Abstract
Nuclear small RNA pathways safeguard genome integrity by establishing transcription-repressing heterochromatin at transposable elements. This inevitably also targets the transposon-rich source loci of the small RNAs themselves. How small RNA source loci are efficiently transcribed while transposon promoters are potently silenced is not understood. Here we show that, in Drosophila, transcription of PIWI-interacting RNA (piRNA) clusters—small RNA source loci in animal gonads—is enforced through RNA polymerase II pre-initiation complex formation within repressive heterochromatin. This is accomplished through Moonshiner, a paralogue of a basal transcription factor IIA (TFIIA) subunit, which is recruited to piRNA clusters via the heterochromatin protein-1 variant Rhino. Moonshiner triggers transcription initiation within piRNA clusters by recruiting the TATA-box binding protein (TBP)-related factor TRF2, an animal TFIID core variant. Thus, transcription of heterochromatic small RNA source loci relies on direct recruitment of the core transcriptional machinery to DNA via histone marks rather than sequence motifs, a concept that we argue is a recurring theme in evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Fedoroff, N. V. Transposable elements, epigenetics, and genome evolution. Science 338, 758–767 (2012)
Castel, S. E. & Martienssen, R. A. RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat. Rev. Genet. 14, 100–112 (2013)
Holoch, D. & Moazed, D. RNA-mediated epigenetic regulation of gene expression. Nat. Rev. Genet. 16, 71–84 (2015)
Siomi, M. C., Sato, K., Pezic, D. & Aravin, A. A. PIWI-interacting small RNAs: the vanguard of genome defence. Nat. Rev. Mol. Cell Biol. 12, 246–258 (2011)
Czech, B. & Hannon, G. J. One loop to rule them all: the ping-pong cycle and piRNA-guided silencing. Trends Biochem. Sci. 41, 324–337 (2016)
Mohn, F., Sienski, G., Handler, D. & Brennecke, J. The rhino-deadlock-cutoff complex licenses noncanonical transcription of dual-strand piRNA clusters in Drosophila. Cell 157, 1364–1379 (2014)
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007)
Le Thomas, A. et al. Transgenerationally inherited piRNAs trigger piRNA biogenesis by changing the chromatin of piRNA clusters and inducing precursor processing. Genes Dev. 28, 1667–1680 (2014)
Klattenhoff, C. et al. The Drosophila HP1 homolog Rhino is required for transposon silencing and piRNA production by dual-strand clusters. Cell 138, 1137–1149 (2009)
Zhang, Z. et al. The HP1 homolog rhino anchors a nuclear complex that suppresses piRNA precursor splicing. Cell 157, 1353–1363 (2014)
Sainsbury, S., Bernecky, C. & Cramer, P. Structural basis of transcription initiation by RNA polymerase II. Nat. Rev. Mol. Cell Biol. 16, 129–143 (2015)
Buratowski, S., Hahn, S., Guarente, L. & Sharp, P. A. Five intermediate complexes in transcription initiation by RNA polymerase II. Cell 56, 549–561 (1989)
Papai, G. et al. TFIIA and the transactivator Rap1 cooperate to commit TFIID for transcription initiation. Nature 465, 956–960 (2010)
Chen, Y. A. et al. Cutoff suppresses RNA polymerase II termination to ensure expression of piRNA precursors. Mol. Cell 63, 97–109 (2016)
Gu, W. et al. CapSeq and CIP-TAP identify Pol II start sites and reveal capped small RNAs as C. elegans piRNA precursors. Cell 151, 1488–1500 (2012)
Kaufmann, J. & Smale, S. T. Direct recognition of initiator elements by a component of the transcription factor IID complex. Genes Dev. 8, 821–829 (1994)
Purnell, B. A., Emanuel, P. A. & Gilmour, D. S. TFIID sequence recognition of the initiator and sequences farther downstream in Drosophila class II genes. Genes Dev. 8, 830–842 (1994)
Gunawardane, L. S. et al. A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315, 1587–1590 (2007)
Czech, B., Preall, J. B., McGinn, J. & Hannon, G. J. A transcriptome-wide RNAi screen in the Drosophila ovary reveals factors of the germline piRNA pathway. Mol. Cell 50, 749–761 (2013)
Geiger, J. H., Hahn, S., Lee, S. & Sigler, P. B. Crystal structure of the yeast TFIIA/TBP/DNA complex. Science 272, 830–836 (1996)
