Abstract
Homologous recombination repairs DNA double-strand breaks and must function even on actively transcribed DNA. Because break repair prevents chromosome loss, the completion of repair is expected to outweigh the transcription of broken templates. However, the interplay between DNA break repair and transcription processivity is unclear. Here we show that the transcription factor GreA inhibits break repair in Escherichia coli. GreA restarts backtracked RNA polymerase and hence promotes transcription fidelity. We report that removal of GreA results in markedly enhanced break repair via the classic RecBCD–RecA pathway. Using a deep-sequencing method to measure chromosomal exonucleolytic degradation, we demonstrate that the absence of GreA limits RecBCD-mediated resection. Our findings suggest that increased RNA polymerase backtracking promotes break repair by instigating RecA loading by RecBCD, without the influence of canonical Chi signals. The idea that backtracked RNA polymerase can stimulate recombination presents a DNA transaction conundrum: a transcription fidelity factor that compromises genomic integrity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Hersh, M. N., Ponder, R. G., Hastings, P. J. & Rosenberg, S. M. Adaptive mutation and amplification in Escherichia coli: two pathways of genome adaptation under stress. Res. Microbiol. 155, 352–359 (2004)
Dillingham, M. S. & Kowalczykowski, S. C. RecBCD enzyme and the repair of double-stranded DNA breaks. Microbiol. Mol. Biol. Rev. 72, 642–671 (2008)
Dixon, D. A. & Kowalczykowski, S. C. The recombination hotspot χ is a regulatory sequence that acts by attenuating the nuclease activity of the E. coli RecBCD enzyme. Cell 73, 87–96 (1993)
Anderson, D. G. & Kowalczykowski, S. C. The translocating RecBCD enzyme stimulates recombination by directing RecA protein onto ssDNA in a χ-regulated manner. Cell 90, 77–86 (1997)
Finkelstein, I. J., Visnapuu, M. L. & Greene, E. C. Single-molecule imaging reveals mechanisms of protein disruption by a DNA translocase. Nature 468, 983–987 (2010)
Park, J. S., Marr, M. T. & Roberts, J. W. E. coli transcription repair coupling factor (Mfd protein) rescues arrested complexes by promoting forward translocation. Cell 109, 757–767 (2002)
Epshtein, V. et al. UvrD facilitates DNA repair by pulling RNA polymerase backwards. Nature 505, 372–377 (2014)
Erie, D. A., Hajiseyedjavadi, O., Young, M. C. & von Hippel, P. H. Multiple RNA polymerase conformations and GreA: control of the fidelity of transcription. Science 262, 867–873 (1993)
Komissarova, N. & Kashlev, M. Transcriptional arrest: Escherichia coli RNA polymerase translocates backward, leaving the 3′ end of the RNA intact and extruded. Proc. Natl Acad. Sci. USA 94, 1755–1760 (1997)
Borukhov, S., Sagitov, V. & Goldfarb, A. Transcript cleavage factors from E. coli. Cell 72, 459–466 (1993)
Koulich, D. et al. Domain organization of Escherichia coli transcript cleavage factors GreA and GreB. J. Biol. Chem. 272, 7201–7210 (1997)
Paul, B. J. et al. DksA: a critical component of the transcription initiation machinery that potentiates the regulation of rRNA promoters by ppGpp and the initiating NTP. Cell 118, 311–322 (2004)
Tehranchi, A. K. et al. The transcription factor DksA prevents conflicts between DNA replication and transcription machinery. Cell 141, 595–605 (2010)
Vinella, D., Potrykus, K., Murphy, H. & Cashel, M. Effects on growth by changes of the balance between GreA, GreB, and DksA suggest mutual competition and functional redundancy in Escherichia coli. J. Bacteriol. 194, 261–273 (2012)
Ponder, R. G., Fonville, N. C. & Rosenberg, S. M. A switch from high-fidelity to error-prone DNA double-strand break repair underlies stress-induced mutation. Mol. Cell 19, 791–804 (2005)
Amundsen, S. K., Taylor, A. F. & Smith, G. R. The RecD subunit of the Escherichia coli RecBCD enzyme inhibits RecA loading, homologous recombination, and DNA repair. Proc. Natl Acad. Sci. USA 97, 7399–7404 (2000)
