Abstract
Concomitant activation of the Wnt pathway and suppression of Mapk signalling by two small molecule inhibitors (2i) in the presence of leukaemia inhibitory factor (LIF) (hereafter termed 2i/L) induces a naive state in mouse embryonic stem (ES) cells that resembles the inner cell mass (ICM) of the pre-implantation embryo1. Since the ICM exists only transiently in vivo, it remains unclear how sustained propagation of naive ES cells in vitro affects their stability and functionality. Here we show that prolonged culture of male mouse ES cells in 2i/L results in irreversible epigenetic and genomic changes that impair their developmental potential. Furthermore, we find that female ES cells cultured in conventional serum plus LIF medium phenocopy male ES cells cultured in 2i/L. Mechanistically, we demonstrate that the inhibition of Mek1/2 is predominantly responsible for these effects, in part through the downregulation of DNA methyltransferases and their cofactors. Finally, we show that replacement of the Mek1/2 inhibitor with a Src inhibitor preserves the epigenetic and genomic integrity as well as the developmental potential of ES cells. Taken together, our data suggest that, although short-term suppression of Mek1/2 in ES cells helps to maintain an ICM-like epigenetic state, prolonged suppression results in irreversible changes that compromise their developmental potential.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Sequence Read Archive
Change history
09 August 2017
The name of the PKC inhibitor in the main text and Methods was corrected.
References
Ying, Q.-L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008)
Raggioli, A., Junghans, D., Rudloff, S. & Kemler, R. β-catenin is vital for the integrity of mouse embryonic stem cells. PLoS One 9, e86691 (2014)
Chen, H. et al. Erk signaling is indispensable for genomic stability and self-renewal of mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 112, E5936–E5943 (2015)
Chen, R. Z., Pettersson, U., Beard, C., Jackson-Grusby, L. & Jaenisch, R. DNA hypomethylation leads to elevated mutation rates. Nature 395, 89–93 (1998)
Gaudet, F. et al. Induction of tumors in mice by genomic hypomethylation. Science 300, 489–492 (2003)
Habibi, E. et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 13, 360–369 (2013)
Ficz, G. et al. FGF signaling inhibition in ES cells drives rapid genome-wide demethylation to the epigenetic ground state of pluripotency. Cell Stem Cell 13, 351–359 (2013)
Leitch, H. G. et al. Naive pluripotency is associated with global DNA hypomethylation. Nat. Struct. Mol. Biol. 20, 311–316 (2013)
Schulz, E. G. et al. The two active X chromosomes in female ES cells block exit from the pluripotent state by modulating the ES cell signaling network. Cell Stem Cell 14, 203–216 (2014)
Zvetkova, I. et al. Global hypomethylation of the genome in XX embryonic stem cells. Nat. Genet. 37, 1274–1279 (2005)
Choi, J. et al. DUSP9 modulates DNA hypomethylation in female mouse pluripotent stem cells. Cell Stem Cell 20, 706–719.e7 (2017)
Hackett, J. A. et al. Synergistic mechanisms of DNA demethylation during transition to ground-state pluripotency. Stem Cell Rep. 1, 518–531 (2013)
Wu, T. et al. Histone variant H2A.X deposition pattern serves as a functional epigenetic mark for distinguishing the developmental potentials of iPSCs. Cell Stem Cell 15, 281–294 (2014)
Marks, H. et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 149, 590–604 (2012)
Plasschaert, R. N. & Bartolomei, M. S. Genomic imprinting in development, growth, behavior and stem cells. Development 141, 1805–1813 (2014)
Nagy, A., Rossant, J., Nagy, R., Abramow-Newerly, W. & Roder, J. C. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc. Natl Acad. Sci. USA 90, 8424–8428 (1993)
Celeste, A. et al. H2AX haploinsufficiency modifies genomic stability and tumor susceptibility. Cell 114, 371–383 (2003)
Wray, J. et al. Inhibition of glycogen synthase kinase-3 alleviates Tcf3 repression of the pluripotency network and increases embryonic stem cell resistance to differentiation. Nat. Cell Biol. 13, 838–845 (2011)
Tamm, C., Pijuan Galitó, S. & Annerén, C. A comparative study of protocols for mouse embryonic stem cell culturing. PLoS One 8, e81156 (2013)
Meyenn, von, F. et al. Impairment of DNA methylation maintenance is the main cause of global demethylation in naive embryonic stem cells. Mol. Cell 62, 848–861 (2016)