Tan, S., Hunziker, Y., Sargent, D. F. & Richmond, T. J. Crystal structure of a yeast TFIIA/TBP/DNA complex. Nature 381, 127–134 (1996)
Dantonel, J. C., Quintin, S., Lakatos, L., Labouesse, M. & Tora, L. TBP-like factor is required for embryonic RNA polymerase II transcription in C. elegans. Mol. Cell 6, 715–722 (2000)
Kaltenbach, L., Horner, M. A., Rothman, J. H. & Mango, S. E. The TBP-like factor CeTLF is required to activate RNA polymerase II transcription during C. elegans embryogenesis. Mol. Cell 6, 705–713 (2000)
Kopytova, D. V. et al. Two isoforms of Drosophila TRF2 are involved in embryonic development, premeiotic chromatin condensation, and proper differentiation of germ cells of both sexes. Mol. Cell. Biol. 26, 7492–7505 (2006)
Müller, F., Lakatos, L., Dantonel, J., Strähle, U. & Tora, L. TBP is not universally required for zygotic RNA polymerase II transcription in zebrafish. Curr. Biol. 11, 282–287 (2001)
Veenstra, G. J., Weeks, D. L. & Wolffe, A. P. Distinct roles for TBP and TBP-like factor in early embryonic gene transcription in Xenopus. Science 290, 2312–2315 (2000)
Martianov, I. et al. Late arrest of spermiogenesis and germ cell apoptosis in mice lacking the TBP-like TLF/TRF2 gene. Mol. Cell 7, 509–515 (2001)
Zhang, D., Penttila, T. L., Morris, P. L., Teichmann, M. & Roeder, R. G. Spermiogenesis deficiency in mice lacking the Trf2 gene. Science 292, 1153–1155 (2001)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Yokomori, K. et al. Drosophila TFIIA directs cooperative DNA binding with TBP and mediates transcriptional activation. Genes Dev. 8, 2313–2323 (1994)
Isogai, Y., Keles, S., Prestel, M., Hochheimer, A. & Tjian, R. Transcription of histone gene cluster by differential core-promoter factors. Genes Dev. 21, 2936–2949 (2007)
Rothbauer, U. et al. Targeting and tracing antigens in live cells with fluorescent nanobodies. Nat. Methods 3, 887–889 (2006)
Kloc, A., Zaratiegui, M., Nora, E. & Martienssen, R. RNA interference guides histone modification during the S phase of chromosomal replication. Curr. Biol. 18, 490–495 (2008)
Chen, E. S. et al. Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451, 734–737 (2008)
Law, J. A. et al. Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1. Nature 498, 385–389 (2013)
Law, J. A., Vashisht, A. A., Wohlschlegel, J. A. & Jacobsen, S. E. SHH1, a homeodomain protein required for DNA methylation, as well as RDR2, RDM4, and chromatin remodeling factors, associate with RNA polymerase IV. PLoS Genet. 7, e1002195 (2011)
Zhai, J. et al. A one precursor one siRNA model for Pol IV-dependent siRNA biogenesis. Cell 163, 445–455 (2015)
Hochheimer, A., Zhou, S., Zheng, S., Holmes, M. C. & Tjian, R. TRF2 associates with DREF and directs promoter-selective gene expression in Drosophila. Nature 420, 439–445 (2002)
Goriaux, C., Desset, S., Renaud, Y., Vaury, C. & Brasset, E. Transcriptional properties and splicing of the flamenco piRNA cluster. EMBO Rep. 15, 411–418 (2014)
Hur, J. K. et al. Splicing-independent loading of TREX on nascent RNA is required for efficient expression of dual-strand piRNA clusters in Drosophila. Genes Dev. 30, 840–855 (2016)
Zhang, F. et al. UAP56 couples piRNA clusters to the perinuclear transposon silencing machinery. Cell 151, 871–884 (2012)
Zeidler, M. P., Yokomori, K., Tjian, R. & Mlodzik, M. Drosophila TFIIA-S is up-regulated and required during Ras-mediated photoreceptor determination. Genes Dev. 10, 50–59 (1996)
Ni, J.-Q. et al. A genome-scale shRNA resource for transgenic RNAi in Drosophila. Nat. Methods 8, 405–407 (2011)
Schmitz, M. L., Stelzer, G., Altmann, H., Meisterernst, M. & Baeuerle, P. A. Interaction of the COOH-terminal transactivation domain of p65 NF-κ B with TATA-binding protein, transcription factor IIB, and coactivators. J. Biol. Chem. 270, 7219–7226 (1995)
Lieberman, P. M. & Berk, A. J. A mechanism for TAFs in transcriptional activation: activation domain enhancement of TFIID-TFIIA--promoter DNA complex formation. Genes Dev. 8, 995–1006 (1994)
Kobayashi, N., Boyer, T. G. & Berk, A. J. A class of activation domains interacts directly with TFIIA and stimulates TFIIA-TFIID-promoter complex assembly. Mol. Cell. Biol. 15, 6465–6473 (1995)
