Abstract
In many prokaryotes, type III clustered regularly interspaced short palindromic repeat (CRISPR)–CRISPR-associated (Cas) systems detect and degrade invasive genetic elements by an RNA-guided, RNA-targeting multisubunit interference complex. The CRISPR-associated protein Csm6 additionally contributes to interference by functioning as a standalone RNase that degrades invader RNA transcripts, but the mechanism linking invader sensing to Csm6 activity is not understood. Here we show that Csm6 proteins are activated through a second messenger generated by the type III interference complex. Upon target RNA binding by the interference complex, its Cas10 subunit converts ATP into a cyclic oligoadenylate product, which allosterically activates Csm6 by binding to its CRISPR-associated Rossmann fold (CARF) domain. CARF domain mutations that abolish allosteric activation inhibit Csm6 activity in vivo, and mutations in the Cas10 Palm domain phenocopy loss of Csm6. Together, these results point to an unprecedented mechanism for regulation of CRISPR interference that bears striking conceptual similarity to oligoadenylate signalling in mammalian innate immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Mohanraju, P. et al. Diverse evolutionary roots and mechanistic variations of the CRISPR-Cas systems. Science 353, aad5147 (2016)
Marraffini, L. A. CRISPR–Cas immunity in prokaryotes. Nature 526, 55–61 (2015)
Sorek, R., Lawrence, C. M. & Wiedenheft, B. CRISPR-mediated adaptive immune systems in bacteria and archaea. Annu. Rev. Biochem. 82, 237–266 (2013)
Koonin, E. V., Makarova, K. S. & Zhang, F. Diversity, classification and evolution of CRISPR-Cas systems. Curr. Opin. Microbiol. 37, 67–78 (2017)
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015)
Samai, P. et al. Co-transcriptional DNA and RNA cleavage during Type III CRISPR-Cas immunity. Cell 161, 1164–1174 (2015)
Goldberg, G. W., Jiang, W., Bikard, D. & Marraffini, L. A. Conditional tolerance of temperate phages via transcription-dependent CRISPR–Cas targeting. Nature 514, 633–637 (2014)
Deng, L., Garrett, R. A., Shah, S. A., Peng, X. & She, Q. A novel interference mechanism by a type IIIB CRISPR-Cmr module in Sulfolobus. Mol. Microbiol. 87, 1088–1099 (2013)
Elmore, J. R. et al. Bipartite recognition of target RNAs activates DNA cleavage by the type III-B CRISPR-Cas system. Genes Dev. 30, 447–459 (2016)
Estrella, M. A., Kuo, F.-T. & Bailey, S. RNA-activated DNA cleavage by the type III-B CRISPR-Cas effector complex. Genes Dev. 30, 460–470 (2016)
Kazlauskiene, M., Tamulaitis, G., Kostiuk, G., Venclovas, Č. & Siksnys, V. Spatiotemporal control of type III-A CRISPR-Cas immunity: coupling DNA degradation with the target RNA recognition. Mol. Cell 62, 295–306 (2016)
Tamulaitis, G., Venclovas, Č. & Siksnys, V. Type III CRISPR-Cas immunity: major differences brushed aside. Trends Microbiol. 25, 49–61 (2017)
Makarova, K. S., Anantharaman, V., Grishin, N. V., Koonin, E. V. & Aravind, L. CARF and WYL domains: ligand-binding regulators of prokaryotic defense systems. Front. Genet. 5, 102 (2014)
Anantharaman, V., Makarova, K. S., Burroughs, A. M., Koonin, E. V. & Aravind, L. Comprehensive analysis of the HEPN superfamily: identification of novel roles in intra-genomic conflicts, defense, pathogenesis and RNA processing. Biol. Direct 8, 15 (2013)
Hatoum-Aslan, A., Maniv, I., Samai, P. & Marraffini, L. A. Genetic characterization of antiplasmid immunity through a type III-A CRISPR-Cas system. J. Bacteriol. 196, 310–317 (2014)
Jiang, W., Samai, P. & Marraffini, L. A. Degradation of phage transcripts by CRISPR-associated RNases enables type III CRISPR-Cas immunity. Cell 164, 710–721 (2016)
Niewoehner, O. & Jinek, M. Structural basis for the endoribonuclease activity of the type III-A CRISPR-associated protein Csm6. RNA 22, 318–329 (2016)
Sheppard, N. F., Glover, C. V. C., III, Terns, R. M. & Terns, M. P. The CRISPR-associated Csx1 protein of Pyrococcus furiosus is an adenosine-specific endoribonuclease. RNA 22, 216–224 (2016)
Staals, R. H. J. et al. RNA targeting by the type III-A CRISPR-Cas Csm complex of Thermus thermophilus. Mol. Cell 56, 518–530 (2014)
Tamulaitis, G. et al. Programmable RNA shredding by the type III-A CRISPR-Cas system of Streptococcus thermophilus. Mol. Cell 56, 506–517 (2014)
