Abstract
In contrast to the DNA-based viruses in prokaryotes, the emergence of eukaryotes provided the necessary compartmentalization and membranous environment for RNA viruses to flourish, creating the need for an RNA-targeting antiviral system1,2. Present day eukaryotes employ at least two main defence strategies that emerged as a result of this viral shift, namely antiviral RNA interference and the interferon system2. Here we demonstrate that Drosha and related RNase III ribonucleases from all three domains of life also elicit a unique RNA-targeting antiviral activity. Systemic evolution of ligands by exponential enrichment of this class of proteins illustrates the recognition of unbranched RNA stem loops. Biochemical analyses reveal that, in this context, Drosha functions as an antiviral clamp, conferring steric hindrance on the RNA-dependent RNA polymerases of diverse positive-stranded RNA viruses. We present evidence for cytoplasmic translocation of RNase III nucleases in response to virus in diverse eukaryotes including plants, arthropods, fish, and mammals. These data implicate RNase III recognition of viral RNA as an antiviral defence that is independent of, and possibly predates, other known eukaryotic antiviral systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Koonin, E. V., Wolf, Y. I., Nagasaki, K. & Dolja, V. V. The Big Bang of picorna-like virus evolution antedates the radiation of eukaryotic supergroups. Nat. Rev. Microbiol. 6, 925–939 (2008)
tenOever, B. R. The evolution of antiviral defense systems. Cell Host Microbe 19, 142–149 (2016)
tenOever, B. R. RNA viruses and the host microRNA machinery. Nat. Rev. Microbiol. 11, 169–180 (2013)
Cerutti, H. & Casas-Mollano, J. A. On the origin and functions of RNA-mediated silencing: from protists to man. Curr. Genet. 50, 81–99 (2006)
Obbard, D. J., Jiggins, F. M., Halligan, D. L. & Little, T. J. Natural selection drives extremely rapid evolution in antiviral RNAi genes. Curr. Biol. 16, 580–585 (2006)
Shapiro, J. S., Langlois, R. A., Pham, A. M. & Tenoever, B. R. Evidence for a cytoplasmic microprocessor of pri-miRNAs. RNA 18, 1338–1346 (2012)
Shapiro, J. S., Varble, A., Pham, A. M. & Tenoever, B. R. Noncanonical cytoplasmic processing of viral microRNAs. RNA 16, 2068–2074 (2010)
Rouha, H., Thurner, C. & Mandl, C. W. Functional microRNA generated from a cytoplasmic RNA virus. Nucleic Acids Res. 38, 8328–8337 (2010)
Bogerd, H. P., Whisnant, A. W., Kennedy, E. M., Flores, O. & Cullen, B. R. Derivation and characterization of Dicer- and microRNA-deficient human cells. RNA 20, 923–937 (2014)
Han, J. et al. The Drosha–DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004)
Triboulet, R., Chang, H. M., Lapierre, R. J. & Gregory, R. I. Post-transcriptional control of DGCR8 expression by the Microprocessor. RNA 15, 1005–1011 (2009)
Tang, X., Zhang, Y., Tucker, L. & Ramratnam, B. Phosphorylation of the RNase III enzyme Drosha at serine300 or serine302 is required for its nuclear localization. Nucleic Acids Res. 38, 6610–6619 (2010)
Court, D. L. et al. RNase III: genetics and function; structure and mechanism. Annu. Rev. Genet. 47, 405–431 (2013)
Shirako, Y., Strauss, E. G. & Strauss, J. H. Modification of the 5′ terminus of Sindbis virus genomic RNA allows nsP4 RNA polymerases with nonaromatic amino acids at the N terminus to function in RNA replication. J. Virol. 77, 2301–2309 (2003)
Tsetsarkin, K. A., Liu, G., Shen, K. & Pletnev, A. G. Kissing-loop interaction between 5′ and 3′ ends of tick-borne Langat virus genome ‘bridges the gap’ between mosquito- and tick-borne flaviviruses in mechanisms of viral RNA cyclization: applications for virus attenuation and vaccine development. Nucleic Acids Res. 44, 3330–3350 (2016)
Bick, M. J. et al. Expression of the zinc-finger antiviral protein inhibits alphavirus replication. J. Virol. 77, 11555–11562 (2003)
Rupp, J. C., Jundt, N. & Hardy, R. W. Requirement for the amino-terminal domain of sindbis virus nsP4 during virus infection. J. Virol. 85, 3449–3460 (2011)
