Abstract
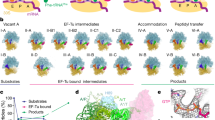
Gene translation depends on accurate decoding of mRNA, the structural mechanism of which remains poorly understood. Ribosomes decode mRNA codons by selecting cognate aminoacyl-tRNAs delivered by elongation factor Tu (EF-Tu). Here we present high-resolution structural ensembles of ribosomes with cognate or near-cognate aminoacyl-tRNAs delivered by EF-Tu. Both cognate and near-cognate tRNA anticodons explore the aminoacyl-tRNA-binding site (A site) of an open 30S subunit, while inactive EF-Tu is separated from the 50S subunit. A transient conformation of decoding-centre nucleotide G530 stabilizes the cognate codon–anticodon helix, initiating step-wise ‘latching’ of the decoding centre. The resulting closure of the 30S subunit docks EF-Tu at the sarcin–ricin loop of the 50S subunit, activating EF-Tu for GTP hydrolysis and enabling accommodation of the aminoacyl-tRNA. By contrast, near-cognate complexes fail to induce the G530 latch, thus favouring open 30S pre-accommodation intermediates with inactive EF-Tu. This work reveals long-sought structural differences between the pre-accommodation of cognate and near-cognate tRNAs that elucidate the mechanism of accurate decoding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ogle, J. M., Murphy, F. V., IV, Tarry, M. J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002)
Rodnina, M. V., Pape, T., Fricke, R., Kuhn, L. & Wintermeyer, W. Initial binding of the elongation factor Tu·GTP·aminoacyl-tRNA complex preceding codon recognition on the ribosome. J. Biol. Chem. 271, 646–652 (1996)
Pape, T., Wintermeyer, W. & Rodnina, M. V. Conformational switch in the decoding region of 16S rRNA during aminoacyl-tRNA selection on the ribosome. Nat. Struct. Mol. Biol. 7, 104–107 (2000)
Pape, T., Wintermeyer, W. & Rodnina, M. Induced fit in initial selection and proofreading of aminoacyl-tRNA on the ribosome. EMBO J. 18, 3800–3807 (1999)
Gromadski, K. B. & Rodnina, M. V. Kinetic determinants of high-fidelity tRNA discrimination on the ribosome. Mol. Cell 13, 191–200 (2004)
Gromadski, K. B., Daviter, T. & Rodnina, M. V. A uniform response to mismatches in codon–anticodon complexes ensures ribosomal fidelity. Mol. Cell 21, 369–377 (2006)
Zhang, J., Ieong, K.-W., Johansson, M. & Ehrenberg, M. Accuracy of initial codon selection by aminoacyl-tRNAs on the mRNA-programmed bacterial ribosome. Proc. Natl Acad. Sci. USA 112, 9602–9607 (2015)
Blanchard, S. C., Gonzalez, R. L., Kim, H. D., Chu, S. & Puglisi, J. D. tRNA selection and kinetic proofreading in translation. Nat. Struct. Mol. Biol. 11, 1008–1014 (2004)
Ieong, K.-W., Uzun, Ü., Selmer, M. & Ehrenberg, M. Two proofreading steps amplify the accuracy of genetic code translation. Proc. Natl Acad. Sci. USA 113, 13744–13749 (2016)
Voorhees, R. M. & Ramakrishnan, V. Structural basis of the translational elongation cycle. Annu. Rev. Biochem. 82, 203–236 (2013)
Maracci, C. & Rodnina, M. V. Review: translational GTPases. Biopolymers 105, 463–475 (2016)
Pape, T., Wintermeyer, W. & Rodnina, M. V. Complete kinetic mechanism of elongation factor Tu-dependent binding of aminoacyl-tRNA to the A site of the E. coli ribosome. EMBO J. 17, 7490–7497 (1998)
Rodnina, M. V., Fricke, R., Kuhn, L. & Wintermeyer, W. Codon-dependent conformational change of elongation factor Tu preceding GTP hydrolysis on the ribosome. EMBO J. 14, 2613–2619 (1995)
