Abstract
The constant regeneration of stomach epithelium is driven by long-lived stem cells1,2,3, but the mechanism that regulates their turnover is not well understood. We have recently found that the gastric pathogen Helicobacter pylori can activate gastric stem cells and increase epithelial turnover4, while Wnt signalling is known to be important for stem cell identity and epithelial regeneration in several tissues5. Here we find that antral Wnt signalling, marked by the classic Wnt target gene Axin2, is limited to the base and lower isthmus of gastric glands, where the stem cells reside. Axin2 is expressed by Lgr5+ cells, as well as adjacent, highly proliferative Lgr5− cells that are able to repopulate entire glands, including the base, upon depletion of the Lgr5+ population. Expression of both Axin2 and Lgr5 requires stroma-derived R-spondin 3 produced by gastric myofibroblasts proximal to the stem cell compartment. Exogenous R-spondin administration expands and accelerates proliferation of Axin2+/Lgr5− but not Lgr5+ cells. Consistent with these observations, H. pylori infection increases stromal R-spondin 3 expression and expands the Axin2+ cell pool to cause hyperproliferation and gland hyperplasia. The ability of stromal niche cells to control and adapt epithelial stem cell dynamics constitutes a sophisticated mechanism that orchestrates epithelial regeneration and maintenance of tissue integrity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Barker, N. et al. Lgr5+ve stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell 6, 25–36 (2010)
Hayakawa, Y. et al. CCK2R identifies and regulates gastric antral stem cell states and carcinogenesis. Gut 64, 544–553 (2015)
Matsuo, J. et al. Identification of stem cells in the epithelium of the stomach corpus and antrum of mice. Gastroenterology 152, 218–231 (2017)
Sigal, M. et al. Helicobacter pylori activates and expands Lgr5+ stem cells through direct colonization of the gastric glands. Gastroenterology 148, 1392–404.e21 (2015)
Clevers, H., Loh, K. M. & Nusse, R. An integral program for tissue renewal and regeneration: Wnt signaling and stem cell control. Science 346, 1248012 (2014)
Leushacke, M., Ng, A., Galle, J., Loeffler, M. & Barker, N. Lgr5+ gastric stem cells divide symmetrically to effect epithelial homeostasis in the pylorus. Cell Reports 5, 349–356 (2013)
Lee, E. R. & Leblond, C. P. Dynamic histology of the antral epithelium in the mouse stomach: II. Ultrastructure and renewal of isthmal cells. Am. J. Anat. 172, 205–224 (1985)
Koo, B. K. et al. Tumour suppressor RNF43 is a stem-cell E3 ligase that induces endocytosis of Wnt receptors. Nature 488, 665–669 (2012)
de Lau, W. et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature 476, 293–297 (2011)
Barker, N., Bartfeld, S. & Clevers, H. Tissue-resident adult stem cell populations of rapidly self-renewing organs. Cell Stem Cell 7, 656–670 (2010)
Stange, D. E. et al. Differentiated Troy+ chief cells act as reserve stem cells to generate all lineages of the stomach epithelium. Cell 155, 357–368 (2013)
Muñoz, J. et al. The Lgr5 intestinal stem cell signature: robust expression of proposed quiescent ‘+4’ cell markers. EMBO J. 31, 3079–3091 (2012)
Neufeld, S. et al. A conditional allele of Rspo3 reveals redundant function of R-spondins during mouse limb development. Genesis 50, 741–749 (2012)
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007)
Tian, H. et al. A reserve stem cell population in small intestine renders Lgr5-positive cells dispensable. Nature 478, 255–259 (2011)
Yan, K. S. et al. The intestinal stem cell markers Bmi1 and Lgr5 identify two functionally distinct populations. Proc. Natl. Acad. Sci. USA 109, 466–471 (2012)
Tomasetti, C. & Vogelstein, B. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science 347, 78–81 (2015)
Seshagiri, S. et al. Recurrent R-spondin fusions in colon cancer. Nature 488, 660–664 (2012)
Wilhelm, F. et al. Novel insights into gastric cancer: methylation of R-spondins and regulation of LGR5 by SP1. Mol. Cancer Res. 15, 776–785 (2017)
van Amerongen, R., Bowman, A. N. & Nusse, R. Developmental stage and time dictate the fate of Wnt/β-catenin-responsive stem cells in the mammary gland. Cell Stem Cell 11, 387–400 (2012)
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007)
Ueno, H. & Weissman, I. L. Clonal analysis of mouse development reveals a polyclonal origin for yolk sac blood islands. Dev. Cell 11, 519–533 (2006)
Herring, B. P., Hoggatt, A. M., Burlak, C. & Offermanns, S. Previously differentiated medial vascular smooth muscle cells contribute to neointima formation following vascular injury. Vasc. Cell 6, 21 (2014)
Schlaermann, P. et al. A novel human gastric primary cell culture system for modelling Helicobacter pylori infection in vitro. Gut 65, 202–213 (2016)
Churchill, G. A. Fundamentals of experimental design for cDNA microarrays. Nat. Genet. 32 (Suppl.), 490–495 (2002)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Acknowledgements
We thank K. Hoffmann, S. Jackisch, D. Frahm, S. Müllerke, C. Fung, and R. Cooper for technical assistance, R. Zietlow for editing the manuscript, and D. Zhao for comments. Financial support for this work was provided by the Deutsche Forschungsgemeinschaft to M.S. (SI 1983/2-1 and SI 1983/3-1) and T.F.M. (ME 705/18-1) and the Innovation Fund of the Max Planck Society to T.F.M. M.S. was supported by the WILL Foundation and is a participant in the Berlin Institute of Health Charité Clinician Scientist Program funded by the Charité University Medicine Berlin and the Berlin Institute of Health.