Moy, R. H. et al. Stem-loop recognition by DDX17 facilitates miRNA processing and antiviral defense. Cell 158, 764–777 (2014)
Shapiro, J. S. et al. Drosha as an interferon-independent antiviral factor. Proc. Natl Acad. Sci. USA 111, 7108–7113 (2014)
Sabin, L. R. et al. Dicer-2 processes diverse viral RNA species. PLoS ONE 8, e55458 (2013)
Sarkar, D., Desalle, R. & Fisher, P. B. Evolution of MDA-5/RIG-I-dependent innate immunity: independent evolution by domain grafting. Proc. Natl Acad. Sci. USA 105, 17040–17045 (2008)
Maillard, P. V. et al. Inactivation of the type I interferon pathway reveals long double-stranded RNA-mediated RNA interference in mammalian cells. EMBO J. 35, 2505–2518 (2016)
Seo, G. J. et al. Reciprocal inhibition between intracellular antiviral signaling and the RNAi machinery in mammalian cells. Cell Host Microbe 14, 435–445 (2013)
Varble, A. et al. An in vivo RNAi screening approach to identify host determinants of virus replication. Cell Host Microbe 14, 346–356 (2013)
Cheng, E. H., Levine, B., Boise, L. H., Thompson, C. B. & Hardwick, J. M. Bax-independent inhibition of apoptosis by Bcl-XL. Nature 379, 554–556 (1996)
Zhang, F. & Simon, A. E. A novel procedure for the localization of viral RNAs in protoplasts and whole plants. Plant J. 35, 665–673 (2003)
Perez, J. T. et al. MicroRNA-mediated species-specific attenuation of influenza A virus. Nat. Biotechnol. 27, 572–576 (2009)
Cherry, S. & Perrimon, N. Entry is a rate-limiting step for viral infection in a Drosophila melanogaster model of pathogenesis. Nat. Immunol. 5, 81–87 (2004)
Chong, M. M., Rasmussen, J. P., Rudensky, A. Y. & Littman, D. R. The RNAseIII enzyme Drosha is critical in T cells for preventing lethal inflammatory disease. J. Exp. Med. 205, 2005–2017 (2008)
Ossovskaya, V., Lim, S. T., Ota, N., Schlaepfer, D. D. & Ilic, D. FAK nuclear export signal sequences. FEBS Lett. 582, 2402–2406 (2008)
Pall, G. S. & Hamilton, A. J. Improved northern blot method for enhanced detection of small RNA. Nat. Protocols 3, 1077–1084 (2008)
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011)
Sedano, C. D. & Sarnow, P. Hepatitis C virus subverts liver-specific miR-122 to protect the viral genome from exoribonuclease Xrn2. Cell Host Microbe 16, 257–264 (2014)
Manley, J. L. SELEX to identify protein-binding sites on RNA. Cold Spring Harb. Protoc. 2013, 156–163 (2013)
Rio, D. C. Electrophoretic mobility shift assays for RNA-protein complexes. Cold Spring Harb. Protoc. 2014, 435–440 (2014)
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003)
Hardy, R. W. & Rice, C. M. Requirements at the 3′ end of the sindbis virus genome for efficient synthesis of minus-strand RNA. J. Virol. 79, 4630–4639 (2005)
McCormack, J. C. & Simon, A. E. in Current Protocols in Microbiology Ch. 16, Unit 16D.1 (Wiley, 2006)
Shaffer, S. M., Wu, M. T., Levesque, M. J. & Raj, A. Turbo FISH: a method for rapid single molecule RNA FISH. PLoS ONE 8, e75120 (2013)
Acknowledgements
We thank J. K. Lim and A. G. Pletnev for Langat virus reagents, B. Lee for Sendai virus reagents, R. W. Hardy for SINV reagents, B. R. Cullen for NoDice cells, K. K. Conzelmann for BSR-T7 cells, and B. Ramratnam for pEGFP–Drosha. Recombinant IFN-β was provided by the National Institute of Health’s Biodefense and Emerging Infections Research Resources Repository (HuIFN-β, NR-3080). This material is based upon work supported in part by the Burroughs Wellcome Fund, which provides support for both S.C. and B.R.T. S.C. is also supported by the National Institute of Allergy and Infectious Diseases (NIAID) (R01A1074951). J.P.L. is supported by the DIM Malinf, Conseil Regional d’Ile-de-France. L.C.A. is partly supported by the American Heart Association (15PRE24930012). A.E.S. is supported by National Science Foundation (MCB-1411836) and NIAID (R21AI117882). J.M. is supported by National Institute of General Medicine (F32 GM119235). B.R.T. is also supported by NIAID (R01AI110575).