Schmeing, T. M. et al. The crystal structure of the ribosome bound to EF-Tu and aminoacyl-tRNA. Science 326, 688–694 (2009)
Voorhees, R. M., Schmeing, T. M., Kelley, A. C. & Ramakrishnan, V. The mechanism for activation of GTP hydrolysis on the ribosome. Science 330, 835–838 (2010)
Hausner, T. P., Atmadja, J. & Nierhaus, K. H. Evidence that the G2661 region of 23S rRNA is located at the ribosomal binding sites of both elongation factors. Biochimie 69, 911–923 (1987)
Bilgin, N. & Ehrenberg, M. Mutations in 23 S ribosomal RNA perturb transfer RNA selection and can lead to streptomycin dependence. J. Mol. Biol. 235, 813–824 (1994)
Daviter, T., Wieden, H.-J. & Rodnina, M. V. Essential role of histidine 84 in elongation factor Tu for the chemical step of GTP hydrolysis on the ribosome. J. Mol. Biol. 332, 689–699 (2003)
Moazed, D., Robertson, J. M. & Noller, H. F. Interaction of elongation factors EF-G and EF-Tu with a conserved loop in 23S RNA. Nature 334, 362–364 (1988)
Ogle, J. M. et al. Recognition of cognate transfer RNA by the 30S ribosomal subunit. Science 292, 897–902 (2001)
Jenner, L., Demeshkina, N., Yusupova, G. & Yusupov, M. Structural rearrangements of the ribosome at the tRNA proofreading step. Nat. Struct. Mol. Biol. 17, 1072–1078 (2010)
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006)
Demeshkina, N., Jenner, L., Westhof, E., Yusupov, M. & Yusupova, G. A new understanding of the decoding principle on the ribosome. Nature 484, 256–259 (2012)
Zeng, X., Chugh, J., Casiano-Negroni, A., Al-Hashimi, H. M. & Brooks, C. L., III . Flipping of the ribosomal A-site adenines provides a basis for tRNA selection. J. Mol. Biol. 426, 3201–3213 (2014)
Schrode, P., Huter, P., Clementi, N. & Erlacher, M. Atomic mutagenesis at the ribosomal decoding site. RNA Biol. 14, 104–112 (2016)
Khade, P. K., Shi, X. & Joseph, S. Steric complementarity in the decoding center is important for tRNA selection by the ribosome. J. Mol. Biol. 425, 3778–3789 (2013)
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013)
Grigorieff, N. Frealign: an exploratory tool for single-particle cryo-EM. Methods Enzymol. 579, 191–226 (2016)
Loveland, A. B. et al. Ribosome•RelA structures reveal the mechanism of stringent response activation. eLife 5, e17029 (2016)
Wimberly, B. T. et al. Structure of the 30S ribosomal subunit. Nature 407, 327–339 (2000)
Schuwirth, B. S. et al. Structures of the bacterial ribosome at 3.5 Å resolution. Science 310, 827–834 (2005)
Nissen, P. et al. Crystal structure of the ternary complex of Phe-tRNAPhe, EF-Tu, and a GTP analog. Science 270, 1464–1472 (1995)
Fischer, N. et al. Structure of the E. coli ribosome-EF-Tu complex at <3 Å resolution by Cs-corrected cryo-EM. Nature 520, 567–570 (2015)
Valle, M. et al. Cryo-EM reveals an active role for aminoacyl-tRNA in the accommodation process. EMBO J. 21, 3557–3567 (2002)
Stark, H. et al. Visualization of elongation factor Tu on the Escherichia coli ribosome. Nature 389, 403–406 (1997)
Ledoux, S. & Uhlenbeck, O. C. Different aa-tRNAs are selected uniformly on the ribosome. Mol. Cell 31, 114–123 (2008)
Fahlman, R. P., Dale, T. & Uhlenbeck, O. C. Uniform binding of aminoacylated transfer RNAs to the ribosomal A and P sites. Mol. Cell 16, 799–805 (2004)
