Abstract
Cytosolic DNA arising from intracellular pathogens triggers a powerful innate immune response1,2. It is sensed by cyclic GMP–AMP synthase (cGAS), which elicits the production of type I interferons by generating the second messenger 2′3′-cyclic-GMP–AMP (cGAMP)3,4,5. Endogenous nuclear or mitochondrial DNA can also be sensed by cGAS under certain conditions, resulting in sterile inflammation. The cGAS dimer binds two DNA ligands shorter than 20 base pairs side-by-side6,7,8,9, but 20-base-pair DNA fails to activate cGAS in vivo and is a poor activator in vitro. Here we show that cGAS is activated in a strongly DNA length-dependent manner both in vitro and in human cells. We also show that cGAS dimers form ladder-like networks with DNA, leading to cooperative sensing of DNA length: assembly of the pioneering cGAS dimer between two DNA molecules is ineffective; but, once formed, it prearranges the flanking DNA to promote binding of subsequent cGAS dimers. Remarkably, bacterial and mitochondrial nucleoid proteins HU and mitochondrial transcription factor A (TFAM), as well as high-mobility group box 1 protein (HMGB1), can strongly stimulate long DNA sensing by cGAS. U-turns and bends in DNA induced by these proteins pre-structure DNA to nucleate cGAS dimers. Our results suggest a nucleation-cooperativity-based mechanism for sensitive detection of mitochondrial DNA10 and pathogen genomes11, and identify HMGB/TFAM proteins as DNA-structuring host factors. They provide an explanation for the peculiar cGAS dimer structure and suggest that cGAS preferentially binds incomplete nucleoid-like structures or bent DNA.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Rodero, M. P. & Crow, Y. J. Type I interferon-mediated monogenic autoinflammation: the type I interferonopathies, a conceptual overview. J. Exp. Med. 213, 2527–2538 (2016)
Barber, G. N. STING: infection, inflammation and cancer. Nat. Rev. Immunol. 15, 760–770 (2015)
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013)
Zhang, X. et al. Cyclic GMP-AMP containing mixed phosphodiester linkages is an endogenous high-affinity ligand for STING. Mol. Cell 51, 226–235 (2013)
Chen, Q., Sun, L. & Chen, Z. J. Regulation and function of the cGAS–STING pathway of cytosolic DNA sensing. Nat. Immunol. 17, 1142–1149 (2016)
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP-AMP synthase. Cell 153, 1094–1107 (2013)
Civril, F. et al. Structural mechanism of cytosolic DNA sensing by cGAS. Nature 498, 332–337 (2013)
Li, X. et al. Cyclic GMP-AMP synthase is activated by double-stranded DNA-induced oligomerization. Immunity 39, 1019–1031 (2013)
Zhang, X. et al. The cytosolic DNA sensor cGAS forms an oligomeric complex with DNA and undergoes switch-like conformational changes in the activation loop. Cell Reports 6, 421–430 (2014)
West, A. P. et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 520, 553–557 (2015)
Hansen, K. et al. Listeria monocytogenes induces IFNβ expression through an IFI16-, cGAS- and STING-dependent pathway. EMBO J. 33, 1654–1666 (2014)
Gao, D. et al. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science 341, 903–906 (2013)
Reinert, L. S. et al. Sensing of HSV-1 by the cGAS–STING pathway in microglia orchestrates antiviral defence in the CNS. Nat. Commun. 7, 13348 (2016)
Collins, A. C. et al. Cyclic GMP-AMP synthase is an innate immune DNA sensor for Mycobacterium tuberculosis. Cell Host Microbe 17, 820–828 (2015)
Wassermann, R. et al. Mycobacterium tuberculosis differentially activates cGAS- and inflammasome-dependent intracellular immune responses through ESX-1. Cell Host Microbe 17, 799–810 (2015)
Watson, R. O. et al. The cytosolic sensor cGAS detects Mycobacterium tuberculosis DNA to induce type I interferons and activate autophagy. Cell Host Microbe 17, 811–819 (2015)
Herzner, A. M. et al. Sequence-specific activation of the DNA sensor cGAS by Y-form DNA structures as found in primary HIV-1 cDNA. Nat. Immunol. 16, 1025–1033 (2015)
Holm, C. K. et al. Virus-cell fusion as a trigger of innate immunity dependent on the adaptor STING. Nat. Immunol. 13, 737–743 (2012)
Weiss, J. N. The Hill equation revisited: uses and misuses. FASEB J. 11, 835–841 (1997)
Ngo, H. B., Kaiser, J. T. & Chan, D. C. The mitochondrial transcription and packaging factor Tfam imposes a U-turn on mitochondrial DNA. Nat. Struct. Mol. Biol. 18, 1290–1296 (2011)