Author information
Authors and Affiliations
Contributions
L.C.A. and S.S. designed and conducted experiments. M.P. performed SELEX. J.M., L.C.A., and A.E.S. were responsible for plant data. L.S. and S.C. generated the Drosophila data. J.V.S. generated the RNaseIII−/− cells. J.P.L. and L.C.A. performed the zebrafish work. D.S. and D.B.M. were responsible for all bioinformatics. B.R.T., L.C.A., and S.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks B. Cullen, L. Maraffini and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Characterization of RNaseIII−/− cells.
a, Sequence alignment of genetic alterations in the two alleles encoding Drosha in RNaseIII−/− cells. The deletion and insertion result in a frameshift and early stop codon. b, The ten most abundant miRNAs in each condition, parental HEK293T (WT) or RNaseIII−/− either mock-treated or SINV-infected for 24 h as determined by Illumina small RNA deep sequencing. c, Quantitative PCR (qPCR) analysis of DGCR8 mRNA levels in mock-treated and SINV-infected (MOI = 0.1, 8 h.p.i.) NoDice and RNaseIII−/− cells; error bars, s.d. from three independent experiments; all conditions had P < 0.05 by Student’s t-test. d, Transcriptome profiling and correlation analyses of NoDice cells at baseline and RNaseIII−/− cells transfected with GFP-tagged human Drosha for 72 h. Graph depicts data from two biological replicates per condition.
Extended Data Figure 2 Drosha depletion does not alter the response to IFN-I.
a, qPCR analysis of IFIT1 mRNA levels in NoDice and RNaseIII−/− fibroblasts treated with IFN-β (100 U ml−1, 8 h); error bars, s.d. from three independent experiments; all conditions had P < 0.05 by Student’s t-test. b, Western blot of NoDice and RNaseIII−/− fibroblasts infected with SINV for 1 h before administration of indicated amounts of IFN-β for 24 h. c, d, Northern blot (c) and western blot (d) of primary Rnasenf/f ear-derived fibroblasts treated with indicated AdVs for 2–3 days and then treated with either 100 U IFN-β for 6 h (c) or infected with SINV for 24 h (d). e, mRNA sequencing of total RNA from samples in c. Heatmap depicts known mouse ISGs and IFN downregulated genes with a log2(fold change) greater than 1, as defined by the Interferome database (www.interferome.org).
Extended Data Figure 3 Characterization of Drosha-2A cells.
a, Immunofluorescence of human fibroblasts stably expressing GFP-tagged Drosha-WT or Drosha-2A (S300A/S302A). b, The indicated Flag-tagged proteins were immunoprecipitated from whole-cell extracts (WCE) and incubated at 37 °C with in vitro transcribed genome of SIN124. Production of pre-miR-124 was determined by small RNA northern blot. c, Indicated cell types were infected with SINV at an MOI of 0.001 and viral titres were determined at 16 h.p.i. Shown is the average and s.d. of three independent experiments, with P < 0.05 as determined using a two-tailed Student’s t-test.
Extended Data Figure 4 Drosha-RB-RIIIDmut recognizes stem-loop structures in SINV RNA.
a, Immunoprecipitation of exogenously expressed Flag-tagged proteins. Shown is protein expression in the whole-cell extract and after immunoprecipitation (IP) with a Flag-specific antibody. b, Cells were transfected with Flag-tagged SeV-N or Drosha-RBmut and infected at 36 h.p.t. with SINV at an MOI of 3. At 8 h.p.i., Flag-tagged proteins were immunoprecipitated and bound RNA was isolated to perform qPCR. Graph shows SINV RNA levels relative to input and normalized to tubulin. The average of three independent experiments is shown. Error bars, s.d.; *P < 0.05 using a one-tailed Student’s t-test. c, Prediction of the structure of the 5′ 200 nucleotides of the SINV genome using RNAfold. d, EMSA was performed with the indicated immunoprecipitated proteins and radio-labelled in vitro transcribed RNA comprising the 5′ 200 nucleotides of the SINV genome. Unbound genome is indicated as ‘Free RNA’.