Zhang, J., Ieong, K.-W., Mellenius, H. & Ehrenberg, M. Proofreading neutralizes potential error hotspots in genetic code translation by transfer RNAs. RNA 22, 896–904 (2016)
Powers, T. & Noller, H. F. Evidence for functional interaction between elongation factor Tu and 16S ribosomal RNA. Proc. Natl Acad. Sci. USA 90, 1364–1368 (1993)
Cochella, L., Brunelle, J. L. & Green, R. Mutational analysis reveals two independent molecular requirements during transfer RNA selection on the ribosome. Nat. Struct. Mol. Biol. 14, 30–36 (2007)
Abdi, N. M. & Fredrick, K. Contribution of 16S rRNA nucleotides forming the 30S subunit A and P sites to translation in Escherichia coli. RNA 11, 1624–1632 (2005)
Santer, M. et al. Functional effects of a G to U base change at position 530 in a highly conserved loop of Escherichia coli 16S RNA. Biochemistry 32, 5539–5547 (1993)
Taliaferro, D. L. & Farabaugh, P. J. Testing constraints on rRNA bases that make nonsequence-specific contacts with the codon-anticodon complex in the ribosomal A site. RNA 13, 1279–1286 (2007)
Westhof, E. Isostericity and tautomerism of base pairs in nucleic acids. FEBS Lett. 588, 2464–2469 (2014)
Fischer, N. et al. The pathway to GTPase activation of elongation factor SelB on the ribosome. Nature 540, 80–85 (2016)
Budkevich, T. V. et al. Regulation of the mammalian elongation cycle by subunit rolling: a eukaryotic-specific ribosome rearrangement. Cell 158, 121–131 (2014)
Brilot, A. F., Korostelev, A. A., Ermolenko, D. N. & Grigorieff, N. Structure of the ribosome with elongation factor G trapped in the pretranslocation state. Proc. Natl Acad. Sci. USA 110, 20994–20999 (2013)
Moazed, D. & Noller, H. F. Interaction of tRNA with 23S rRNA in the ribosomal A, P, and E sites. Cell 57, 585–597 (1989)
Moazed, D. & Noller, H. F. Transfer RNA shields specific nucleotides in 16S ribosomal RNA from attack by chemical probes. Cell 47, 985–994 (1986)
Traub, P., Mizushima, S., Lowry, C. V. & Nomura, M. in Nucleic Acids and Protein Synthesis, Part C (eds Kaplan, N. & Colowick, N. ) Vol. 20, 391–407 (Academic Press, 1971)
Walker, S. E. & Fredrick, K. Preparation and evaluation of acylated tRNAs. Methods 44, 81–86 (2008)
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005)
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996)
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015)
Grant, T. & Grigorieff, N. Automatic estimation and correction of anisotropic magnification distortion in electron microscopes. J. Struct. Biol. 192, 204–208 (2015)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Chen, J. Z. & Grigorieff, N. SIGNATURE: a single-particle selection system for molecular electron microscopy. J. Struct. Biol. 157, 168–173 (2007)
Gabashvili, I. S. et al. Solution structure of the E. coli 70S ribosome at 11.5 Å resolution. Cell 100, 537–549 (2000)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Cardone, G., Heymann, J. B. & Steven, A. C. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 184, 226–236 (2013)
Korostelev, A. et al. Crystal structure of a translation termination complex formed with release factor RF2. Proc. Natl Acad. Sci. USA 105, 19684–19689 (2008)
Rozov, A., Westhof, E., Yusupov, M. & Yusupova, G. The ribosome prohibits the G•U wobble geometry at the first position of the codon-anticodon helix. Nucleic Acids Res. 44, 6434–6441 (2016)