Author information
Authors and Affiliations
Contributions
The study was designed by M.S., C.Y.L., M.R.A., and T.F.M. M.S. performed most of the experiments. C.Y.L. performed ISH studies. M.K. and M.S. performed lineage-tracing studies and analysis. Data were analysed by M.S. and M.K. H.J.M. performed microarray studies, and H.B. and H.J.M. analysed the microarray data. R.N. and B.W. gave advice for experimental design and data interpretation. The manuscript was written by M.S. and T.F.M.; all the authors provided comments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. Hatakeyama, J. Mills, T. Wang, and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Axin2 expression and Wnt signalling intensities differ in distinct regions of the gastrointestinal tract.
a–c, Confocal microscopy images of mouse antral glands from an Axin2CreErt2/Rosa26-Rainbow mouse treated with tamoxifen 7 days (a) or 6 weeks (b) before being euthanized, and number of clones per section (c) (n = 2–3 mice per time point). d, e, In the corpus, Axin2+ does not fully trace glands: Confocal microscopy image from the corpus of an Axin2CreErt2/Rosa26-tdTomato mouse treated with tamoxifen 7 days before being euthanized. e, Confocal microscopy image from the corpus of an Axin2CreErt2/Rosa26-tdTomato mouse treated with tamoxifen 120 days before being euthanized. f–h, Image of ISH for Axin2 in the stomach corpus (f), antrum (g), and duodenum (h). Red dots mark Axin2 mRNA transcripts. Note that expression in the duodenum is substantially higher but not restricted to the crypt base. Experiments were repeated at least twice in the laboratory. Data represent mean ± s.e.m. from two or three mice analysed by Student’s t-test. Scale bar, 100 μm.
Extended Data Figure 2 Co-localization of Lgr5 and Axin2.
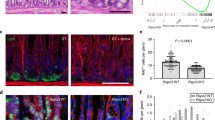
a, Double-ISH for Axin2 (red) and Lgr5 (blue) in the mouse antrum. b, Confocal microscopy images of antrum from Axin2CreErt2/Rosa26-mTmG mice treated with tamoxifen for 1, 2, 3, or 4 days, as well as 6 months, before being euthanized. c, Average numbers of Axin2+ cells in different cell positions counted from the gland base at day 1, 2, 3 and 4 after tamoxifen treatment. A rapid expansion of the signal is observed. d, Confocal microscopy images of antrum from Axin2CreErt2/Rosa26-mTmG and Lgr5CreErt2/Rosa26-mTmG mice treated with tamoxifen 7 days before being euthanized. e, Haematoxylin and eosin images of antral glands. The gland base contains mucous cells that are more differentiated (arrowheads), whereas cells in the lower isthmus above appear less differentiated and have larger nuclear to cytoplasmic ratios (arrows). f, Confocal images of antrum from Lgr5DTR/Axin2CreErt2/Rosa26-mTmG mice treated with tamoxifen 24 h before being euthanized (left), tamoxifen and diphtheria toxin 24 h before being euthanized (middle), and tamoxifen and diphtheria toxin 30 days before being euthanized (right). Arrows indicate Axin2+/Lgr5− cells in the isthmus, arrowheads indicate Lgr5+ cells in the base. Images represent findings that were reproduced at least twice in the laboratory in independent biological replicates. Scale bar, 100 μm.