Morimatsu, K. & Kowalczykowski, S. C. RecFOR proteins load RecA protein onto gapped DNA to accelerate DNA strand exchange: a universal step of recombinational repair. Mol. Cell 11, 1337–1347 (2003)
Oppenheim, A. B., Kobiler, O., Stavans, J., Court, D. L. & Adhya, S. Switches in bacteriophage lambda development. Annu. Rev. Genet. 39, 409–429 (2005)
Friedberg, E. C., Walker, G. C., Siede, W., Wood, R. & Schultz, R. DNA Repair and Mutagenesis 2nd edn (ASM, 2006)
Kamarthapu, V. et al. ppGpp couples transcription to DNA repair in E. coli. Science 352, 993–996 (2016)
Potrykus, K. & Cashel, M. (p)ppGpp: still magical? Annu. Rev. Microbiol. 62, 35–51 (2008)
Ross, W., Vrentas, C. E., Sanchez-Vazquez, P., Gaal, T. & Gourse, R. L. The magic spot: a ppGpp binding site on E. coli RNA polymerase responsible for regulation of transcription initiation. Mol. Cell 50, 420–429 (2013)
Ross, W. et al. ppGpp binding to a site at the RNAP-DksA interface accounts for its dramatic effects on transcription initiation during the stringent response. Mol. Cell 62, 811–823 (2016)
Kuzminov, A. Collapse and repair of replication forks in Escherichia coli. Mol. Microbiol. 16, 373–384 (1995)
Motamedi, M. R., Szigety, S. K. & Rosenberg, S. M. Double-strand-break repair recombination in Escherichia coli: physical evidence for a DNA replication mechanism in vivo. Genes Dev. 13, 2889–2903 (1999)
Churchill, J. J., Anderson, D. G. & Kowalczykowski, S. C. The RecBC enzyme loads RecA protein onto ssDNA asymmetrically and independently of χ, resulting in constitutive recombination activation. Genes Dev. 13, 901–911 (1999)
Anderson, D. G., Churchill, J. J. & Kowalczykowski, S. C. A single mutation, RecBD1080A, eliminates RecA protein loading but not Chi recognition by RecBCD enzyme. J. Biol. Chem. 274, 27139–27144 (1999)
Ivancic´-Bac´e, I. et al. RecFOR function is required for DNA repair and recombination in a RecA loading-deficient recB mutant of Escherichia coli. Genetics 163, 485–494 (2003)
Spies, M. et al. A molecular throttle: the recombination hotspot χ controls DNA translocation by the RecBCD helicase. Cell 114, 647–654 (2003)
García-Rubio, M. et al. Different physiological relevance of yeast THO/TREX subunits in gene expression and genome integrity. Mol. Genet. Genomics 279, 123–132 (2008)
Jeon, C. & Agarwal, K. Fidelity of RNA polymerase II transcription controlled by elongation factor TFIIS. Proc. Natl Acad. Sci. USA 93, 13677–13682 (1996)
Hutchison, C. A., III et al. Design and synthesis of a minimal bacterial genome. Science 351, aad6253 (2016)
Gordon, A. J., Satory, D., Halliday, J. A. & Herman, C. Heritable change caused by transient transcription errors. PLoS Genet. 9, e1003595 (2013)
Vermulst, M. et al. Transcription errors induce proteotoxic stress and shorten cellular lifespan. Nat. Commun. 6, 8065 (2015)
Krisko, A. & Radman, M. Biology of extreme radiation resistance: the way of Deinococcus radiodurans. Cold Spring Harb. Perspect. Biol. 5, a012765 (2013)
Handa, N. et al. Molecular determinants responsible for recognition of the single-stranded DNA regulatory sequence, χ, by RecBCD enzyme. Proc. Natl Acad. Sci. USA 109, 8901–8906 (2012)
Shee, C. et al. Engineered proteins detect spontaneous DNA breakage in human and bacterial cells. Elife 2, e01222 (2013)
Lloyd, R. G. & Buckman, C. Overlapping functions of recD, recJ and recN provide evidence of three epistatic groups of genes in Escherichia coli recombination and DNA repair. Biochimie 73, 313–320 (1991)
Pennington, J. M. & Rosenberg, S. M. Spontaneous DNA breakage in single living Escherichia coli cells. Nat. Genet. 39, 797–802 (2007)
Drolet, M. et al. Overexpression of RNase H partially complements the growth defect of an Escherichia coli ΔtopA mutant: R-loop formation is a major problem in the absence of DNA topoisomerase I. Proc. Natl Acad. Sci. USA 92, 3526–3530 (1995)