Rubio-Cosials, A. et al. Human mitochondrial transcription factor A induces a U-turn structure in the light strand promoter. Nat. Struct. Mol. Biol. 18, 1281–1289 (2011)
Yanai, H. et al. HMGB proteins function as universal sentinels for nucleic-acid-mediated innate immune responses. Nature 462, 99–103 (2009)
Allain, F. H. T. et al. Solution structure of the HMG protein NHP6A and its interaction with DNA reveals the structural determinants for non-sequence-specific binding. EMBO J. 18, 2563–2579 (1999)
Ngo, H. B., Lovely, G. A., Phillips, R. & Chan, D. C. Distinct structural features of TFAM drive mitochondrial DNA packaging versus transcriptional activation. Nat. Commun. 5, 3077 (2014)
Rongvaux, A. et al. Apoptotic caspases prevent the induction of type I interferons by mitochondrial DNA. Cell 159, 1563–1577 (2014)
White, M. J. et al. Apoptotic caspases suppress mtDNA-induced STING-mediated type I IFN production. Cell 159, 1549–1562 (2014)
Bonaldi, T. et al. Monocytic cells hyperacetylate chromatin protein HMGB1 to redirect it towards secretion. EMBO J. 22, 5551–5560 (2003)
Yoh, S. M. et al. PQBP1 is a proximal sensor of the cGAS-dependent innate response to HIV-1. Cell 161, 1293–1305 (2015)
Jønsson, K. L. et al. IFI16 is required for DNA sensing in human macrophages by promoting production and function of cGAMP. Nat. Commun. 8, 14391 (2017)
Sohn, J. & Hur, S. Filament assemblies in foreign nucleic acid sensors. Curr. Opin. Struct. Biol. 37, 134–144 (2016)
Ablasser, A. et al. Cell intrinsic immunity spreads to bystander cells via the intercellular transfer of cGAMP. Nature 503, 530–534 (2013)
Rapino, F. et al. C/EBPα induces highly efficient macrophage transdifferentiation of B lymphoma and leukemia cell lines and impairs their tumorigenicity. Cell Reports 3, 1153–1163 (2013)
Schmidt, T., Schmid-Burgk, J. L. & Hornung, V. Synthesis of an arrayed sgRNA library targeting the human genome. Sci. Rep. 5, 14987 (2015)
Schmid-Burgk, J. L. et al. OutKnocker: a web tool for rapid and simple genotyping of designer nuclease edited cell lines. Genome Res. 24, 1719–1723 (2014)
Gaidt, M. M. et al. Human monocytes engage an alternative inflammasome pathway. Immunity 44, 833–846 (2016)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Tickle, I. J. et al. STARANISO (Global Phasing, 2016)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Read, R. J. Pushing the boundaries of molecular replacement with maximum likelihood. Acta Crystallogr. D 57, 1373–1382 (2001)
Gerber, P. R. & Müller, K. MAB, a generally applicable molecular force field for structure modelling in medicinal chemistry. J. Comput. Aided Mol. Des. 9, 251–268 (1995)
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011)
Yoneyama, M. et al. Autocrine amplification of type I interferon gene expression mediated by interferon stimulated gene factor 3 (ISGF3). J. Biochem. 120, 160–169 (1996)
Lässig, C. et al. ATP hydrolysis by the viral RNA sensor RIG-I prevents unintentional recognition of self-RNA. eLife 4, e10859 (2015)
Schrödinger. The PyMOL molecular graphics system, version 1.8 (2010)
Waterhouse, A. M. et al. Jalview version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009)
Acknowledgements
We thank E. Kremmer for the generation of antibodies, A. Butryn for help with structure determination, K. Lammens and G. Witte for help with crystallization, S. Somarokov for help with protein co-localization studies, S. Bauernfried for help with cell studies, C. Isakaj and O. Fettscher for technical assistance, H. Harz and F. Schüder for advice on staining, F. Civril for cGAS constructs, T. Graf for BLaER1 cells, T. Cremer for fibroblasts, and T. Fujita for the p-125luc reporter plasmid. We thank the Swiss Light Source (Villigen), the European Synchrotron Radiation Facility (Grenoble), and the DESY Petra III (Hamburg) for technical assistance. This work was funded by German Research Foundation grant HO2489/8-1 to K.-P.H., and the Center for Integrated Protein Sciences to K.-P.H., H.L., and V.H. L.A. acknowledges the International Max Planck Research School for Molecular Life Sciences. C.L. and K.-P.H. acknowledge support from BioSysNet (Bavarian Ministry of Education). D.J.D. and C.C.O.M. acknowledge German Research Foundation RTG1721.