Extended Data Figure 5 Using virus engineering to discern Drosha’s antiviral mechanism.
a, Schematic of the SINV replicon encoding Gaussia luciferase in place of the structural polyprotein used in Fig. 3e–g. b, Schematic of the SINV temperature-sensitive mutant (SIN-RdRpts). Star denotes ts point mutant. c, NoDice and RNaseIII−/− cells were infected with virus depicted in b, at an MOI of 10 and incubated at 40 °C, a temperature at which the mutant viral RdRp is completely inactive. Levels of genomic (g) SINV RNA were determined by qPCR at the indicated times after infection. Data are representative of two independent experiments where each condition was done in triplicate; error bars, s.d. d, Schematic of the SINV encoding firefly luciferase in the nsP3 region and an inactive RdRp (SIN-nsP3Luc) e, Graph depicts levels of in vitro translation of firefly luciferase produced from virus in d, in the presence of membrane fractions from control or Drosha-2A. The data shown are the average of three independent experiments; error bars, s.d.
Extended Data Figure 6 Localization of and miRNA production from cytoplasmic viruses in diverse eukaryotes.
a, Zebrafish embryos were inoculated with SINV for 24 h and then analysed by FISH using a probe complementary to the capsid region of the genome. b, qPCR analysis of zebrafish embryos treated with the indicated morpholinos for 2 days (n = 4); error bars, s.d. c, A. thaliana protoplasts were mock-treated or TCV-infected for 40 h and then analysed by FISH using a Cy3-labelled probe complementary to bases 1210–1259 of the TCV genome. d, Quantification of mature miR-124 production from recombinant TCV was performed using the TaqMan miRNA assay on RNA from Fig. 4b. All samples were normalized to endogenous snoR66. Quantifications of each sample were performed in triplicate; error bars, s.d. from two biological replicates.
Extended Data Figure 7 The impact of diverse RNase III members on virus infection.
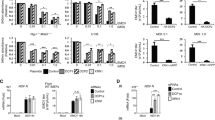
a, Schematic depicting core domains of human Drosha, C-terminal region of C. intestinalis Drosha, or full-length RNase III of S. pombe, M. maripaludis, and S. pyogenes. Domains depicted include Proline-rich (P-rich), arginine–serine-rich (RS-rich), conserved central domain (CED), RNaseIII domain (RIIID), and double-stranded RNA binding domain (dsRBD). b, Western blots from BSR-T7 cells, co-transfected with the indicated RNase-III-expression plasmids and SeV rescue plasmids encoding SeV-GFP genome, SeV-N, SeV-P, and SeV-L genes. RNase III expression was determined at 48 h.p.t. and virus replication at 72 h.p.t. c, Western blot of DL1 cells treated with indicated dsRNA for 3 days and subsequently infected with SINV (MOI = 1) for 96 h. d, HEK293T (WT) or RNaseIII−/− cells were infected with SINV for 24 h. Graphs depict the number of SINV reads mapping to indicated positions along the viral genomes from the small RNA deep sequencing performed in Extended Data Fig. 1b.
Supplementary information
Supplementary Information
This file contains the uncropped blots and Supplementary Tables 1-2. (PDF 3967 kb)
Rights and permissions
About this article
Cite this article
Aguado, L., Schmid, S., May, J. et al. RNase III nucleases from diverse kingdoms serve as antiviral effectors. Nature 547, 114–117 (2017). https://doi.org/10.1038/nature22990
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22990
This article is cited by
-
Inhibition of RNA-binding proteins with small molecules
Nature Reviews Chemistry (2020)
-
Study of the role of Mg2+ in dsRNA processing mechanism by bacterial RNase III through QM/MM simulations
JBIC Journal of Biological Inorganic Chemistry (2020)
-
A-to-I editing of Malacoherpesviridae RNAs supports the antiviral role of ADAR1 in mollusks
BMC Evolutionary Biology (2019)
-
Dgcr8 knockout approaches to understand microRNA functions in vitro and in vivo
Cellular and Molecular Life Sciences (2019)
-
A billion-year arms race against viruses shaped our evolution
Nature (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.