Artsimovitch, I. & Landick, R. Pausing by bacterial RNA polymerase is mediated by mechanistically distinct classes of signals. Proc. Natl Acad. Sci. USA 97, 7090–7095 (2000)
Landick, R., Stewart, J. & Lee, D. N. Amino acid changes in conserved regions of the β-subunit of Escherichia coli RNA polymerase alter transcription pausing and termination. Genes Dev. 4, 1623–1636 (1990)
Cockram, C. A., Filatenkova, M., Danos, V., El Karoui, M. & Leach, D. R. Quantitative genomic analysis of RecA protein binding during DNA double-strand break repair reveals RecBCD action in vivo. Proc. Natl Acad. Sci. USA 112, E4735–E4742 (2015)
Kuzminov, A. & Stahl, F. W. Stability of linear DNA in recA mutant Escherichia coli cells reflects ongoing chromosomal DNA degradation. J. Bacteriol. 179, 880–888 (1997)
Jockovich, M. E. & Myers, R. S. Nuclease activity is essential for RecBCD recombination in Escherichia coli. Mol. Microbiol. 41, 949–962 (2001)
Acknowledgements
We thank A. Gordon, G. Ira, D. Bates, H. Dierick, G. Shaulsky, and A. Barker for comments; I. Campbell for assistance with figure design; R. Nehring and M. Joshi for technical help; and S. Amundsen, W. Ross, A. Šimatovic´, C. Rudolph, M. Gottesman, J. Wang, and A. Poteete for sharing strains. The study was supported by National Institutes of Health (NIH) grant R01-GM088653 (C.H.), Dan L. Duncan Cancer Center and P30 CA125123 pilot grant (C.H., S.M.R.), a gift from the W. M. Keck Foundation (S.M.R.), NIH grants R35-GM122598 and DP1-CA174424 (S.M.R.), NIH grant RO1-GM082837 (I.G.), National Science Foundation PHY 1147498, PHY 1430124, PHY 1427654 (I.G.), the Welch Foundation (Q-1759), and the John S. Dunn Foundation (I.G.).
Author information
Authors and Affiliations
Contributions
P.S. and C.H. conceived the study. P.S., J.A.H., and J.L. performed the experiments. M.A.B.N. and L.A.S. analysed the sequencing data. L.A.S. and I.G. performed the mathematical modelling of RecBCD. P.S., C.H., L.A.S., I.G., and S.M.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks N. Savery and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 DSB repair by the RecBCD–RecA pathway, TCR, and RNAP backtracking.
a, The RecBCD tri-subunit complex binds to blunt DSB ends with high affinity. The enzyme uses translocase, helicase, and nuclease activities to move along the DNA while unwinding and degrading the DNA. RecB is a helicase with 3′→5′ translocation polarity, whereas RecD acts on the other strand2. Initially, the 3′ end of DNA is degraded more vigorously than the 5′ end3. When RecBCD encounters an 8-bp Chi site that is recognized by the RecC subunit36, the nuclease polarity is switched and unwinding proceeds at a slower rate resulting in the formation of ssDNA overhangs onto which RecA is loaded. The RecA–ssDNA filament performs the homology search and invades the homology donor (blue). DNA from the donor is copied by DNA polymerase III (DnaE). b, TCR is initiated when RNAP is stalled at a bulky or helix-distorting lesion. UvrD and ppGpp together can pull RNAP backwards20, or alternatively Mfd can promote RNAP forward translocation6. Either of these processes expose the DNA lesion, allowing it to be accessed and repaired by UvrA (blue) and other proteins (not shown) in the nucleotide excision repair pathway19. c, Elongating RNAP can undergo reverse translocation or backtracking along the DNA template. When this happens, the 3′ end of the nascent transcript, which was originally within the active site of RNAP, gets extruded into the secondary channel and transcription elongation cannot continue9. GreA and GreB can independently stimulate the internal hydrolytic cleavage of the RNA, realigning the 3′ end within the active site of RNAP, allowing the restart of transcription elongation9,10.
Extended Data Figure 2 DSB generation and phleomycin phenotype of GreB overexpression.