Extended Data Figure 3 Wnt ligand expression in the antrum.
In situ hybridization of Wnt ligands in the mouse antrum. For the probes that detected a signal, a higher magnification image is also displayed. Red dots mark the mRNA of the respective target. Scale bar, 100 μm. Images represent findings that were reproduced at least twice in the laboratory on independent biological replicates. Scale bar, 100 μm.
Extended Data Figure 4 Wnt receptors and inhibitors expression in the antrum.
a–c, In situ hybridization of the Wnt receptors Frizzled (Fz)1–10 (a), Dkk1–4 (b), as well as Troy and RNF-43 (c) in the mouse antrum. Images represent findings that were reproduced at least twice in the laboratory on independent biological replicates. Scale bar, 100 μm.
Extended Data Figure 5 Rspo expression in the antrum.
a, In situ hybridization of Rspo1–4 in the mouse antrum with an inset displaying the Rspo3 signal in the stroma at high magnification. Scale bar, 100 μm. b, Expression of Rspo1 was observed in the forestomach at lower level, but not in the antrum on the same slide. c, ISH for Rspo1–4 in embryonic day 10.5 mouse embryo, displaying distinct patterns for all four probes in the developing brain and limb bud. Pol2 and DapB served as positive and negative controls. d, e, qPCR from cultured myofibroblasts and epithelial organoids isolated from mouse antrum for α-SMA, vimentin, and collagen 1A1 (d) and Rspo homologues (e). f, Western blot for Rspo3 from epithelial organoids and cultured myofibroblasts (Con, positive control, HEK293T cells). Epi, antral epithelial three-dimensional organoids; Str, cultures of antral stroma. Experiments were repeated at least twice in the laboratory. Data represent mean ± s.e.m. from n = 3 biological replicates, analysed by Student’s t-test. Scale bar, 100 μm.
Extended Data Figure 6 a
, Human gastric organoids require Rspo for active Wnt signalling: images from human gastric antrum-derived organoids grown from FACS sorted RFP+ cells from a culture infected with BAT-RED lentiviral construct to visualize Wnt signalling. Cells were grown either in full medium (left column), without Rspo (middle column), or without either Rspo or Wnt3a (right column). b, c, Expression of Wnt target genes in organoids requires Rspo qPCR analysis of Axin2 (b) and Lgr5 (c) expression in the human organoids grown in full medium or medium lacking Rspo or Rspo and Wnt3a. d, Organoid growing capacity is higher in antral cells from Rspo1 pre-treated mice plated at low density. Images show cells at passage one from two independent biological replicates per group. e, Single Axin2+ cells generate clonal organoids that can be expanded and passaged over months: GFP+ cells from Axin2CreErt2/Rosa26-mTmG mice, pre-treated with tamoxifen 48 h before being euthanized and seeded at clonal density, generate organoids that can be expanded and grown for several passages. Shown are organoids at passages 2 and 6. Organoids were cultured for 3 months and passaged weekly at a ratio of 1:2 or 1:3. Experiments were repeated at least twice in the laboratory. Data represent mean ± s.e.m. from three independent experiments, analysed by one-way ANOVA.
Extended Data Figure 7 Rspo treatment accelerates Axin2+ cell tracing.
a, Axin2CreErt2/Rosa26-mTmG mice were treated with tamoxifen 4 h before administration of Rspo1 or PBS and euthanized 48 h later. Tile scans and high magnification of antral glands are shown. The tracing ratio (distance between the lowest and highest tdTomato+ cell divided by the gland height) of antral glands was quantified (n = 2 mice per group). b, Axin2CreErt2/Rosa26-mTmG mice were treated with tamoxifen 24 h before Rspo1, Rspo3, or PBS and euthanized 24 h later; shown are antral glands and their tracing ratio (n = 3 mice per group). c, Rspo does not expand the Lgr5eGPF+ cell population. Lgr5eGFP mice were treated with PBS or 100 μg Rspo3 48 h before being euthanized. Right: graph demonstrating the highest position of GFP+ cells per gland in PBS versus Rspo3-treated animals. Data represent mean ± s.e.m. from antral glands analysed in two mice per group. d, Confocal microscopy images of Ki67-labelled antrum of C57Bl6 mice treated either with PBS or Rspo1 48 h before being euthanized (left) and Lgr5-eGFP mice treated either with PBS or Rspo3 48 h before being euthanized (right). Experiments were repeated at least twice in the laboratory. Except where indicated, data represent mean ± s.e.m. from n = 3 mice, analysed by Student’s t-test. Scale bar, 100 μm.