Aoyagi, N. & Wassarman, D. A. Genes encoding Drosophila melanogaster RNA polymerase II general transcription factors: diversity in TFIIA and TFIID components contributes to gene-specific transcriptional regulation. J. Cell Biol. 150, F45–F50 (2000)
Duttke, S. H. C., Doolittle, R. F., Wang, Y.-L. & Kadonaga, J. T. TRF2 and the evolution of the bilateria. Genes Dev. 28, 2071–2076 (2014)
Martianov, I. et al. Distinct functions of TBP and TLF/TRF2 during spermatogenesis: requirement of TLF for heterochromatic chromocenter formation in haploid round spermatids. Development 129, 945–955 (2002)
Oyama, T. et al. Cleavage of TFIIA by Taspase1 activates TRF2-specified mammalian male germ cell programs. Dev. Cell 27, 188–200 (2013)
Martianov, I., Velt, A., Davidson, G., Choukrallah, M.-A. & Davidson, I. TRF2 is recruited to the pre-initiation complex as a testis-specific subunit of TFIIA/ALF to promote haploid cell gene expression. Sci. Rep. 6, 32069 (2016)
Brancorsini, S., Davidson, I. & Sassone-Corsi, P. TIPT, a male germ cell-specific partner of TRF2, is chromatin-associated and interacts with HP1. Cell Cycle 7, 1415–1422 (2008)
Venken, K. J. T. et al. Versatile P[acman] BAC libraries for transgenesis studies in Drosophila melanogaster. Nat. Methods 6, 431–434 (2009)
Handler, D. et al. The genetic makeup of the Drosophila piRNA pathway. Mol. Cell 50, 762–777 (2013)
Sarot, E., Payen-Groschêne, G., Bucheton, A. & Pélisson, A. Evidence for a piwi-dependent RNA silencing of the gypsy endogenous retrovirus by the Drosophila melanogaster flamenco gene. Genetics 166, 1313–1321 (2004)
Gokcezade, J., Sienski, G. & Duchek, P. Efficient CRISPR/Cas9 plasmids for rapid and versatile genome editing in Drosophila. G3 (Bethesda) 4, 2279–2282 (2014)
Maggert, K. A., Gong, W. J. & Golic, K. G. Methods for homologous recombination in Drosophila. Methods Mol. Biol. 420, 155–174 (2008)
Dorfer, V. et al. MS Amanda, a universal identification algorithm optimized for high accuracy tandem mass spectra. J. Proteome Res. 13, 3679–3684 (2014)
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015)
Stampfel, G. et al. Transcriptional regulators form diverse groups with context-dependent regulatory functions. Nature 528, 147–151 (2015)
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012)
Lee, T. I., Johnstone, S. E. & Young, R. A. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nat. Protocols 1, 729–748 (2006)
Brown, J. B. et al. Diversity and dynamics of the Drosophila transcriptome. Nature 512, 393–399 (2014)
Dobin, A . et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017)
Nechaev, S. et al. Global analysis of short RNAs reveals widespread promoter-proximal stalling and arrest of Pol II in Drosophila. Science 327, 335–338 (2010)
Jayaprakash, A. D., Jabado, O., Brown, B. D. & Sachidanandam, R. Identification and remediation of biases in the activity of RNA ligases in small-RNA deep sequencing. Nucleic Acids Res. 39, e141 (2011)
Eddy, S. R. Accelerated profile HMM Searches. PLoS Comput. Biol. 7, e1002195 (2011)
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997)
Katoh, K. & Toh, H. Recent developments in the MAFFT multiple sequence alignment program. Brief. Bioinform. 9, 286–298 (2008)
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009)
Cole, C., Barber, J. D. & Barton, G. J. The Jpred 3 secondary structure prediction server. Nucleic Acids Res. 36, W197–W201 (2008)
Wootton, J. C. & Federhen, S. Analysis of compositionally biased regions in sequence databases. Methods Enzymol. 266, 554–571 (1996)
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, 2016)
Wickham, H. ggplot2 (Springer, 2016)
Wickham, H. Reshaping data with the reshape package. J. Stat. Softw. 21 (12), 1–20 (2007)
Wickham, H. scales: scale functions for visualization. R package version 0.4.0. https://CRAN.R-project.org/package=scales (2016)
Bolstad, B. M. preprocessCore: a collection of pre-processing functions. R package version 1.28.0. https://github.com/bmbolstad/preprocessCore (2013)