Author information
Authors and Affiliations
Contributions
L.A. performed crystallographic and biochemical studies. B.H. performed enzyme-linked immunosorbent assay (ELISA) assays and IFN-β mRNA expression analysis. D.K. built and refined the structure. C.L. performed co-immunopurification studies. C.C.O.M. established staining protocols for three-dimensional structured illumination microscopy (3D SIM). D.J.D. performed luciferase reporter assays and analysed cGAS products. A.M. performed microscopy. M.G. generated cGAS-deficient BLaER1 cells. H.L., C.C.O.M., C.L., and L.A. designed and interpreted microscopy experiments. V.H., B.H., and C.L. designed and interpreted cell-based experiments. K.-P.H. designed the study, derived the mathematical model, and analysed data. K.-P.H. and L.A. wrote the paper with contributions from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks O. Nureki and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 cGAS activity increases with DNA length in vitro and in vivo.
a, b, CXCL10 cytokine production of WT (a) and cGAS-KO (b) transdifferentiated BLaER1 cells. Cells were transfected with 20, 40, and 60 ng DNA of increasing length (20–100 bp in 5 bp intervals) and herring testis (HT) DNA, and CXCL10 concentration in the supernatant was measured by ELISA. The first two bars of each series represent unstimulated cells and Lipofectamine controls. Shown are mean values ± s.d., n = 3. c, Fold change of IFN-β mRNA expression in WT and cGAS-KO BLaER1 cells transfected with 20–100 bp and herring testis DNA. IFN-β mRNA expression levels were normalized to GAPDH mRNA and the fold change was calculated on the basis of the unstimulated control. Shown are mean values ± s.d., n = 3. d, Radiolabelled cGAMP production of cGAS stimulated with DNA of different lengths (20, 35, 55 bp, and plasmid). cGAS reactions in the presence of [α32P]ATP were stopped at the indicated time points and radiolabelled compounds (shown with black arrows) were visualized. e, f, Radiolabelled cGAMP production by (left to right) hcGAScd, hcGAS, and mcGAScd stimulated with 14, 16, 18, 20, 50 bp and plasmid DNA. cGAS (e, 5 μM; f, 10 μM) was incubated with 325 ng μl−1 (e) or 650 ng μl−1 (f) DNA of indicated length in the presence of ATP, GTP, and [α32P]ATP at 37 °C. The reactions were stopped at the indicated time points and radiolabelled compounds (shown with black arrows) were visualized. g, mcGAScd activity measured by the rate of fATP incorporation into fGAMP (see Extended Data Fig. 2 for the assays) in the presence of linearized or circular plasmid DNA. Mean values of initial cGAS reaction rates (∆F/∆t) are plotted against DNA constructs ± s.d., n = 3. No significant difference between linearized and circular plasmid could be detected.