Dutta, D. et al. Self-renewal versus lineage commitment of embryonic stem cells: protein kinase C signaling shifts the balance. Stem Cells 29, 618–628 (2011)
Shimizu, T. et al. Dual inhibition of Src and GSK3 maintains mouse embryonic stem cells, whose differentiation is mechanically regulated by Src signaling. Stem Cells 30, 1394–1404 (2012)
Kolodziejczyk, A. A. et al. Single cell RNA-sequencing of pluripotent states unlocks modular transcriptional variation. Cell Stem Cell 17, 471–485 (2015)
Theunissen, T. W. et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 15, 471–487 (2014)
Takashima, Y. et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 158, 1254–1269 (2014)
Pastor, W. A. et al. Naive human pluripotent cells feature a methylation landscape devoid of blastocyst or germline memory. Cell Stem Cell 18, 323–329 (2016)
Choi, J ., Huebner, A. J . & Hochedlinger, K. Derivation and long-term propagation of mouse embryonic stem cells. Protoc. Exch. http://dx.doi.org/10.1038/protex.2017.070 (2017)
Stadtfeld, M., Maherali, N., Borkent, M. & Hochedlinger, K. A reprogrammable mouse strain from gene-targeted embryonic stem cells. Nat. Methods 7, 53–55 (2010)
Kaneda, M. et al. Essential role for de novo DNA methyltransferase Dnmt3a in paternal and maternal imprinting. Nature 429, 900–903 (2004)
Dodge, J. E. et al. Inactivation of Dnmt3b in mouse embryonic fibroblasts results in DNA hypomethylation, chromosomal instability, and spontaneous immortalization. J. Biol. Chem. 280, 17986–17991 (2005)
Boyle, P. et al. Gel-free multiplexed reduced representation bisulfite sequencing for large-scale DNA methylation profiling. Genome Biol. 13, R92 (2012)
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008)
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Anders, S. et al. Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nat. Protocols 8, 1765–1786 (2013)
Dobin, A . et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011)
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011)
Treff, N. R. et al. Next generation sequencing-based comprehensive chromosome screening in mouse polar bodies, oocytes, and embryos. Biol. Reprod. 94, 76 (2016)
Xiao, A. et al. WSTF regulates the H2A.X DNA damage response via a novel tyrosine kinase activity. Nature 457, 57–62 (2009)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Song, Q. & Smith, A. D. Identifying dispersed epigenomic domains from ChIP-Seq data. Bioinformatics 27, 870–871 (2011)
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010)
Eggan, K. et al. Hybrid vigor, fetal overgrowth, and viability of mice derived by nuclear cloning and tetraploid embryo complementation. Proc. Natl Acad. Sci. USA 98, 6209–6214 (2001)
Huttlin, E. L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010)
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994)
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007)
Acknowledgements
We thank members of the Hochedlinger laboratory for suggestions as well as L. Barrett for critical reading of the manuscript. Additionally, we thank A. Galvin, M. Gesner and M. Handley at the Massachusetts General Hospital flow cytometry core. We also thank A. L. Hawkins, S. Wang, A. Aggarwal and C. C. Morton at Brigham and Woman’s Hospital CytoGenomics Core Laboratory for karyotyping analysis. We are grateful to K. Shioda from the Shioda laboratory for technical support for genome sequencing. We are also grateful to W. Yu at The Sidney Kimmel Cancer Center Microarray Core Facility at Johns Hopkins University for aCGH analysis, who was supported by NIH grant P30 CA006973 entitled Regional Oncology Research Center. We thank T. Theunissen and R. Jaenisch for sharing human OCT4–eGFP ES cell lines and T. Graf for providing the Rex1–GFP ES cells. A.J.H. is supported by an American Cancer Society–New England Division–Ellison Foundation Postdoctoral Fellowship (PF-15-130-01-DDC). B.D.S. is supported by an EMBO long-term Fellowship (ALTF 1143-2015). T.S. is supported by NIH grant R21ES024861. A.Me. is a New York Stem Cell Foundation Robertson Investigator and supported by NIH grant 1P50HG006193 and the New York Stem Cell Foundation. K.H. was supported by funds from the MGH, HHMI, NIH (R01 HD058013-06) and the Gerald and Darlene Jordan Chair in Regenerative Medicine.