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002)
Raney, B. J. et al. Track data hubs enable visualization of user-defined genome-wide annotations on the UCSC Genome Browser. Bioinformatics 30, 1003–1005 (2014)
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44 (D1), D447–D456 (2016)
Acknowledgements
We thank K. Meixner for experimental support, D. Handler and D. Jurczak for bioinformatics help, P. Duchek and J. Gokcezade for generating CRISPR-edited and transgenic flies, K. Mechtler and his team for mass spectrometry, T. Lendl for RNA FISH quantification, A. Schleiffer and M. Novatchkova for Moonshiner phylogenetic analysis, the Vienna Biocenter Core Facilities Next Generation Sequencing unit for deep sequencing, M. Elmaghraby for the Deadlock antigen, the Max F. Perutz Laboratories monoclonal facility for the Deadlock antibody, and the Vienna Drosophila RNAi Center, TRiP, and Bloomington stock centers for flies. We thank A. Ordonez, D. Handler, F. Mohn, F. Muerdter, M. Bühler, and especially O. Wueseke (Impulse Science) and Life Science Editors for comments on the manuscript. This work was supported by the Austrian Academy of Sciences and the European Community (ERC grant 260711EU and ERC-2015-CoG-682181). P.R.A. is supported by fellowships from the Alfred Benzon Foundation and the Novo Nordisk Foundation.
Author information
Authors and Affiliations
Contributions
P.R.A. performed the experiments except the genetic bypass and promoter deletion experiments (both L.T.), and the S2 cell-based protein interaction assays (M.V.). P.R.A. and J.B. analysed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks E. Brasset, T. Juven-Gershon and P. Zamore for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Characterization of transcription initiation events at piRNA clusters.
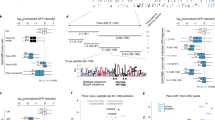
a, Size profile histograms of small RNAs mapping to the Pld gene locus from ovaries with indicated genotypes. siRNAs (21 nt) are highlighted in orange and piRNAs (23–29 nt) are highlighted in green. b, UCSC genome browser panels showing cluster80F for which flanking promoter dependency was investigated by deletion of the promoter region of alpha Catenin. Shown are Pol II occupancy (red), Rhino occupancy (blue), and piRNA levels (black/grey). Flanking transcription units are shown in grey, light grey shading indicates the experimental promoter deletion. As alpha Catenin is an essential gene, a cDNA rescue transgene was expressed from another locus. c, UCSC genome browser panels showing the Cap-seq profile at the promoter of a canonical gene. d, DNA sequence motif at 5′ ends of capped RNAs mapping to Rhino-bound genomic loci (Rhino ChIP-seq reads per kilobase per million mappers > 300; cluster80F and 42AB excluded) outside known transcription units. e, DNA sequence motif at 5′ ends of 5′-monophosphorylated RNAs mapping to cluster42AB or cluster80F. The schematic to the right shows how the ‘ping-pong’ amplification loop involving Aub- and Ago3-mediated cleavages gives rise to the observed sequence biases at positions +1 and +10. f, Histogram of the ‘YR’ dinucleotide occurrence around cluster42AB and cluster80F transcription start sites (expected chance occurrence 25%).
Extended Data Figure 2 CG12721/Moonshiner is a germline-specific TFIIA-L paralogue.
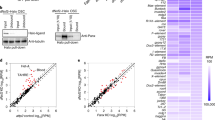
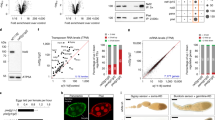
a, Expression levels of indicated genes in larval/adult tissues on the basis of modENCODE RNA-seq data; rpkm, reads per kilobase per million mappers. b, The top schematic denotes the two regions of homology shown in Fig. 2a. Shown below is the amino-acid sequence alignment of these two regions from drosophilid species (Moonshiner) and selected insect species (TFIIA-L). The alignment was created using JalView with standard ClustalX colour coding and conservation score calculation. c, d, Western blot analyses of Flag–Moonshiner co-immunoprecipitation from lysates of S2 cells transfected with indicated expression constructs (IN, input; UB, unbound; IP, immunoprecipitate; asterisk, signal from anti-Flag heavy chain).