Extended Data Figure 8 Rspo transiently affects turnover dynamics in the gland and could interact through Lgr4 receptors.
a, Confocal microscopy images from the antrum of Lgr5CreErt2/Rosa26-mTmG mice treated with tamoxifen and PBS (top) or tamoxifen and Rspo1 (bottom) 7 days before being euthanized. b, Tracing ratio from all glands with membrane-bound GFP signal in a longitudinal section from mice treated with either Rspo1 or PBS, as well as tamoxifen at 7 and 14 days before being euthanized. In addition, one group received four injections of Rspo1 for 4 consecutive days, as well as tamoxifen together with the last Rspo injection 7 days before being euthanized. Further, one group received tamoxifen at 7 days and Rspo at 5 days before being euthanized. Each dot represents a single gland, Data represent mean ± s.e.m. from antral glands of two mice per group. c, Rspo-driven Axin2 expansion is a transient response. Confocal microscopy image from a mouse antrum of an Axin2CreErt2/Rosa26-tdTomato mouse treated with 100 μg Rspo1 (left) or PBS (right) 8 days and tamoxifen 48 h before being euthanized (n = 2 mice per group). d, Single-molecule ISH for the Lgr homologues Lgr4, Lgr5, and Lgr6 in mouse antrum. Images represent findings that were reproduced at least twice in the laboratory in independent biological replicates.
Extended Data Figure 9 Expression of Axin2 is increased upon H. pylori infection.
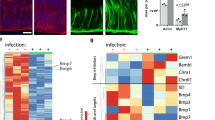
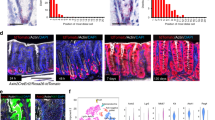
a, Quantification of Axin2+/Ki67+ cells and Axin2+/Ki67− cells in glandular positions 1–3 and 4–12 in the antrum of uninfected mice and mice infected with H. pylori for 2 months; n = 3 mice per group. b, Single-molecule ISH for Axin2 in uninfected and 1-month infected mouse antrum and (c) quantification of the ISH images (number of Axin2mRNA transcripts per image, n = 3 mice per group). d, e, Images and quantification of short-term Axin2 lineage tracing (48 h) from uninfected and 2-month infected Axin2CreErt2/Rosa26-mTmG mice infected with either wild type (WT) H. pylori strain PMSS1 or an isogenic strain with a mutation in the CagE gene. f, Images from uninfected and 2-month infected Myh11CreErt2/Rosa26-mTmG mice. Tamoxifen was injected 7 days before mice were euthanized to visualize myofibroblasts/smooth muscle cells in the stomach (red, actin; blue, nuclei; green, Myh11+ myofibroblasts). g, Upon infection with H. pylori, Rspo3 is expressed in α-SMA+ myofibroblasts but not in CD45+ blood cells or in PECAM+ endothelial cells: double-ISH for Rspo3 (blue) and either α-SMA, CD45, or PECAM (red) in mouse antrum infected with H. pylori 2 months before being euthanized. Scale bar, 100 μm. h, Quantification of Rspo3 transcripts between glands detected by ISH. Images represent findings that were reproduced at least twice in the laboratory in independent biological replicates. Data represent mean ± s.e.m. from n = 3 mice, analysed by Student’s t-test.
Supplementary information
Supplementary Information
This file contains uncropped blots. (PDF 675 kb)
Source data
Rights and permissions
About this article
Cite this article
Sigal, M., Logan, C., Kapalczynska, M. et al. Stromal R-spondin orchestrates gastric epithelial stem cells and gland homeostasis. Nature 548, 451–455 (2017). https://doi.org/10.1038/nature23642
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23642
This article is cited by
-
Decoding spatiotemporal transcriptional dynamics and epithelial fibroblast crosstalk during gastroesophageal junction development through single cell analysis
Nature Communications (2024)
-
Impact of media compositions and culture systems on the immunophenotypes of patient-derived breast cancer cells
BMC Cancer (2023)
-
A synergistic partnership between IL-33/ST2 and Wnt pathway through Bcl-xL drives gastric cancer stemness and metastasis
Oncogene (2023)
-
Role of PDGFRA+ cells and a CD55+ PDGFRALo fraction in the gastric mesenchymal niche
Nature Communications (2023)
-
Intracellular bacteria in cancer—prospects and debates
npj Biofilms and Microbiomes (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.