Author information
Authors and Affiliations
Contributions
J. Choi and A.J.H. performed the bulk of the cell culture, teratoma assays, dot blot and flow cytometry analyses; J. Choi, A.J.H. and S.-Y.K. performed the blastocyst injections; J. Choi, A.J.H. and J. Charron generated the Mek1/2 double-knockout iPS cell lines; R.M.W. generated the XGXT ES cell line; J.S. and H.K. generated the Dnmt triple-knockout ES cell line; J. Choi, A.J.H. and J.B. conducted the western blot analysis; J.B., C.M.R. and S.G. performed the proteomics analysis; B.D.S. performed experiments related to human ES cell culture; J.O. and T.S. performed the genome sequencing analyses and analysed the RNA-seq data of interspecies iPS cell lines; K.C., H.G., A.Mo., A.G. and A.Me. performed the RRBS analyses; A.S. and R.I.S. performed the RNA-seq analyses; K.L. and A.X. conducted the H2A.X ChIP–seq analysis.; J. Choi, A.J.H. and K.H. designed the experiments, interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks T. Zwaka and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Effects of different culture conditions on DNA methylation in ES cells.
a–c, Violin plots showing global methylation levels of ES cells at p10 (a), p20 (b) and after being placed back into S/L at p23 and p30 (c) by RRBS. Hypomethylated Dnmt1 knockout (KO) ES cells were included as control. White dots denote median value. d, Heat map of DNA methylation levels at SINE elements in ES cells. e, Median methylation levels of different genomic elements in the indicated conditions. f, Box plot showing methylation levels of ICRs cultured in each condition. Thick horizontal lines represent median values. Box shows the interquartile range of the data. Whiskers extend to the most extreme data point that is no more than 1.5 times the interquartile range. Outliers beyond this range are shown as points. g, ICR methylation levels after being re-exposed to S/L for the indicated passages. h, RNA sequencing was performed on a F1 Mus musculus × Mus spretus stem-cell line cultured in 2i/L for 6 passages. Normalized read counts of mRNA transcripts expressed from the paternal or maternal alleles are shown based on multiple allele-discriminating single nucleotide polymorphisms (SNPs) for each gene (6 or more SNPs per gene). Statistical analysis (two-tailed t-tests) for allelic biases of the SNPs is shown. i, Karyotyping results from two female ES cell lines cultured in S/L for 10 passages.
Extended Data Figure 2 Effects of prolonged 2i/L culture on H2A.X deposition in ES cells.
a, Heat map showing global H2A.X deposition loss relative to male ES cells cultured in S/L. Three ES cell lines per condition (male S/L, male 2i/L, female S/L) at p10 and two ES cell lines (male 2i/L to S/L) at p30 were analysed (see Fig. 1a). b, Bar graph quantifying the percentage of H2A.X-depleted regions relative to male ES cells cultured in S/L. Each bar represents median values of biological replicates of samples in a. c, Relative H2A.X deposition on a representative chromosome. Relative losses (green bars) or gains (grey bars) of H2A.X are mapped to their location on chromosome 2. d, Overlap of devoid H2A.X regions common to both female ES cells cultured in S/L and male cells cultured in 2i/L. e, H2A.X-devoid regions present in the overlap identified in d were significantly enriched for genes involved in the listed developmental pathways. The GREAT bioinformatic database was used to bin the genes identified into transcriptional networks.