Extended Data Figure 3 Moonshiner forms an alternative TFIIA–TRF2 complex enriched at piRNA clusters.
a, TRF2 isoform characterization by total wild-type ovary RNA-seq (top) and LAP–Moonshiner co-immunoprecipitation mass spectrometry (bottom). The identified TRF2 peptides show that Moonshiner is in complex only with the shorter TRF2 isoform. We therefore specifically investigated this isoform, also known as TRF2S, in the remainder of the study. b, c, Absolute peptide peak intensities for the main protein interactors identified in Fig. 2b, c. Peak area intensities are displayed as immunoprecipitation values subtracted that of the paired control immunoprecipitation experiment. On the basis of this, we conclude that TFIIA-S and TRF2 are robust Moonshiner interactors (supportive of an alternative TFIIA–TRF2 complex), while only a small fraction of Moonshiner is bound to Deadlock. Furthermore, the data show that, in ovaries, TRF2 interacts predominantly with canonical TFIIA, but also clearly with Moonshiner. Black dots represent individual biological replicate values. Orange bars show median values. d, Western blot analyses as Extended Data Fig. 2d, but addressing interaction with HA–TRF2 (lower bands probably represent TRF2 decay intermediates). e, Schematic of a developing Drosophila ovariole with germline cells in beige and somatic support cells in green. Confocal images were typically taken from egg chambers of stage 7 (highlighted by a dashed box). f, Whole egg chamber confocal image stained for DNA (DAPI; blue), LAP–Moonshiner (GFP auto-fluorescence; green), Rhino (magenta), and Deadlock (cyan). The circled nucleus is shown in Fig. 2d. g, Fluorescence images of nurse cell nuclei (depleted for indicated factors using sh-lines) indicating levels and localization of Moonshiner and Rhino (scale bar, 5 μm). h, Western blot showing levels of LAP–Moonshiner in ovaries where the indicated factors were depleted in the germline via sh-lines (ATP synthase serves as loading control).
Extended Data Figure 4 Moonshiner mutants reveal highly specific function at Rhino-bound piRNA clusters.
a, Schematic of the moonshiner frameshift alleles generated by CRISPR/Cas9. b, piRNA levels from ovaries with indicated genotype (relative to wild type) mapping uniquely to indicated piRNA clusters. c, Left: deregulation of steady-state transposon transcript levels (RNA-seq; sense only) in ovaries of the indicated mutants fly strains. Right: changes in corresponding piRNA levels (antisense only). The y axis values show log2(fold change) of transcripts per million values relative to wild type. Each bar represents one transposon consensus sequence (n = 73; shown are only transposons with minimum expression of RNA-seq transcripts per million > 5 in any library). Sorting of transposons in all panels is identical. The plotted values are available as figure source data. d, Rhino occupancy at indicated major piRNA clusters as well as all other Rhino-bound loci is shown as boxplot quantification (n = 1-kb windows analysed for each group) of Rhino ChIP-seq read coverage in the indicated genotypes. Boxplots are defined as in Fig. 3c; ***P < 0.0001 based on Mann–Whitney–Wilcoxon non-parametric tests. e, Genome browser panel showing read coverage at cluster80F of the data underlying the log2(fold change) tracks shown in Fig. 3b. Shown are RNA-seq (green), Pol II ChIP-seq (red), and ChIP-seq input samples (purple) generated from the indicated genotypes. f, RNA-seq transcripts per million values for canonical genes compared between control and moonshiner−/− (left) or rhino−/− (right); key genes related to Moonshiner biology are highlighted in orange. Abbreviation rSpearman denotes the Spearman correlation coefficient for each data set pair. g, Representative confocal images underlying the quantitative RNA FISH-based detection of piRNA precursors from cluster20A (Rhino-independent) and cluster42AB (Rhino-dependent) in germline nuclei of wild-type and moonshiner mutant ovaries. h, Example confocal images of germline nuclei stained for of DNA (DAPI) and nuclear pore complexes (wheat germ agglutinin, WGA-488), which were used to define the nuclear region in whole-nucleus Z-stack images acquired in parallel with images of RNA FISH signal. i, Example single-plane images of dual-channel RNA FISH quantification of whole-germline nuclei. RNA FISH signal within the nuclear regions (left, segmented using DAPI and WGA-488 signal) was used to define regions of interest (right), representing active sites of piRNA cluster transcription6. Signal in the foci was subsequently quantified for whole nuclei.