Hatoum-Aslan, A., Samai, P., Maniv, I., Jiang, W. & Marraffini, L. A. A ruler protein in a complex for antiviral defense determines the length of small interfering CRISPR RNAs. J. Biol. Chem. 288, 27888–27897 (2013)
Burroughs, A. M., Zhang, D., Schäffer, D. E., Iyer, L. M. & Aravind, L. Comparative genomic analyses reveal a vast, novel network of nucleotide-centric systems in biological conflicts, immunity and signaling. Nucleic Acids Res. 43, 10633–10654 (2015)
Yan, X., Guo, W. & Yuan, Y. A. Crystal structures of CRISPR-associated Csx3 reveal a manganese-dependent deadenylation exoribonuclease. RNA Biol. 12, 749–760 (2015)
Topuzlu, E. & Lawrence, C. M. Recognition of a pseudo-symmetric RNA tetranucleotide by Csx3, a new member of the CRISPR associated Rossmann fold superfamily. RNA Biol. 13, 254–257 (2016)
Carroll, S. S. et al. Activation of RNase L by 2′,5′-oligoadenylates. Kinetic characterization. J. Biol. Chem. 272, 19193–19198 (1997)
Prischi, F., Nowak, P. R., Carrara, M. & Ali, M. M. U. Phosphoregulation of Ire1 RNase splicing activity. Nat. Commun. 5, 3554 (2014)
Staals, R. H. J. et al. Structure and activity of the RNA-targeting Type III-B CRISPR-Cas complex of Thermus thermophilus. Mol. Cell 52, 135–145 (2013)
Hale, C. R., Cocozaki, A., Li, H., Terns, R. M. & Terns, M. P. Target RNA capture and cleavage by the Cmr type III-B CRISPR-Cas effector complex. Genes Dev. 28, 2432–2443 (2014)
Makarova, K. S. et al. Evolution and classification of the CRISPR–Cas systems. Nat. Rev. Microbiol. 9, 467–477 (2011)
Osawa, T., Inanaga, H. & Numata, T. Crystal structure of the Cmr2-Cmr3 subcomplex in the CRISPR-Cas RNA silencing effector complex. J. Mol. Biol. 425, 3811–3823 (2013)
Kalia, D. et al. Nucleotide, c-di-GMP, c-di-AMP, cGMP, cAMP, (p)ppGpp signaling in bacteria and implications in pathogenesis. Chem. Soc. Rev. 42, 305–341 (2013)
Hornung, V., Hartmann, R., Ablasser, A. & Hopfner, K.-P. OAS proteins and cGAS: unifying concepts in sensing and responding to cytosolic nucleic acids. Nat. Rev. Immunol. 14, 521–528 (2014)
East-Seletsky, A. et al. Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature 538, 270–273 (2016)
He, F., Vestergaard, G., Peng, W., She, Q. & Peng, X. CRISPR-Cas type I-A Cascade complex couples viral infection surveillance to host transcriptional regulation in the dependence of Csa3b. Nucleic Acids Res. 45, 1902–1913 (2017)
Lintner, N. G. et al. The structure of the CRISPR-associated protein Csa3 provides insight into the regulation of the CRISPR/Cas system. J. Mol. Biol. 405, 939–955 (2011)
Geertsma, E. R. FX cloning: a simple and robust high-throughput cloning method for protein expression. Methods Mol. Biol. 1116, 153–164 (2014)
Kreiswirth, B. N. et al. The toxic shock syndrome exotoxin structural gene is not detectably transmitted by a prophage. Nature 305, 709–712 (1983)
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D 60, 2256–2268 (2004)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
Gouet, P., Courcelle, E., Stuart, D. I. & Métoz, F. ESPript: analysis of multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999)
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015)
Acknowledgements
We thank members of the Jinek and Marraffini laboratories for discussions and comments on the manuscript. We thank D. Swarts and P. Sledz for technical assistance and sharing reagents. We thank U. Manzau for technical assistance during A4>P synthesis and purification. We thank the Service for Mass Spectrometry of ETH Zurich for support with MS analysis. This study was supported by a Swiss National Science Foundation project grant to M.J. (SNSF 31003A_149393) and by funding from the Swiss National Competence Center for Research (NCCR) ‘RNA & Disease’ (to M.J. and J.H.). M.J. is an International Research Scholar of the Howard Hughes Medical Institute and Vallee Scholar of the Bert L & N Kuggie Vallee Foundation. C.G.-D. was supported by a Long-Term Fellowship from the European Molecular Biology Organization. J.T.R. was supported by a Boehringer Ingelheim Fonds PhD fellowship. L.A.M. is supported by the Rita Allen Scholars Program, a Burroughs Wellcome Fund PATH award, a National Institutes of Health Director’s New Innovator Award (1DP2AI104556-01), and a Howard Hughes Medical Institute-Simons Faculty Scholar Award.
