63-linked poly-ubiquitin and depolymerization kinetics.
a, Schematic diagrams of substrate-bound K48-linked polyubiquitin chains (left) and K63-linked chains (right), and proposed USP7 interactions. Dashed lines indicate residue side chains; filled circle indicates an isopeptide bond between the ubiquitin C terminus and a Lys residue side chain. The proximal ubiquitin subunits (ligated to the substrate) and their K48 or K63 side chains are indicated with ‘a’ subscripts, the next most distal ubiquitin subunits and side chains are indicated with ‘b’ subscripts, and so on. Preferential USP7 binding to free K48 side chains would direct USP7 to the distal ubiquitin subunit of K48 polyubiquitin and promote sequential exo-cleavage, whereas USP7 would bind all subunits of K63-polyubiquitin and promote exo-, endo-, and base-cleavage. b, The 1H/15N SOFAST spectrum of labelled ubiquitin at 100 μM (cyan), superimposed with the spectrum of a 1:1 molar ratio of labelled ubiquitin in the presence of unlabelled USP7 catalytic domain (orange). Ubiquitin residues affected by the USP7 interaction, which results in exchange broadening of the residue cross peaks, are labelled with grey or blue text and correspond to the residues depicted in c. The K48 and K63 residues as well as the N- and C-terminal residues are labelled in red type. The L43 and L50 residues labelled in green type do not broaden upon USP7 binding and serve as internal controls. The box highlights the region depicted in Fig. 4a. Data are representative of three experimental replicates. c, Structure depictions by ribbon diagrams of the covalent complex between USP7 catalytic domain (orange) with ubiquitin (cyan); (PDB accession number 1NBF) (top), K48 linked di-ubiquitin (green; PDB accession number 2KDE) (lower left), and K63 linked di-ubiquitin (purple; PDB accession number 2RR9) (lower right). In all diagrams, highlighted spheres are the residues in ubiquitin that are broadened in the 1H/15N SOFAST spectrum of 1H/15N-labelled protein by addition of unlabelled USP7 catalytic domain (see b for more details). Leu or Thr residues coloured in blue show well-resolved peaks and were amenable to selective 15N labelling of di-ubiquitin, highlighted with asterisks in the corresponding schematic diagrams. Lysine side chains of K48 and K63 are indicated as sticks in red. d, Time-course analysis of TAMRA/peptide-conjugated tetra-ubiquitin chains cleaved by the USP7 catalytic domain (above) and corresponding densitometry plots (below). Left: time course of USP7 catalytic domain-mediated depolymerization of K48-linked tetra-ubiquitin conjugated to a TAMRA-labelled peptide. Right: time course of USP7 catalytic domain-mediated depolymerization of K63-linked tetra-ubiquitin conjugated to a TAMRA-labelled peptide. Data are representative of three experimental replicates. e, A shorter 0–7 min time-course analysis of TAMRA peptide-K63 tetra-ubiquitin conjugate depolymerization by full-length USP7 (top gel) and the corresponding densitometry plot (bottom), representative of three experimental replicates. Gel source data: Supplementary Fig. 1.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-2, Supplementary Tables 1-2 and Supplementary Methods. (PDF 1112 kb)
Supplementary Figures
This file contains Supplementary Figures containing gel source data and graphs, and Supplementary Tables 1 and 2. (PDF 14589 kb)
Rights and permissions
About this article
Cite this article
Kategaya, L., Di Lello, P., Rougé, L. et al. USP7 small-molecule inhibitors interfere with ubiquitin binding. Nature 550, 534–538 (2017). https://doi.org/10.1038/nature24006
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24006
This article is cited by
-
Inhibition of USP7 enhances CD8+ T cell activity in liver cancer by suppressing PRDM1-mediated FGL1 upregulation
Acta Pharmacologica Sinica (2024)
-
Backbone 1H, 13C and 15N resonance assignment of the ubiquitin specific protease 7 catalytic domain (residues 208–554) in complex with a small molecule ligand
Biomolecular NMR Assignments (2024)
-
Targeting ubiquitin specific proteases (USPs) in cancer immunotherapy: from basic research to preclinical application
Journal of Experimental & Clinical Cancer Research (2023)
-
Multimodal data fusion for supervised learning-based identification of USP7 inhibitors: a systematic comparison
Journal of Cheminformatics (2023)
-
SCML2 contributes to tumor cell resistance to DNA damage through regulating p53 and CHK1 stability
Cell Death & Differentiation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.