Extended Data Figure 5 Depletion of Moonshiner, TFIIA-S, or Trf2 activates transposon expression.
a, Percentages of eggs hatching into larvae laid by females expressing sh-constructs against the indicated target genes in their germline cells. Error bars, s.e.m. from four independent countings; n, the sum of counted eggs (see also figure source data). b, Ovarioles from flies expressing indicated piRNA sensors and indicated germline knockdown constructs (sh-lines) stained for β-galactosidase with X-gal. c, Top: deregulation of steady-state transposon transcript levels (sense only; compared with control ovaries) in ovaries expressing the indicated germline knockdown constructs. Each bar represents one transposon consensus sequence (n = 59; shown are only transposons with minimum expression of RNA-seq transcripts per million > 5 in any library). Bottom: changes in corresponding piRNA levels (antisense only). Sorting of transposons in all panels is identical. For plotted values see figure source data.
Extended Data Figure 6 piRNA production from Rhino-bound clusters requires Moonshiner, TFIIA-S, and Trf2.
a, UCSC genome browser panel showing piRNA profiles at cluster80F in ovaries expressing indicated germline knockdown constructs. b, Levels of piRNAs (relative to control) mapping uniquely to indicated Rhino-dependent or Rhino-independent piRNA clusters and derived from ovaries depleted of the indicated factors. c, d, UCSC genome browser panel showing cluster20A (c) or cluster42AB (d) piRNA levels from ovaries expressing indicated germline knockdown constructs.
Extended Data Figure 7 Characterization of Rhino-dependent, but Moonshiner-independent, piRNA production.
a, The log2(fold changes) in levels of piRNAs mapping antisense to transposons are plotted for rhino mutants versus moonshiner mutants. An outlier group of transposons for which the level of antisense piRNAs is decreased in rhino mutants but increased in moonshiner mutants is apparent, and elements enriched in cluster38C1/2 are highlighted in orange. The same transposons are shown as in Extended Data Fig. 4c (n = 73; transposon mRNAs analysed). b, Quantification of relative piRNA levels originating from cluster38C1 in ovaries from flies subjected to the indicated germline knockdowns. Percentages relative to control knockdowns were calculated with the total numbers of piRNA reads mapping uniquely to cluster38C1. c, Representative confocal images underlying the quantitative RNA FISH-based detection of piRNA precursors from cluster20A (Rhino-independent) and cluster38C1 (Rhino-dependent) in germline nuclei of wild-type and moonshiner mutant ovaries (scale bar, 5 μm). d, UCSC genome browser panel showing the most distal part of cluster42AB for which piRNA production dependency on the right flanking promoter was investigated by deletion of the promoter region. Shown are Pol II occupancy (red), Rhino occupancy (blue), and piRNA levels (black/grey). Flanking transcription units are shown in grey; light grey shading indicates the experimental promoter deletion.
Extended Data Figure 8 Moonshiner function can be bypassed by directly connecting Deadlock to Trf2.