Extended Data Figure 2 Description of the fluorescence-based cGAS activity assay.
a, Radiolabelled cGAS products obtained with different NTP combinations. Reactions were stopped at the indicated time points, and products separated by thin-layer chromatography and visualized by radiography. b, The principle of the fluorescence-based cGAS activity assay. cGAS catalyses the conversion of fluorescent ATP analogue fATP (2-aminopurine riboside triphosphate, three orange stars) into less fluorescent fGAMP (fluorescent cGAS product, one orange star), resulting in a gradual decrease in fluorescence intensity during the reaction. c, General workflow for calculating the initial cGAS reaction rates. From initial fluorescence curves (left), the background fluorescence was subtracted and the resulting curve was inverted for better visualization (∆F). Initial rates were calculated as a slope of the linear intervals (red dashed lines) and defined as ∆F/∆t (relative fluorescence units per minute) (right). d, e, fGAMP mobility in anion-exchange chromatography on a MonoQ 5/50 GL column. d, Comparison of fGAMP, produced by cGAS from fATP and GTP (red), 2′3′-cGAMP (black), and 3′3′-cGAMP (blue) mobilities. e, Comparison of fGAMP (red), fATP (black), and GTP (blue) mobilities.
Extended Data Figure 3 Electron densities and comparison with cGAS in complex with 18 bp DNA.
a, 2Fo − Fc electron density of 39 bp DNA within the complex at a contour level of 1σ. b, Close-up view of the cGAS active site (orange) superposed with a previously published structure of cGAS bound to 18 bp DNA (PDB accession number 4LEY, blue). Shown is the 2Fo − Fc electron density of the active-site residues at a contour level of 1σ. c, Superposition of the cGAS complex with 39 bp DNA (green and grey) with 18 bp DNA-bound cGAS (PDB accession number 4LEY, blue). A slight rotation of superposed 18 bp-cGAS (blue) relative to 39 bp-cGAS (grey) is shown with a black arrow. The superposition shows the difference between a hypothetical straight (red dashed line) DNA, leading the previously proposed DNA end preference of cGAS, and the curved DNA observed in our crystal structure. d, Binding mode of the DNA strands to cGAS within the previously published structure (PDB accession number 4LEY). Two neighbouring crystallographic asymmetric units represent a ‘head-to-tail’ cGAS dimer orientation. Elongation of both strands (red dashed line) leads to a steric clash (orange star). e, Schematic model of cGAS binding to continuous DNA in alternating ‘head-to-head’ and ‘tail-to-tail’ arrangements. DNA curves over λ bp (black arrows). f, Schematic model of cGAS binding to continuous DNA in a (not observed) ‘head-to-tail’ arrangement. DNA curves over λ/2 bp (black arrows), if the same density of cGAS per DNA is assumed. Thus DNA must be bent twice more often than in model e to sustain the parallel DNA arrangement and is energetically less favourable.
Extended Data Figure 4 Asymmetric unit of the cGAS–39 bp complex and details of protein–protein and protein–DNA contacts.
a, An overview of the asymmetric unit of 39 bp DNA-bound cGAS. Filled oval-like symbol represents a twofold crystallographic symmetry axis; black dashed lines represent non-crystallographic twofold symmetry axes. The asymmetric unit contains one full and one half ‘head-to-head’-oriented cGAS4–DNA2 complex. Residues mutated to examine four areas (encircled 1–4) are shown as red sticks. b, Close-up view of mutated interfaces: 1, cGAS active site (yellow) with superimposed cGAMP (from PDB accession number 4LEZ, grey) and labelled active-site residues D213 and D307; 2–4, potential protein–protein or DNA–protein interaction sites, respectively. Mutated residues are in red. c, d, Mutational analysis of the described regions of mcGAScd in vitro and hcGAS in vivo, respectively. D307N and D319N correspond to active-site mutations in mouse and human cGAS, respectively. c, cGAS activity measured by the rate of fATP incorporation into fGAMP (∆F/∆t, see Extended Data Fig. 2) and normalized to WT. Mean values represent percentage of WT activity ± s.d., n = 3. d, Percentage change of IFN-β promoter-driven luciferase activity upon the expression of WT or mutant cGAS. IFN-β response was measured as a proportion of firefly (FF) to Renilla (REN) luciferase activity in HEK293T STING-KI cells upon Flag/HA–hcGAS overexpression. All ratios were normalized to WT. Mean values represent the percentage of WT activity ± s.d., n = 3. Mutants are named according to their position in hcGAS: D319N (active site), Q454R, S263K, R166E, K173E, K458E, K258E mutants refer to D307N, Q439R, S249K, K151E, R158E, R443E, and R244E in mcGAS.
Extended Data Figure 5 cGAS affinity to DNA increases with DNA length accompanied by increase in number of cGAS binding sites along the DNA.
