Abstract
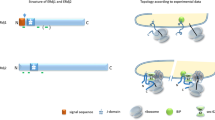
Misfolded endoplasmic reticulum proteins are retro-translocated through the membrane into the cytosol, where they are poly-ubiquitinated, extracted from the membrane, and degraded by the proteasome1,2,3,4—a pathway termed endoplasmic reticulum-associated protein degradation (ERAD). Proteins with misfolded domains in the endoplasmic reticulum lumen or membrane are discarded through the ERAD-L and ERAD-M pathways, respectively. In Saccharomyces cerevisiae, both pathways require the ubiquitin ligase Hrd1, a multi-spanning membrane protein with a cytosolic RING finger domain5,6. Hrd1 is the crucial membrane component for retro-translocation7,8, but it is unclear whether it forms a protein-conducting channel. Here we present a cryo-electron microscopy structure of S. cerevisiae Hrd1 in complex with its endoplasmic reticulum luminal binding partner, Hrd3. Hrd1 forms a dimer within the membrane with one or two Hrd3 molecules associated at its luminal side. Each Hrd1 molecule has eight transmembrane segments, five of which form an aqueous cavity extending from the cytosol almost to the endoplasmic reticulum lumen, while a segment of the neighbouring Hrd1 molecule forms a lateral seal. The aqueous cavity and lateral gate are reminiscent of features of protein-conducting conduits that facilitate polypeptide movement in the opposite direction—from the cytosol into or across membranes9,10,11. Our results suggest that Hrd1 forms a retro-translocation channel for the movement of misfolded polypeptides through the endoplasmic reticulum membrane.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bays, N. W., Wilhovsky, S. K., Goradia, A., Hodgkiss-Harlow, K. & Hampton, R. Y. HRD4/NPL4 is required for the proteasomal processing of ubiquitinated ER proteins. Mol. Biol. Cell 12, 4114–4128 (2001)
Ye, Y., Meyer, H. H. & Rapoport, T. A. The AAA ATPase Cdc48/p97 and its partners transport proteins from the ER into the cytosol. Nature 414, 652–656 (2001)
Jarosch, E. et al. Protein dislocation from the ER requires polyubiquitination and the AAA-ATPase Cdc48. Nat. Cell Biol. 4, 134–139 (2002)
Rabinovich, E., Kerem, A., Fröhlich, K. U., Diamant, N. & Bar-Nun, S. AAA-ATPase p97/Cdc48p, a cytosolic chaperone required for endoplasmic reticulum-associated protein degradation. Mol. Cell. Biol. 22, 626–634 (2002)
Vashist, S. & Ng, D. T. W. Misfolded proteins are sorted by a sequential checkpoint mechanism of ER quality control. J. Cell Biol. 165, 41–52 (2004)
Carvalho, P., Goder, V. & Rapoport, T. A. Distinct ubiquitin-ligase complexes define convergent pathways for the degradation of ER proteins. Cell 126, 361–373 (2006)
Carvalho, P., Stanley, A. M. & Rapoport, T. A. Retrotranslocation of a misfolded luminal ER protein by the ubiquitin-ligase Hrd1p. Cell 143, 579–591 (2010)
Baldridge, R. D. & Rapoport, T. A. Autoubiquitination of the Hrd1 ligase triggers protein retrotranslocation in ERAD. Cell 166, 394–407 (2016)
Park, E. & Rapoport, T. A. Mechanisms of Sec61/SecY-mediated protein translocation across membranes. Annu. Rev. Biophys. 41, 21–40 (2012)
Kumazaki, K. et al. Structural basis of Sec-independent membrane protein insertion by YidC. Nature 509, 516–520 (2014)
Dalbey, R. E. & Kuhn, A. Membrane insertases are present in all three domains of life. Structure 23, 1559–1560 (2015)
Bordallo, J., Plemper, R. K., Finger, A. & Wolf, D. H. Der3p/Hrd1p is required for endoplasmic reticulum-associated degradation of misfolded lumenal and integral membrane proteins. Mol. Biol. Cell 9, 209–222 (1998)
Bays, N. W., Gardner, R. G., Seelig, L. P., Joazeiro, C. A. & Hampton, R. Y. Hrd1p/Der3p is a membrane-anchored ubiquitin ligase required for ER-associated degradation. Nat. Cell Biol. 3, 24–29 (2001)