Author information
Authors and Affiliations
Contributions
O.N., C.G.-D., and M.J. conceived the study. O.N., C.G.-D., J.T.R., L.M., and M.J. designed experiments. O.N. expressed and purified recombinant Csm6 proteins, performed oligoA activation and ATPase assays, and performed enzymatic probing of the cyclic oligoA product. C.G.-D. expressed and purified recombinant EiCsm(1–5) complexes, performed oligoA activation assays, and assisted with LC–MS analysis. J.T.R. performed phage infection assays under supervision of L.M. C.B. synthesized 2′,3′-cyclic phosphate-terminated nucleotides and performed LC–MS analysis under supervision of J.H. F.S. synthesized 2′,3′-cyclic phosphate-terminated A4 nucleotide and advised on nucleotide chemistry. L.B. performed additional LC–MS analyses of Csm6 activators. O.N., C.G.-D., and M.J. wrote the manuscript, with input from the remaining authors.
Corresponding author
Ethics declarations
Competing interests
F.S. is an employee of BIOLOG Life Science Institute GmbH, which markets synthetic nucleotides. The other authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S. Bailey, P. Kranzusch and R. Staals for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 2 Activation of Csm6 ribonucleases by oligoadenylates.
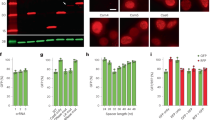
a, TtCsm6 RNase activity assay in the presence of 3′-hydroxylated oligoadenylates ranging from A3 to A10. b, SDS–PAGE analysis of purified WT and mutant Csm6 proteins from T. thermophilus (TtCsm6), M. thermautotrophicus (MtCsm), and E. italicus (EiCsm6). c, d, MtCsm6 and EiCsm6 RNase activity assays in the presence of 3′-hydroxylated oligoadenylates ranging from A3 to A10. e, MtCsm6 RNase activity assay in the presence of oligoadenylates carrying 3′-hydroxyl (-OH) or 2′,3′-cyclic phosphate (>P) groups. f, TtCsm6 and EiCsm6 RNase activity assay in the presence of A4>P and A6>P. All data points represent the mean of three replicates ± s.e.m.
Extended Data Figure 3 Activation of EiCsm6 by A6>P.
a, EiCsm6 RNase assay in the presence of varying concentrations of A6>P. b, The log(dose)-versus-response curve and EC50 derived from assays in a. Errors are indicated as 95% confidence interval. All data points represent the mean of three replicates ± s.e.m.
Extended Data Figure 4 Conservation and structure of the CARF domain motif.
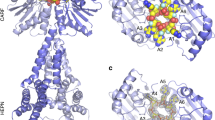
a, Multiple sequence alignment of the CARF domain motifs in Csm6. The polypeptide sequence of EiCsm6 (denoted with an arrow) was used for a BLAST search against the National Center for Biotechnology Information (NCBI) non-redundant protein sequences database and 19 best scoring hits with >95% query coverage were selected. The multiple sequence alignment was performed using Clustal Omega39 and visualized using ESPript 3 (ref. 40). Green asterisk denotes the conserved glutamine residue mutated to abrogate allosteric activation of EiCsm6. b, Zoom-in view of a structural model of the dimeric interface of the CARF domains in the EiCsm6 dimer. The model was generated using Phyre2 (ref. 41) on the basis of the TtCsm6 crystal structure (PDB 5FSH).
Extended Data Figure 5 Purification and crRNA-guided RNA cleavage by the EiCsm(1–5) complex.