ITC measurements of mcGAScd binding to 20, 40, 45, 50, 70, and 80 bp DNA. For each DNA, the power differential (DP) is plotted against time and ∆H is plotted against the molar ratio of DNA:cGAS. Calculated binding parameters are given on each graph.
Extended Data Figure 6 cGAS forms higher-ordered structures on long DNA.
a, Signature plots for each ITC measurement (Extended Data Fig. 5) showing ∆G (green), ∆H (blue), and −T∆S (red) for each binding reaction. b, ∆H and −T∆S components of Gibbs free energy measured with ITC for mcGAScd binding to 20–80 bp DNA are plotted against DNA length ± s.d. of the measured values from the fit. c, Molecular mass distribution within SEC peaks (Fig. 2c–f) containing mcGAScd1–DNA1 (with 20 bp DNA), mcGAScd4–DNA2 (with 50 bp DNA), and mcGAScd6–DNA2 (with 70 and 80 bp DNA) calculated with RALS. Refractive index (red) and estimated molecular mass (black) are plotted against retention volume.
Extended Data Figure 7 TFAM enhances cGAS activity by prearranging DNA into U-shape.
a, DNA ladders (blue) and (b) DNA-bound TFAM structure (PDB 3TMM, red) arrange DNA in a remarkably similar fashion. The two similarly spaced DNA strands flanking cGAS ladders or the TFAM U-turn (black dotted lines) provide a possible explanation for the effect of TFAM on cGAS activity. c, Radiolabelled cGAMP production in the presence of increasing TFAM concentrations. cGAS reactions with ATP, GTP, and [α32P]ATP were stopped at the indicated time points and the radiolabelled compounds (shown with black arrows) were visualized. d, Activation of mcGAScd by mTFAM and DNA of increasing length (20–200 bp or plasmid DNA). Mean values of initial cGAS reaction rates (∆F/∆t, see Extended Data Fig. 2) are plotted against increasing concentrations of mTFAM ± s.d., n = 4–8.
Extended Data Figure 8 HMGB proteins activate cGAS through DNA bending.
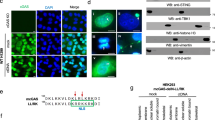
a, Sequence alignment of HMG boxes from different HMG proteins. Positions of intercalating residues, responsible for the DNA-bending activity (red), are shown with respect to their location within helices I, II, or III of HMGB1_boxA (green lines) and HMGB1_boxB (blue lines). b, Structures of box A in complex with DNA (PDB accession number 4QR9) and box B (PDB accession number 1HME) of rat HMGB1 with indicated intercalating residues (red). c, mcGAScd activity measured by the rate of fATP incorporation into fGAMP (see Extended Data Fig. 2) in the presence of increasing concentration of mHMGB1dCTT WT and intercalating residues mutants. Double mutations (F38A in box A, and F103A or I122A in box B) or triple mutants (F38A in box A, and both F103A and I122A in box B) were used. Mean values of initial cGAS reaction rates (∆F/∆t) are plotted against increasing concentrations of mHMGB1dCTT ± s.d., n = 7. d, Electrophoretic mobility shift assay of mHMGB1dCTT WT and point mutants with 2.6 ng μl−1 (~200 nM binding sites) 200 bp DNA. Introduced mutations do not reduce DNA binding of mHMGB1 mutants under conditions used for cGAS activity assays (c). e, mcGAScd activity measured by the rate of fATP incorporation into fGAMP (see Extended Data Fig. 2) in the presence of mTFAM with circular or linearized plasmid. Mean values of initial cGAS reaction rates (∆F/∆t) are plotted against increasing concentrations of mTFAM ± s.d., n = 3. f, ITC of hcGAS with hTFAM at 35, 25, and 15 °C. Power differential (DP) is plotted against time of the experiment. hTFAM (~530 μM) was titrated to ~50 μM hcGAS. No binding is observed, indicating Kd > 100 μM. g, h, cGAS activity measured by the rate of fATP incorporation into fGAMP (see Extended Data Fig. 2) in the presence of increasing TFAM concentrations. Mean values of initial cGAS reaction rates (∆F/∆t) are plotted against increasing concentrations of TFAM ± s.d., n = 3 or 4. mcGAScd (g) and hcGAS (h) are activated by both human and mouse TFAM.