Horn, S. C. et al. Usa1 functions as a scaffold of the HRD-ubiquitin ligase. Mol. Cell 36, 782–793 (2009)
Gardner, R. G. et al. Endoplasmic reticulum degradation requires lumen to cytosol signaling. J. Cell Biol. 151, 69–82 (2000)
Denic, V., Quan, E. M. & Weissman, J. S. A luminal surveillance complex that selects misfolded glycoproteins for ER-associated degradation. Cell 126, 349–359 (2006)
Gauss, R., Jarosch, E., Sommer, T. & Hirsch, C. A complex of Yos9p and the HRD ligase integrates endoplasmic reticulum quality control into the degradation machinery. Nat. Cell Biol. 8, 849–854 (2006)
Stein, A., Ruggiano, A., Carvalho, P. & Rapoport, T. A. Key steps in ERAD of luminal ER proteins reconstituted with purified components. Cell 158, 1375–1388 (2014)
Bai, X.-C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. W. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, 1485 (2015)
Ovchinnikov, S. et al. Protein structure determination using metagenome sequence data. Science 355, 294–298 (2017)
Song, Y. et al. High-resolution comparative modeling with RosettaCM. Structure 21, 1735–1742 (2013)
Deak, P. M. & Wolf, D. H. Membrane topology and function of Der3/Hrd1p as a ubiquitin-protein ligase (E3) involved in endoplasmic reticulum degradation. J. Biol. Chem. 276, 10663–10669 (2001)
Jeong, H. et al. Crystal structure of SEL1L: Insight into the roles of SLR motifs in ERAD pathway. Sci. Rep. 6, 20261 (2016)
Mehnert, M., Sommer, T. & Jarosch, E. Der1 promotes movement of misfolded proteins through the endoplasmic reticulum membrane. Nat. Cell Biol. 16, 77–86 (2014)
Sato, B. K., Schulz, D., Do, P. H. & Hampton, R. Y. Misfolded membrane proteins are specifically recognized by the transmembrane domain of the Hrd1p ubiquitin ligase. Mol. Cell 34, 212–222 (2009)
Gonen, T. & Walz, T. The structure of aquaporins. Q. Rev. Biophys. 39, 361–396 (2006)
Dutzler, R., Campbell, E. B., Cadene, M., Chait, B. T. & MacKinnon, R. X-ray structure of a ClC chloride channel at 3.0 A reveals the molecular basis of anion selectivity. Nature 415, 287–294 (2002)
McCoy, J. G., Levin, E. J. & Zhou, M. Structural insight into the PTS sugar transporter EIIC. Biochim. Biophys. Acta 1850, 577–585 (2015)
Van den Berg, B. et al. X-ray structure of a protein-conducting channel. Nature 427, 36–44 (2004)
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005)
Zheng, S. Q., Palovcak, E., Armache, J.-P., Verba, K. A., Cheng, Y. & Agard, D. A. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017)
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Ru, H. et al. Molecular mechanism of V(D)J recombination from synaptic RAG1–RAG2 complex structures. Cell 163, 1138–1152 (2015)
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Su, H . et al. GeRelion: GPU-enhanced parallel implementation of single particle cryo-EM image processing. Preprint available at http://www.biorxiv.org/content/early/2016/09/19/075887 (2016)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014)
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013)
Ovchinnikov, S., Kamisetty, H. & Baker, D. Robust and accurate prediction of residue-residue interactions across protein interfaces using evolutionary information. eLife 3, e02030 (2014)
Remmert, M., Biegert, A., Hauser, A. & Söding, J. HHblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat. Methods 9, 173–175 (2011)
Nordberg, H. et al. The genome portal of the Department of Energy Joint Genome Institute: 2014 updates. Nucleic Acids Res. 42, D26–D31 (2014)
Tsirigos, K. D., Peters, C., Shu, N., Käll, L. & Elofsson, A. The TOPCONS web server for consensus prediction of membrane protein topology and signal peptides. Nucleic Acids Res. 43, W401–W407 (2015)