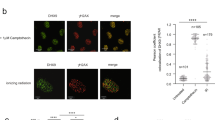
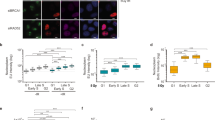
a, DSB formation is unaffected by removal of greA. Mu GamGFP (Gam protein from phage Mu fused to GFP, ref. 37) foci formation in WT and ΔgreA cells after treatment with 20 μg ml−1 phleomycin (PHL) compared with untreated cells (no PHL). Enlarged cells are shown in the upper right corner. Without treatment, 3–5% of WT and ΔgreA cells have only one GamGFP focus. After phleomycin treatment, 87% of WT cells and 77% of ΔgreA cells have multiple GamGFP foci. b, GreB, when present at the appropriate levels, can also modulate phleomycin sensitivity. Representative (one of three) semiquantitative spot assay of tenfold serially diluted log-phase cultures from strains overexpressing GreB from its native promoter on a high copy plasmid at the indicated phleomycin concentrations. GreB overexpression, but not the pBA169 plasmid-only control (–), suppresses the phleomycin resistance of the greA deletion.
Extended Data Figure 3 PFGE analysis.
a, b, Absorption at 600 nm (OD600) (a) and survival (b) measured at each of the time points at which samples were obtained for PFGE in the ΔdksA, ΔdksAΔgreA, and ΔdksAΔrecAΔgreA mutants, described in Fig. 1f, g. c, Intensity of DNA within the well, which is composed of intact circular E. coli genomes, as determined by densitometric analysis. For a–c, n ≥ 3 biological replicates (cultures), data are mean ± s.e.m. d, Representative PFGE of WT and ΔgreA strains before (lanes 1, 7), after 60 min of treatment with 20 μg ml−1 phleomycin (lanes 2, 8), and at indicated times after washing out the drug (lanes 3–6, 9–12). Black bars highlight the observable DNA fragmentation difference between ΔgreA and WT cells at 30 min after removing phleomycin. e, Representative PFGE of the ΔdksAΔrecAΔgreA and ΔdksAΔrecA mutants after treatment with 20 μg ml−1 phleomycin (lanes 2, 7) and at indicated times after washing out the drug (lanes 3–5, 8–10), showing that unrepaired fragmented DNA is eventually degraded in ΔrecA mutants both in the presence and in the absence of GreA. For source data see Supplementary Fig. 1.
Extended Data Figure 4 Factors influencing DSB resistance in the ΔgreA mutant.
a, Deletion of recD does not alter the phleomycin resistance of ΔgreA cells (1 μg ml−1 phleomycin). b, In the absence of RecBCD, an alternative repair mode involving the RecQ helicase, RecJ exonuclease, and RecF-assisted RecA–ssDNA filament formation (by displacing SSB) can repair DSBs2,17,38. c, RecF is not required for the phleomycin (1.5 μg ml−1) resistance of ΔgreA strains. d, RecJ is also not required for the phleomycin (1 μg ml−1) resistance of ΔgreA strains. e, Schematic representation of the I-SceI cutsite engineered within the phage lambda genome (prophage). The I-SceI enzyme is expressed from the doxycycline-induced PN25-tetO promoter. f, Generation of a DSB in the prophage genome reduces the survival of WT and ΔgreA cells equally. g, The DNA damage (SOS) response is activated when RecA bound to ssDNA (RecA*) stimulates the self-proteolytic cleavage of the transcriptional repressor LexA, which is part of the SOS regulon. The SOS response can be monitored using a transcriptional fusion of the promoter of sulA (PsulA) to GFP (ref. 39). h, Flow cytometry analysis of PsulA–GFP expression shows no significant difference between WT and ΔgreA cells, before (0 min) and 60 min after I-SceI DSB induction at the lacA locus. Gating was performed using the lexA3 allele; n = 3, data are mean ± s.e.m.; P ≥ 0.05 (Mann–Whitney U-test). i, Phleomycin resistance of the ΔgreA mutant is unaffected by the inability of cells to activate the SOS response using the lexA3 allele (phleomycin used at 1 μg ml−1). For a, c, d, f, and i, n ≥ 3 biological replicates (cultures), mean ± s.e.m.; **P ≤ 0.01; *P ≤ 0.05; NS, P ≥ 0.05 (Kruskal–Wallis test with multiple testing correction).