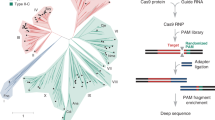
a, E. italicus DSM15952 contains a CRISPR–Cas locus with nine cas genes followed by a repeat-spacer array. For heterologous expression of the EiCsm(1–5) complex, the genomic DNA fragment spanning genes csm1–csm6 and cas6 and the first two repeat-spacer units in the CRISPR array was cloned into the expression plasmid. b, SDS–PAGE analysis of purified wild-type and mutant EiCsm(1–5) complexes. c, Denaturing gel of the products of a target RNase cleavage assay performed using a fluorophore-labelled target ssRNA substrate in the presence of WT or mutant EiCsm(1–5) complexes.
Extended Data Figure 6 The EiCsm(1–5) complex generates cyclic hexaadenylate.
a, Denaturing 20% polyacrylamide gel indicating time-resolved [α-32P]ATP oligomerization by the EiCsm(1–5) complex. b, Mass spectra of LC–MS peaks from the analysis of the products generated by the EiCsm(1–5)–dCsm3D32A and EiCsm(1–5)–dCas10Palm/dCsm3D32A complexes shown in Fig. 4c. The mass spectrum of synthetic A6>P standard is shown for comparison.
Extended Data Figure 7 LC–MS analysis of purified EiCsm(1–5) product.
a, c, e, LC–MS analysis of the HPLC-purified activator product generated by the EiCsm(1–5) complex as well as an A6>P standard and purified product spiked with the A6>P standard. b, d, f, Mass spectra of the peaks indicated in a, c, e.
Extended Data Figure 8 The EiCsm(1–5) complex generates cyclic hexaadenylate.
a, Left: LC–MS analysis of the nuclease S1 digested activator. Right: mass spectrum profile of the peak obtained after S1 digest. b, EiCsm6 RNase assay in the presence of varying concentrations of c-A6. c, Overlay of EC50 values obtained for A6>P and c-A6. Errors are indicated as 95% confidence interval and data points represent mean of three replicates ± s.e.m.
Extended Data Figure 9 The EiCsm(1–5) complex selectively incorporates adenosines into the activator.
a, Stimulation of EiCsm6 RNase activity by products generated by the EiCsm(1–5) complex in the presence of various ribonucleotide triphosphate combinations. b, LC–MS analysis of indicated samples from a. The elution profile (left) and mass spectrum (right) of the product remain unaltered upon addition of all four ribonucleotide triphosphates, indicating ATP selectivity of the EiCsm(1–5) complex.
Extended Data Figure 10 Proposed mechanism of cyclic oligoadenylate formation.
The Cas10 Palm domain initially catalyses 5′–3′ phosphodiester bond formation by a nucleophilic attack of the 3′-hydroxyl group of one ATP substrate molecule onto the α-phosphate group of a second ATP molecule, yielding a 5′-triphosphorylated dinucleotide and inorganic pyrophosphate (PPi). The dinucleotide is extended by addition of successive AMP moieties to the 3′ end to generate a linear oligonucleotide. Final cyclization involves the 3′-hydroxyl group attacking the α-phosphate group of the 5′-triphosphate moiety in the linear oligoadenylate intermediate. Notably, all reaction steps use the same catalytic mechanism and thus could be performed by a single catalytic active site in the Palm domain of Cas10.
Supplementary information
Supplementary Figure
This file contains the uncropped gels. (PDF 941 kb)
Supplementary Data
This file contains Supplementary Table 1 (XLSX 11 kb)
Supplementary Data
This file contains Supplementary Table 2. (XLSX 11 kb)
Rights and permissions
About this article
Cite this article
Niewoehner, O., Garcia-Doval, C., Rostøl, J. et al. Type III CRISPR–Cas systems produce cyclic oligoadenylate second messengers. Nature 548, 543–548 (2017). https://doi.org/10.1038/nature23467
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23467
This article is cited by
-
Activation of Thoeris antiviral system via SIR2 effector filament assembly
Nature (2024)
-
The CRISPR effector Cam1 mediates membrane depolarization for phage defence
Nature (2024)
-
Conservation and similarity of bacterial and eukaryotic innate immunity
Nature Reviews Microbiology (2024)
-
RNA targeting and cleavage by the type III-Dv CRISPR effector complex
Nature Communications (2024)
-
Antiviral type III CRISPR signalling via conjugation of ATP and SAM
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.