DiMaio, F., Tyka, M. D., Baker, M. L., Chiu, W. & Baker, D. Refinement of protein structures into low-resolution density maps using rosetta. J. Mol. Biol. 392, 181–190 (2009)
Ovchinnikov, S. et al. Large-scale determination of previously unsolved protein structures using evolutionary information. eLife 4, e09248 (2015)
Finn, R. D. et al. HMMER web server: 2015 update. Nucleic Acids Res. 43, W30–W38 (2015)
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013)
Wang, S., Sun, S., Li, Z., Zhang, R. & Xu, J. Accurate de novo prediction of protein contact map by ultra-deep learning model. PLOS Comput. Biol. 13, e1005324 (2017)
Wang, S., Li, W., Zhang, R., Liu, S. & Xu, J. CoinFold: a web server for protein contact prediction and contact-assisted protein folding. Nucleic Acids Res. 44, W361–W366 (2016)
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009)
Gemmill, R. M. et al. The hereditary renal cell carcinoma 3;8 translocation fuses FHIT to a patched-related gene, TRC8. Proc. Natl Acad. Sci. USA 95, 9572–9577 (1998)
The PyMOL Molecular Graphics System, Version 1.8 (Schrödinger, LLC, 2017)
Acknowledgements
We thank Z. Yu, R. Huang and C. Hong at the HHMI Janelia Cryo-EM Facility for help in microscope operation and data collection, Z. Li for technical support at our in-house EM facility, and S. Andrei Anghel, for help with experiments, and X. Wu, P. Carvalho and T. Walz for comments on the manuscript. This work was supported by the European Molecular Biology Organization (EMBO LTF #1437-2012) to S. S., by the European Research Council (ERC) under the Horizon 2020 research and innovation program (grant # 677770) to A.S., by the National Key Research and Development Program of China (grant # 2016YFB1000101) to D.L., and by NIGMS Award R01GM052586 to T.A.R. T.A.R. is a Howard Hughes Medical Institute Investigator.
Author information
Authors and Affiliations
Contributions
S.S. prepared protein and built the models, W.M. and M.L. collected and analysed EM data, A.S. designed the construct and performed sequence alignments, S.O. and R.P. and their advisors F.D. and D.B. built models based on evolutionary couplings and energy minimization, M.G.C. helped with EM data collection, H.S. and D.L. developed DSS in GeRelion, T.A.R. and M.L. supervised the project and T.A.R. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Purification and cryo-EM of the Hrd1–Hrd3 complex.
a, In the last purification step, the Hrd1–Hrd3 complex was subjected to gel filtration on a Superdex 200 10/300GL Increase column. Shown is the UV elution profile. b, SDS–PAGE gel of the peak fraction, stained with Coomassie blue. For gel source data, see Supplementary Fig. 1. c, Representative cryo-EM image with a few particles marked by circles. A total of 5,361 images were collected. d, 2D class averages of cryo-EM particles.
Extended Data Figure 2 3D classification and refinement procedure for the Hrd1–Hrd3 complex.
Views parallel to the membrane of 3D reconstructions are shown, and percentages of the particles in each class indicated. Three different classes selected from the first round of 3D classification are circled with dashed lines in different colours and were used for further analysis, as indicated by correspondingly coloured arrows. The four final maps are labelled A–D and shown with their resolutions and particle numbers. Maps C and D were used for model building. To obtain the best 3D classification focusing on the Hrd1 dimer, we compared DSS and conventional signal subtraction. Only with DSS was a particle class obtained that resulted in a reconstruction showing clear densities for the TM7–TM8 and TM5–TM6 loops of Hrd1.