Extended Data Figure 3 Developmental potential of ES cells cultured in S/L and 2i/L.
a, Representative image of a chimaeric pup derived from female ES cells cultured in S/L (left) compared to a pup obtained through natural mating (right). Body weight of each pup was shown below. b, Representative image of a low-grade adult chimaera with patches of agouti hairs generated using female ES cells in S/L (white arrow). Her agouti germline offspring (red arrowhead) was generated by crossing her to a C57B6 wild-type mouse. c, Representative image of all-ES cell adult mice generated from male ES cells cultured in S/L for 10 passages. d, Bar graphs showing percentages of transferred 4n blastocysts that survived to birth (full-term), established regular breathing or survived past 5 weeks (adult) using male ES cells grown in either S/L (blue bars) or 2i/L (red bars) until p20. The number of animals obtained per total number of transferred embryos is shown. e, Representative images of all-ES cell neonates produced from male ES cells cultured in S/L (left) and 2i/L (middle) at p20. Representative image of all-ES cell adult mice generated from ES cells cultured in S/L at p20 (right).
Extended Data Figure 4 Potential of male ES cells cultured in 2i/L to produce teratomas.
a, Karyotype analysis of male ES cell lines grown in 2i/L at p10. b, Representative images of teratomas produced with three male ES cell lines grown in 2i/L at p20 with depiction of germ layer differentiation where detectable.
Extended Data Figure 5 Effects of prolonged 2i/L culture on chromosomal stability in ES cells.
a, aCGH analysis of the J-35 ES cell line cultured in S/L or 2i/L condition for the indicated passage numbers. b, Karyotype analysis of Rex1–GFP ES cell line grown in 2i/L for 8 passages after receiving the line. Red boxes indicate abnormal chromosomes detected. c, Karyotype analysis of J35 ES cell line grown in 2i/L in the absence of a layer of MEF feeder cells for 16 passages (p20). Red boxes indicate abnormal chromosomes detected. d, aCGH analysis on DNA isolated from a newborn pup generated by 4n blastocyst injection using ES cells cultured in S/L or 2i/L at 20 passages. e, Combined dendrogram and heat map depicting ICR methylation levels in ES cells (n = 3 biological replicates) and keratinocytes using RRBS analysis. ‘2i/L keratinocytes’ were explanted from chimaeric adult mice derived from male 2i/L ES cell p10 and purified by drug selection (G418).
Extended Data Figure 6 Characterization of the XGXT ES cell line.
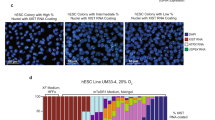
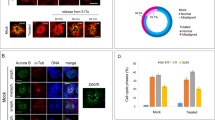
a, Fluorescence microscopic images of XGXT ES cells. GFP/tdTomato double-positive cells (yellow cells in merge 1 and 2) indicate two active X chromosomes while GFP+Tomato− colony depicts cells that lost one of the X chromosomes. b, GFP/tdTomato double-positive XGXT ES cells were sorted at passage 5 and plated in S/L (p6) before measuring the percentage of double-positive and single-positive cells at p8, p10, p13 and p16 using flow cytometry. c, Karyotype analysis of undifferentiated, XGO (GFP-positive) or XTO (tdTomato-positive) ES cells (left) and GFP/tdTomato double-positive XGXT ES cells (right). GFP/tdTomato double-positive XGXT ES cells were sorted at passage 5 and maintained in S/L for 9 passages before analysis. This result confirms that the progressive loss of the GFP or TdTomato signal was due to the loss of an X chromosome, and not X-chromosome inactivation due to differentiation.
Extended Data Figure 7 Consequences of PD0325901 and CHIR99021 treatment on DNA methylation.
a, Global methylation levels of were analysed by 5mC dot blot analysis for male and female ES cell lines cultured in the indicated conditions for 6 passages. b, Median methylation levels at the indicated genetic elements following inhibition or loss of Mek1/2 or Gsk3α/β, respectively. c, Violin plots showing global methylation levels of male and female ES cells cultured in S/L for 6 passages and then cultured for an additional 3 passages in S/L supplemented with PD0325901 (PD). White boxes indicate median methylation levels. d, Heat map and dendrogram of ICR methylation levels in male and female ES cells shown in c.
Extended Data Figure 8 Chromosomal aberrations that occur in 2i/L-cultured ES cells are in part due to Mek1/2 inhibition.