Extended Data Figure 9 TFAM relocalizes from mitochondria into cytosol during mitochondrial stress.
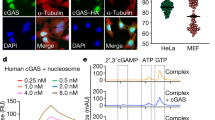
a, b, TFAM (green) localization upon mitochondrial stress induction. Cells were incubated with caspase and Bcl-2 inhibitors for indicated time and stained for super-resolution fluorescence microscopy (3D SIM). Mitochondria were visualized by TOM20 (mitochondrial import receptor subunit, red) staining, cell nuclei by DAPI (blue). a, TFAM cytosolic localization under mitochondrial stress conditions in HEK293T cells. b, TFAM decondensation in mitochondria and its leakage into cytoplasm in human primary fibroblasts in response to mitochondrial stress. c, Example of statistical analysis in human primary fibroblasts (b). Cells were incubated with caspase and Bcl-2 inhibitors for 6 h and stained for super-resolution fluorescence microscopy (3D SIM). Mitochondria were visualized by TOM20 (mitochondrial import receptor subunit) staining, cell nuclei by DAPI. Areas stained for TFAM (green), TOM20 (red), and nucleus (blue) are defined with yellow line. Cytosolic TFAM spots are depicted on the top left view. d, Comparison of cytosolic TFAM signal in control cells (0 h) and under mitochondrial stress conditions (6 h) (b, c). Plotted are mean values of cytosolic TFAM signal calculated as area (top) or number of spots (bottom) ± s.d. Three cells of each type and three segments per cell were used (n = 9). Single asterisk indicates a statistically significant difference (P < 0.05, P = 0.0267), double asterisk indicates a statistically significant difference (P < 0.01, P = 0.0050), compared with control. The TFAM signal in the cytosol of control cells (0 h) represents background signal. e, Co-immunopurification of cGAS and TFAM. Cell lysates with overexpressed Flag/HA–cGAS and HA–TFAM were incubated with anti-hcGAS or with a control antibody (A/b) and the proteins in total lysates (TL) and elution fractions (IP) were visualized by immunoblotting (IB).
Extended Data Figure 10 HMGB1 but not eGFP co-localizes with cytosolic DNA.
a, b, HMGB1 (red), eGFP (green), and DNA (blue) localization in WT or HMGB1-KO MEFs. DNA was stained by DAPI (blue). Images were obtained by wide-field fluorescence microscopy (pDV). Fluorescent signal within enlarged images is enhanced until saturation of the nuclear signal to better visualize cytoplasmic structures. a, Cells were transfected with eGFP expression construct. HMGB1 co-localizes with DAPI-staining in WT but not HMGB1-KO cells. eGFP does not co-localize with DNA or HMGB1. b, In non-transfected cells, HMGB1 and DAPI stainings are present only in the nucleus. The HMGB1 staining in the cytosol of non-transfected and HMGB1-KO cells represents background signals that become visible with the artificial signal amplification beyond saturation, as shown in the zoom-in images.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Supplementary Tables 1-3. The Supplementary Methods contain a detailed explanation of the DPL model and the derivation of equations for its mathematical description. They also contain a description of the global fitting of the experimental data with DPL and generic Hill equations. (PDF 794 kb)
Rights and permissions
About this article
Cite this article
Andreeva, L., Hiller, B., Kostrewa, D. et al. cGAS senses long and HMGB/TFAM-bound U-turn DNA by forming protein–DNA ladders. Nature 549, 394–398 (2017). https://doi.org/10.1038/nature23890
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23890
This article is cited by
-
Mitochondrial DNA replication stress triggers a pro-inflammatory endosomal pathway of nucleoid disposal
Nature Cell Biology (2024)
-
cGAMP-activated cGAS–STING signaling: its bacterial origins and evolutionary adaptation by metazoans
Nature Structural & Molecular Biology (2023)
-
Molecular mechanisms of mitochondrial DNA release and activation of the cGAS-STING pathway
Experimental & Molecular Medicine (2023)
-
Possible frequent multiple mitochondrial DNA copies in a single nucleoid in HeLa cells
Scientific Reports (2023)
-
Computational design of constitutively active cGAS
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.