Extended Data Figure 3 Single particle cryo-EM analysis of Hrd1–Hrd3 complexes.
a, Density maps were generated for the Hrd1–Hrd3 dimer, the Hrd1 dimer with one associated Hrd3 molecule, the Hrd1 dimer, and Hrd3 (Extended Data Fig. 2). Left, maps in a side view, coloured according to local resolution; middle, gold-standard Fourier shell correlation (FSC) curve (blue) with indicated resolution at FSC = 0.143; right, Euler angle distribution in two different views. Dashed grey FCS curves (bottom two rows) were calculated between the atomic model and the corresponding final cryo-EM map. b, The density map for the Hrd1–Hrd3 dimer was filtered to a resolution of 6.8 Å without amplitude modification and is displayed at two different isosurface levels. At a low level (left), the weak amphipol density is visible and encloses the density of Hrd1 dimer. The amphipathic helix of Hrd3 associates only with the outer surface of the amphipol density. At a high isosurface level (middle and right), the density for the amphipathic helix is clearly connected with that of the preceding Sel1 domains and well separated from that of TM1 and TM2 of the nearby Hrd1 molecule. The region in the dashed black box (middle) is displayed as a sectional view on the right.
Extended Data Figure 4 Examples of the fit of the model and density maps.
a, Amino acids for which side chain density was observed are indicated in side and top views of the Hrd1 model. b, Central interface between the Hrd1 molecules. H79 and F83 from the two Hrd1 molecules (orange and green) probably form cation-pi interactions. c, TM3 and TM8 of Hrd1. d, Densities for the transmembrane segments of Hrd1. Amino acids with clear side chain densities are indicated. e, Selected areas in Hrd3. Blue, N-terminal domain; yellow, central domain; purple, C-terminal domain.
Extended Data Figure 6 Sequence similarities between Hrd1 and other multi-spanning ubiquitin ligases.
Multiple sequence alignment showing amino acid conservation in TM3–TM8 of Hrd1, TM3–TM8 of gp78 (also called AMFR), and TM9–TM14 of TRC8 (also called RNF139) and RNF145. Left, Uniprot codes for individual sequences. Numbers after Uniprot codes indicate the depicted amino acid range. Black bars above the sequences indicate the locations of the most C-terminal six transmembrane segments of human gp78 (top) and human TRC8 (bottom) as predicted by TOPCONS. Below bars, amino acid numbering for Hrd1p from S. cerevisiae is given. Colouring was edited in JalView according to conservation of hydrophobicity49. Residues highlighted in green and with green dots are conserved among Hrd1 and gp78 molecules and are involved in the interaction of TM2, TM3 and TM4 on the cytosolic side of the membrane (Extended Data Fig. 7c). Species abbreviations in Uniprot codes: YEAST S. cerevisiae, USTMA Ustilago maydis, CAPO3 Capsaspora owczarzaki, MONBE Monosiga brevicollis, AMPQE Amphimedon queenslandica, SCHMA Schistosoma mansoni, STRPU Strongylocentrotus purpuratus, CAEEL Caenorhabditis elegans, DROME Drosophila melanogaster, DANRE Danio rerio, THETB Thecamonas trahens, PLABS Plasmodiophora brassicae, ECTSI Ectocarpus siliculosus, PLAF7 Plasmodium falciparum, PARTE Paramecium tetraurelia, GUITH Guillardia theta, GALSU Galdieria sulphuraria, OSTLU Ostreococcus lucimarinus, ARATH Arabidopsis thaliana, LEIMA Leishmania major, DICDI Dictyostelium discoideum, DAPPU Daphnia pulex, CIOIN Ciona intestinalis, SELML Selaginella moellendorffii, STRMM Strigamia maritima.