Bairoch, A., Boeckmann, B., Ferro, S. & Gasteiger, E. Swiss-Prot: juggling between evolution and stability. Brief. Bioinform. 5, 39–55 (2004)
Tishchenko, S. et al. High-resolution crystal structure of the isolated ribosomal L1 stalk. Acta Crystallogr. D 68, 1051–1057 (2012)
Pettersen, E. F. et al. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Korostelev, A., Bertram, R. & Chapman, M. S. Simulated-annealing real-space refinement as a tool in model building. Acta Crystallogr. D 58, 761–767 (2002)
Chapman, M. S. Restrained real-space macromolecular atomic refinement using a new resolution-dependent electron-density function. Acta Crystallogr. A 51, 69–80 (1995)
Gonen, T. et al. Lipid-protein interactions in double-layered two-dimensional AQP0 crystals. Nature 438, 633–638 (2005)
Zhou, G., Wang, J., Blanc, E. & Chapman, M. S. Determination of the relative precision of atoms in a macromolecular structure. Acta Crystallogr. D 54, 391–399 (1998)
Laurberg, M. et al. Structural basis for translation termination on the 70S ribosome. Nature 454, 852–857 (2008)
Adams, P. D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011)
Brunger, A. T. Version 1.2 of the crystallography and NMR system. Nat. Protocols 2, 2728–2733 (2007)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
DeLano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, 2002)
Tinoco, I. Jr & Gonzalez, R. L. Jr. Biological mechanisms, one molecule at a time. Genes Dev. 25, 1205–1231 (2011)
Gonzalez, R. L. Jr, Chu, S. & Puglisi, J. D. Thiostrepton inhibition of tRNA delivery to the ribosome. RNA 13, 2091–2097 (2007)
Engh, R. A. & Huber, R. Accurate bond and angle parameters for X-ray protein structure refinement. Acta Cryst A 47, 392–400 (1991)
Acknowledgements
We thank A. Park for help with preparing ribosome complexes; C. Xu and M. Rigney for help with preparing and screening cryo-EM grids at the cryo-EM facility at Brandeis University; Z. Yu, C. Hong, A. Rohou and R. Diaz-Avalos for data collection at Janelia Research Campus; D. Ermolenko for sharing an EF-Tu-overexpression plasmid and helpful comments on the manuscript; A. Korennykh, D. Conte Jr and members of the Korostelev laboratories for comments on the manuscript. This study was supported by NIH Grants R01 GM106105 and GM107465 (to A.A.K.) and P01 GM62580 (to N.G.). A.B.L. performed this work as a Howard Hughes Medical Institute Fellow of the Helen Hay Whitney Foundation.
Author information
Authors and Affiliations
Contributions
A.B.L., N.G. and A.A.K. designed the project; A.B.L. prepared ribosome complexes, collected and analysed cryo-EM data; G.D. assisted with protein purification and prepared ribosomes; N.G. and A.A.K. oversaw cryo-EM data processing; A.B.L. and A.A.K. built and refined structural models; A.B.L. and A.A.K. wrote the manuscript; all authors contributed to manuscript finalization.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. Erlacher, R. Gillet and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Overview of classification procedures and resolution curves for all structures.
a, Scheme of refinement and classification procedures for the cognate dataset. b, Fourier shell correlation (FSC) curves for the cognate structures. c, Scheme of refinement and classification procedures for the near-cognate dataset. d, FSC curves for the near-cognate structures.