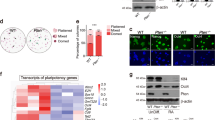
a, Quantification of relative RNA and protein levels of pluripotency-related genes between male ES cells in S/L (n = 2 technical replicates) and male ES cells in S/L + PD (n = 2 technical replicates). b, Quantification of relative RNA and protein levels of genes that facilitate DNA methylation between male ES cells in S/L (n = 2 technical replicates) and male ES cells in S/L + PD (n = 2 technical replicates) (see Fig. 4d). c, d, Karyotype analysis of male ES cells cultured in S/L for 4 passages and subsequently cultured in either S/L + PD (c) or S/L + CHIR (d) for additional 16 passages. Red boxes indicate abnormal chromosomes detected. e, Western blot analysis of a Dnmt conditional triple-knockout (cTKO) ES cell line for Dnmt1, Dnmt3a and Dnmt3b after treatment with either 4-OHT or ethanol (EtOH) for one passage. f, Karyotype analysis of a Dnmt conditional triple-knockout ES cell line that was treated with either 4-OHT or ethanol for one passage and subsequently cultured for 15 passages in S/L.
Extended Data Figure 9 Characterization of alternative culture conditions for ES cells.
a, Representative images of XY ES cells cultured in the indicated conditions. b, Heat map and dendrogram of global gene expression levels in male ES cells cultured in the indicated conditions. c, Principal component analysis of ES cells cultured in S/L, 2i/L, a2i/L or PKCi/L using a previously published set of known pluripotency and differentiation genes23. Two male ES cell lines maintained in S/L for 4 passages were switched to each condition and cultured for additional 6 passages (final passage 10). d, Heat map and dendrogram showing expression levels of pluripotency and developmental genes23 in male ES cell lines cultured in the indicated conditions. e, Relative expression levels of transcripts associated with naive pluripotency in d. f, Violin plots of global methylation levels in the indicated ES cell lines. White dots, median value. g, Heat map and dendrogram of ICR methylation levels in the indicated ES cells lines. h, Flow-cytometric analysis of GFP and TdTomato after culturing XGXT ES cells in S/L, S/L+PD and a2i/L for the indicated passages. i, Agouti germline offspring (red arrows), obtained from across between an all-ES cell male generated from p10 a2i/L-cultured ES cells and a C57B6 female. j, k, Karyotype analysis of male ES cells cultured in a2i/L for 6 or 16 passages (j), or PKCi/L for 6 or 16 passages (k). Red boxes indicate abnormal chromosomes detected.
Extended Data Figure 10 Replacement of MEK1/2 inhibitor with SRC inhibitor using naive human ES cells.
a, Representative image of human ES cells carrying a naive-specific OCT4–eGFP reporter (ΔPE-OCT4-eGFP)24 cultured in the presence of small molecule inhibitors of the ROCK, MEK1/2, GSK3 and PKC kinases (t2iLGöY)25. Bright field image (left) and GFP expression (right). b, Representative image of ΔPE-OCT4-eGFP human ES cells cultured in the presence of small molecule inhibitors of the ROCK, SRC, GSK3 and PKC kinases (a2iLGöY). Bright field image (left) and GFP expression (right). c, Flow-cytometric analysis of eGFP expression in ΔPE-Oct4GFP human ES cells cultured in t2iLGöY or a2iLGöY.
Supplementary information
Supplementary Figure
This file contains the uncropped western blots corresponding to the labelled figures. (PDF 696 kb)
Supplementary Tables
This file contains Supplementary Tables 1-3. (XLSX 50 kb)
Rights and permissions
About this article
Cite this article
Choi, J., Huebner, A., Clement, K. et al. Prolonged Mek1/2 suppression impairs the developmental potential of embryonic stem cells. Nature 548, 219–223 (2017). https://doi.org/10.1038/nature23274
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23274
This article is cited by
-
Molecular versatility during pluripotency progression
Nature Communications (2023)
-
Enhanced cultivation of chicken primordial germ cells
Scientific Reports (2023)
-
Generation of functional oocytes from male mice in vitro
Nature (2023)
-
Young LINE-1 transposon 5′ UTRs marked by elongation factor ELL3 function as enhancers to regulate naïve pluripotency in embryonic stem cells
Nature Cell Biology (2023)
-
RNA stability controlled by m6A methylation contributes to X-to-autosome dosage compensation in mammals
Nature Structural & Molecular Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.