Extended Data Figure 5 Pathways of RNAP backtracking and their influence on phleomycin resistance.
a, Overexpression of RnaseH does not affect the phleomycin phenotype of WT or ΔgreA mutants; RnaseH plasmid (pSK760 rnhA) and control plasmid (pSK762 –) (one of three representative biological replicates (cultures)). Plasmid pSK760 was confirmed as RnaseH overexpressing by transforming into a dnaAtsΔrnhA mutant, growth of which would be rescued only if the plasmid expressed WT rnhA40. b, Phleomycin survival phenotypes of the indicated mutants at 1.5 μg ml−1 phleomycin; n ≥ 3 biological replicates (cultures); mean ± s.e.m.; ***P ≤ 0.001; **P ≤ 0.01; *P ≤ 0.05 (Kruskal–Wallis test with multiple testing correction). c, Resistance of ΔgreA mutants to 1.5 μg ml−1 phleomycin is not affected by deletion of uvrA (nucleotide excision repair pathway) or mutS (mismatch repair); n ≥ 2 biological replicates (cultures); mean ± s.e.m. d, e, Representative (one of three) semiquantitative spot assay of cultures grown to absorbance at 600 nm ~ 0.4 in LB, tenfold serially diluted, and plated on LB agar at the indicated phleomycin concentrations. In d, rpoB8, a slow transcribing RNAP mutant prone to backtracking41, is resistant to phleomycin whereas rpoB2, a mutation that makes RNAP elongate faster42, is sensitive to phleomycin. In e, phleomycin resistance of rpoB8 is suppressed by deletion of uvrD.
Extended Data Figure 6 Genome levels effects of XO-seq.
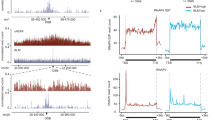
a, Normalized read counts of WT cells across the 4.6 Mb E. coli genome at the indicated times after I-SceI DSB generation at lacA. Read loss is restricted to regions on either side of the break site. However, DNA content changes at oriC and ter can be seen, caused by normalization of each time point relative to the uninduced (0 min) sample. DNA degradation seen around the break site is greater than previously reported for in vivo DSB generation at lac43. This discrepancy could arise from the Tagmentation procedure (Illumina) used to fragment and tag DNA, which captures only double-stranded DNA and may miss ssDNA substrates formed after Chi recognition, or from the differences in the repairability of the two systems. b, c, Normalized read counts of ΔrecD and ΔrecA mutants across the genome at the indicated times after I-SceI induction at lacA, showing reduced DNA degradation in the absence of RecD (b), but extensive ‘Rec-less’44 DNA loss without RecA (c). d, Whole-genome reads from the dnaEts mutant at the indicated times showing read pattern changes at oriC (higher copy number than WT) and ter (lower copy number than WT) consistent with the inactivation of replication by DNA polymerase III (dnaE)25. e, Position and orientation of transcribed genes around the lacA::I-SceI cutsite. f, Quantification of asymmetry in the indicated strains and times after DSB induction. Window size is ±150 kb for all strains and times except for ΔrecD, where window size is ±50 kb.
Extended Data Figure 7 Chi asymmetry effects, mathematical modelling, and extensive DNA degradation identified by XO-seq.
a, Location of 920 kb inversion that surrounds the yjaZ::I-SceI cutsite, but does not include oriC. b, Representative (one of two biological replicates) curves of sequencing in WT and ΔrecD strains compared with strains after inversion of part of the genome (WT and ΔrecD), 60 min after DSB generation at yjaZ. c, Parameters of the mathematical model of RecBCD resection obtained from fitting the degradation curves at 15 min for WT, ΔrecD, and ΔrecA mutants. Also shown are the Chi sensitivities (ε) from each fit; n = 2 biological replicates (cultures); data are mean ± s.d. d, The mathematical model of RecBCD resection fits well with the degradation curve obtained 60 min after I-SceI break induction at lacA in the dnaEts background. e, Representative (one of four) plot of read patterns obtained after I-SceI break induction at lacA in the ΔrecA mutant at the indicated time points. f, XO-seq plots comparing ΔrecA and ΔrecAΔgreA mutants; representative (one of four) plot shown, inset bar plot is from n ≥ 4 biological replicates (cultures); data are mean ± s.d.; NS, P ≥ 0.05.