Extended Data Figure 7 Structural homology between the ubiquitin ligases Hrd1 and TRC8.
a, Domain organization of human gp78 and TRC8. Black bars indicate the positions of transmembrane segments as predicted by TOPCONS. The positions of the VIM (VCP-interacting motif), CUE (coupling of ubiquitin to ER degradation) and RING finger domains were taken from Uniprot entries Q9UKV5 and Q8WU17. The sterol-sensing domain (blue line) in TRC8 is positioned according to ref. 50. Regions with homology to Hrd1 are indicated by red lines. b, Structural model of human TRC8 (residues 323–516) as generated by RaptorX-Contact44,45. The structure of S. cerevisiae Hrd1 is shown in brown and the model for TRC in dark blue (residues 345–516). TM9 of TRC8 does not align well with TM3 of Hrd1 and is therefore shown in light blue (residues 323–344). Shown are views from the cytosol (left) and views parallel to the membrane either towards the hydrophilic cavity or from the back (middle and right, respectively). The structure of Hrd1 from S. cerevisiae was aligned with the TRC8 model obtained from RaptorX-Contact44,45 using the command cealign within Pymol51. c, Views of Hrd1 from the cytosol (left) and the side (right), with residues conserved in the ubiquitin ligases Hrd1, gp78, TRC8 and RNF145 shown in orange, and residues conserved in Hrd1 and gp78 in purple.
Extended Data Figure 8 Potential Hrd3-binding surfaces for substrate and partners.
a, Groove at the inner surface of Hrd3 (circled), located at the junction of the N-terminal (residues 187–199), middle (residues 507–521) and C-terminal (residues 539–560, 593–595) domains. Left, space-filling model with conserved amino acids shown in orange to red (scores 7–9 in Consurf). Hrd1 is shown in ribbon presentation. Right, the same view with Hrd3 in cartoon presentation. Helices containing the conserved residues are shown as ribbons. Some residues are labelled for orientation. b, Left, three conserved surface grooves on the outer surface of Hrd3 (circled), with conserved residues coloured as in a. Right, the same view with helices containing conserved residues in ribbon presentation.
Extended Data Figure 9 Sequence conservation of the amphipathic helix of Hrd3.
Consensus sequence of the amphipathic helix of Hrd3, generated with Weblogo 3.0. The size of the letters correlates with their conservation. Black, blue and green letters indicate hydrophobic, hydrophilic and other residues, respectively. Below, an alignment of the amphipathic helix of S. cerevisiae Hrd3 with the consensus sequence (Cons). The blue area indicates the amphipathic helix, with residues in red forming its hydrophobic face.
Supplementary information
Supplementary Figure
This file contains the uncropped scan of SDS-PAGE used in Extended Data Figure 1b. (PDF 662 kb)
Rights and permissions
About this article
Cite this article
Schoebel, S., Mi, W., Stein, A. et al. Cryo-EM structure of the protein-conducting ERAD channel Hrd1 in complex with Hrd3. Nature 548, 352–355 (2017). https://doi.org/10.1038/nature23314
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23314
This article is cited by
-
SEL1L-HRD1 interaction is required to form a functional HRD1 ERAD complex
Nature Communications (2024)
-
Direct observation of autoubiquitination for an integral membrane ubiquitin ligase in ERAD
Nature Communications (2024)
-
Casting Light on the Janus-Faced HMG-CoA Reductase Degradation Protein 1: A Comprehensive Review of Its Dualistic Impact on Apoptosis in Various Diseases
Molecular Neurobiology (2024)
-
Mechanisms of substrate processing during ER-associated protein degradation
Nature Reviews Molecular Cell Biology (2023)
-
TRIM59 guards ER proteostasis and prevents Bortezomib-mediated colorectal cancer (CRC) cells’ killing
Investigational New Drugs (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.