Extended Data Figure 2 Cryo-EM densities for ternary complex in each structure.
a, Cryo-EM density for ternary complex and codon in structure I is shown at 3σ after applying a B-factor of −36 Å2. b, Cryo-EM density for cognate tRNA and codon in structure I is shown as in a. c, Cryo-EM density for the anticodon and codon, which are not base paired, in structure I is shown at 4σ after applying a B-factor of −36 Å2. d, Cryo-EM density for ternary complex and codon in structure II is shown at 3σ after applying a B-factor of −50 Å2. e, Cryo-EM density for cognate tRNA and codon in structure II is shown as in d. f, Cryo-EM density for the anticodon and codon, which are base paired, in structure II is shown at 4.5σ after applying a B-factor of −50 Å2. g, Cryo-EM density for ternary complex and codon in structure III is shown at 4σ after applying a B-factor of −100 Å2. h, Cryo-EM density for cognate tRNA and codon in structure II is shown as in g. i, Cryo-EM density for the anticodon and codon, which are base paired, in structure III is shown at 5σ after applying a B-factor of −150 Å2. j, Cryo-EM density for ternary complex and codon in structure I-nc is shown at 3σ. k, Cryo-EM density for near-cognate tRNA and codon in structure I-nc is shown as in j. l, Cryo-EM density for the anticodon and codon, which are not base paired, in structure I-nc is shown at 3.5σ for T tRNA and 16S rRNA or 4σ for mRNA. m, Cryo-EM density for ternary complex and codon in structure II-nc is shown at 3σ after applying a B-factor of −25 Å2. n, Cryo-EM density for near-cognate A*/T tRNA and codon in structure II-nc is shown as in m. o, Cryo-EM density for the anticodon and codon, which are interacting in structure II-nc is shown at 4.5σ after applying a B-factor of −25 Å2. p, Cryo-EM density for ternary complex and codon in structure III-nc is shown at 4σ after applying a B-factor of −50 Å2. q, Cryo-EM density for near-cognate tRNA and codon in structure II-nc is shown as in p. r, Cryo-EM density for the anticodon and codon, which are base paired, in structure III-nc is shown at 5.2σ after applying a B-factor of −60 Å2.
Extended Data Figure 3 Local resolution of cryo-EM maps of the cognate and near-cognate complexes.
Local resolution of each cryo-EM map was determined using Blocres. a, An overview of the structure I map. The unsharpened map is shown at 5σ, coloured using a scale ranging from 3.5 Å to 8.5 Å (left). b, An overview of the structure II map shown as in a. c, An overview of the structure III map. The unsharpened map is shown at 5σ, coloured using a scale ranging from 3.0 Å to 8.0 Å (left). d–f, Slab views at the ribosome interior in maps corresponding to structure I (d), structure II (e) and structure III (f), prepared and coloured as in a, b and c, respectively. g, Close-up view of decoding centre of structure I. The map was sharpened by applying a B-factor of −36 Å2 and is shown at 4.5σ, coloured as in a. h, Close-up view of decoding centre of structure II. The map was sharpened by applying a B-factor of −50 Å2 and is shown at 5σ, coloured as in a. i, Close-up view of decoding centre of structure III. The map was sharpened by applying a B-factor of −100 Å2 and is shown at 4σ, coloured as in c. j, An overview of the structure I-nc map. The unsharpened map is shown at 5σ and is coloured using a scale ranging from 3.5 Å to 8.5 Å (left). k, An overview of the structure II-nc map, as in j. l, An overview of the structure III-nc map, as in j. m–o, Slab views at the ribosome interior in maps corresponding to structure I-nc (m), structure II-nc (n) and structure III-nc (o), prepared and coloured as in j. p, Close-up view of decoding centre of structure I-nc. The unsharpened map is shown at 4.5σ, coloured as in j. q, Close-up view of decoding centre of structure II-nc. The map was sharpened by applying a B-factor of −25 Å2 and is shown at 5σ, coloured as in j. r, Close-up view of the decoding centre of structure III-nc. The map was sharpened by applying a B-factor of −50 Å2 and is shown at 5σ, coloured as in j.
Extended Data Figure 4 30S domain closure and aminoacyl-tRNA conformations in cognate and near-cognate complexes.