a, Experimental scheme used to recruit GFP or TRF2 to DNA upstream of sequences of interest to test for stimulation of Luciferase transcription. Bar diagram shows fold changes in reporter activity upon tethering of TRF2 versus GFP to wild-type or mutant Histone 1 core promoter or to random piRNA cluster fragments (error bars, s.e.m.; n = 5 biological replicates; *P <0.05 based on two-tailed paired t-tests). b, Firefly luciferase values underlying the relative activities shown in a. Firefly luciferase activity was normalized to Renilla luciferase activity (transfection and viability control) upon tethering of TRF2 versus GFP to wild-type or mutant Histone 1 core promoter or to ten random piRNA cluster fragments (error bars, s.d. of five biological replicates each with six technical replicates. c, Confocal images showing localization of LAP–Moonshiner and Rhino in germline nuclei of ovaries depleted for indicated factors (scale bar, 5 μm). d, Western blot showing levels of LAP–Moonshiner in ovaries where the indicated factors were depleted in the germline via sh-lines (ATP synthase serves as loading control). e, Confocal images showing localization of germline-expressed LAP–TRF2 and endogenous Rhino in control ovaries (top) or in ovaries expressing the Deadlock–GFP-nanobody fusion protein (bottom) (scale bar, 5 μm). The TRF2 accumulations in wild-type nuclei do not overlap with Rhino foci and instead are reported to be TRF2 accumulations at the repetitive histone loci31. We note that TRF2 accumulation at Rhino foci is not visible in wild-type cells, most probably as the levels of this protein are too high to detect this local enrichment, which depends on Moonshiner (a protein expressed at only low levels). f, Representative images of DAPI-stained embryos (inverted monochromatic) assessed for progress of early embryogenesis. Left: two images of normal embryo development at the blastoderm stage (top) and at the extended germband stage (after gastrulation; bottom). Right: a typical moonshiner mutant embryo arrested early in development (no distinct nuclei are visible; the lower image displays the top image at increased brightness). g, Percentages of embryos with the indicated genotype displaying successful hatching. h, Relative levels of steady-state transposon mRNAs underlying the panel displayed in Fig. 5d. Bars show mean levels relative to those measured in moon−/− samples. Error bars, s.d. of three biological replicates. *P < 0.05 from two-tailed t-tests for difference to moonshiner full mutant samples. i, Levels of piRNAs mapping uniquely to the indicated clusters (grey, Rhino-independent; black, Rhino-dependent) in the indicated genotypes (values are normalized to the wild-type control levels). j, The log2(fold changes) in levels of piRNAs mapping antisense to transposons are plotted relative to levels in moonshiner mutants. The green boxes highlight the set of transposons for which mutation of moonshiner results in decreased antisense piRNAs (n = 111; transposons with fewer than 100 antisense piRNAs per million were removed from the analyses).
Extended Data Figure 9 Comparison of canonical enhancer-dependent and heterochromatin-dependent transcription activation pathways.
Schematic comparison of canonical enhancer-dependent transcription and transcription of small RNA source loci in Drosophila and Arabidopsis specified by chromatin marks. Canonical transcription initiation is driven by sequence-specific transcription factor binding to DNA motifs in accessible enhancer and promoter regions, which subsequently leads to positioning of TFIID/TBP onto core promoters (left). In contrast, while Moonshiner-mediated transcription also converges on recruitment of TFIID to DNA, this pathway exclusively utilizes the TBP paralogue TRF2. Furthermore, Moonshiner-mediated transcription gains locus specificity via recognition of heterochromatic histone marks through the HP1 protein Rhino, rather than through DNA motifs, thereby circumventing the transcriptional inhibition imposed by the compact state of heterochromatic DNA (middle). In plants, a conceptually similar pathway has evolved using an entirely different set of proteins (right). Here, the homeodomain protein SHH1 binds H3K9me histone marks and subsequently recruits the Pol IV variant RNA polymerase complex to transcribe small RNA precursors.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-3 and Supplementary Figure 1, the uncropped western blot images. (PDF 1590 kb)
Supplementary Table 1
This file contains Drosophila melanogaster fly strains. (XLSX 23 kb)
Supplementary Table 2
This table contains oligo sequences. (XLSX 24 kb)
Supplementary Table 3
This table contains a list of antibodies used in the study. (XLSX 9 kb)
Supplementary Table 4
This table contains a list of plasmids used in the study. (XLSX 9 kb)
Supplementary Table 5
This table contains Stellaris probe sequences. (XLSX 43 kb)
Supplementary Table 6
This table contains sequence accessions for Extended Data Figure 2b (XLSX 32 kb)
Rights and permissions
About this article
Cite this article
Andersen, P., Tirian, L., Vunjak, M. et al. A heterochromatin-dependent transcription machinery drives piRNA expression. Nature 549, 54–59 (2017). https://doi.org/10.1038/nature23482
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23482
This article is cited by
-
Critical appraisal of the piRNA-PIWI axis in cancer and cancer stem cells
Biomarker Research (2024)
-
The burgeoning importance of PIWI-interacting RNAs in cancer progression
Science China Life Sciences (2024)
-
The emerging role of the piRNA/PIWI complex in respiratory tract diseases
Respiratory Research (2023)
-
Modeling early germline immunization after horizontal transfer of transposable elements reveals internal piRNA cluster heterogeneity
BMC Biology (2023)
-
Themes and variations on piRNA-guided transposon control
Mobile DNA (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.