Extended Data Figure 8 Resection and DNA damage response influence on XO-seq read patterns.
a, Read patterns in WT and the dnaEts mutant 120 min after I-SceI induction at lacA and temperature shift to 42 °C to inactivate dnaE (ref. 25), showing that dnaE function is removed at 120 min. b, Read count patterns in WT and ΔrecB showing greater DNA degradation in the absence of recB. c, Degradation is reduced in the ΔrecBΔrecJ background, suggesting that RecJ contributes to resection when RecB is not present. d, Degradation is similar in ΔrecBΔrecJ and ΔrecBΔrecJΔgreA cells. For a–d, representative plots are shown after inducing I-SceI DSB at lacA for the indicated times, the inset bar plots are from n ≥ 2 biological replicates (cultures); data are mean ± s.d.; **P ≤ 0.01; NS, P ≥ 0.05 (two-tailed two sample t-test). e, f, Read count patterns in WT after treatment with streptomycin (strep) compared with treatment with streptomycin and rifampicin (strep and rif) (e) and WT after treatment with tetracycline (tet) compared with treatment with tetracycline and rifampicin (tet and rif) (f). g, WT after treatment with rifampicin, 15 min after I-SceI induction.
Extended Data Figure 9 Effect of RNAP backtracking on DSB repair in a ‘Chi-free’ region and mechanism of reduced resection in ΔgreA mutants.
a, Comparison of read count patterns after 60 min of DSB induction at the lacA and mhpC loci. Vertical red and green lines show positions of the I-SceI recognition site at lacA and mhpC, respectively, and the pink and green bars show the positions of correctly oriented Chi sites for lacA and mhpC, respectively. b, Read count patterns 60 min after DSB induction at mhpC in WT compared with ΔgreA, and rpoB8ΔgreA mutants, in which increasing backtracked RNAP complexes are formed. c, The rpoB8ΔgreA mutant shows an increased effect on survival after DSB induction at mhpC compared with ΔgreA and WT cells; n = 4 biological replicates (cultures); data are mean ± s.e.m. d, The ΔrecDΔgreA mutant has reduced degradation compared with the ΔrecD mutant alone 60 min after DSB induction at lacA; representative (one of two) plot shown, inset bar plot is from n ≥ 2 biological replicates (cultures); data are mean ± s.d.; **P ≤ 0.01 (two-tailed two sample t-test). e, Substitution of aspartic acid with alanine at position 1080 in the nuclease domain of recB (recBD1080A or recB*) abolishes the ability of RecB to perform nuclease and RecA loading activities16,27,28. RecJ can compensate for the nuclease function and RecF aids in RecA filament formation, resulting in a hybrid resection machine composed of RecB helicase, RecJ exonuclease, and RecF-assisted RecA loading28,45. Suppression of each individual function by the ΔgreA mutant is indicated. f, Phleomycin survival of the indicated mutants as graphed in Fig. 5d. The recB*ΔrecJ and recB*ΔrecJΔgreA mutants produced no countable colonies on phleomycin; n ≥ 3 biological replicates (cultures); data are mean ± s.e.m.; **P ≤ 0.01 (Kruskal–Wallis test with multiple testing correction). g, Representative (one of four) semiqualitative spot assay showing that the ΔgreA deletion can suppress the phleomycin sensitivity of the recB*ΔrecF mutant but not the recB*ΔrecJ mutant at a low concentration of phleomycin (0.5 μg ml−1).
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion, Supplementary Methods, Supplementary References and Supplementary Figure 1. (PDF 11085 kb)
Supplementary Table 1
This file contains a list of bacterial strains used in the study. (XLSX 49 kb)
Rights and permissions
About this article
Cite this article
Sivaramakrishnan, P., Sepúlveda, L., Halliday, J. et al. The transcription fidelity factor GreA impedes DNA break repair. Nature 550, 214–218 (2017). https://doi.org/10.1038/nature23907
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23907
This article is cited by
-
Control of transcription elongation and DNA repair by alarmone ppGpp
Nature Structural & Molecular Biology (2023)
-
Rapid evolution of mutation rate and spectrum in response to environmental and population-genetic challenges
Nature Communications (2022)
-
Mutational analysis of Escherichia coli GreA protein reveals new functional activity independent of antipause and lethal when overexpressed
Scientific Reports (2020)
-
DksA and DNA double-strand break repair
Current Genetics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.