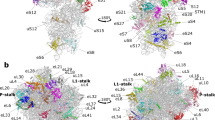
a, Comparison of the 30S conformations among structures I (magenta), II (grey) and III (multi-coloured). Superposition was achieved by structural alignment of 23S rRNA. b, Superposition of structure II (grey) and III (multi-coloured) highlighting the movement of the shoulder including the 530 loop towards the 30S body including h44. c, Different conformations of aminoacyl-tRNA in structures I and II: T tRNA (structure I) is relaxed, whereas A*/T tRNA (structure II) is kinked to base-pair with mRNA. d, Interaction of T tRNA in structure I with the decoding centre is shown in surface representation. All atoms within 15 Å of residues 30–38 of T tRNA are shown except for 16S residues 950–964 and 984–985, which were omitted for clarity. e, Interaction of A*/T tRNA in structure II with the decoding centre is shown in surface representation as in d. f, Cognate tRNA anticodon samples positions between those in structures I and II. Additional focused classification into four classes revealed intermediate classes with A-site tRNA density midway between the T tRNA and A*/T tRNA conformations. The cryo-EM density, within 15 Å of residues 30–38 of T or A*/T tRNA, is shown with exceptions as in d, at 3σ after applying a B-factor of +200 Å2. g, Near-cognate tRNA anticodon samples positions between those in structure I-nc and structure II-nc. Additional focused classification into four classes revealed intermediate classes with A-site tRNA density midway between the T tRNA and A*/T tRNA conformations. The cryo-EM density is shown as in f.
Extended Data Figure 5 Sliding of tRNA elbow along the L11 stalk from structures I–III towards the P-site tRNA agrees with distance changes inferred from smFRET studies of tRNA decoding.
a, Overview of structure III with box highlighting the location of tRNA elbow and L11 stalk. b, tRNA elbow residues G19 and C57 slide along L11 stalk residues 1095 and 1067 from structure II (grey) to structure III (green). Superposition was achieved by aligning on residues 1095 and 1067 of L11 stalk. c, The elbow of T tRNA (green) and L11 stalk in structure I. d, The elbow of A*/T tRNA (green) and L11 stalk in structure II. e, The elbow of A/T tRNA (green) and L11 stalk in structure III. f, The distance between nucleotide 47 of T tRNA (magenta) and nucleotide 8 of P-site tRNA (orange) is shown. These locations were used in smFRET studies of tRNA decoding8,76. g, The distance between nucleotide 47 of A*/T tRNA (grey) and nucleotide 8 of P-site tRNA (orange) is shown. h, The distance between nucleotide 47 of A/T tRNA (green) and nucleotide 8 of P-site tRNA (orange) is shown. The distance changes between T or A*/T tRNA to A/T tRNA are consistent with the change from a low FRET value of 0.35 in the early tRNA decoding states to a mid-FRET value of 0.5 in the GTP activated tRNA decoding state, as described in the Methods.
Extended Data Figure 6 Conformational differences in the decoding centres of cognate and near-cognate structures I–III.
a, Cryo-EM density (shown as mesh) of the decoding centre in structure I. The map was sharpened by applying a B-factor of −36 Å2 and density is shown at 3.5σ for mRNA and anticodon of T tRNA, 5.5σ for G530, 4.0σ for A1492, A1493 and A1913. b, Cryo-EM density for the G530 region in structure I is shown with two conformations of G530-syn (top) and G530-anti (bottom). Both conformations fit with similar local cross-correlation coefficients (Methods). The map was sharpened by applying a B-factor of −75 Å2 and density is shown at 5σ. c, Cryo-EM density for the decoding centre in structure II. The map was sharpened by applying a B-factor of −75 Å2 and density is shown at 5.5σ for G530 or at 4σ for the mRNA and the anticodon of A*/T tRNA, A1492, A1493 and A1913. Density for residue 1492 (shown in grey) is compatible with two conformations, in and out of h44. d, Cryo-EM density for the G530 region in structure II. The map was sharpened by applying a B-factor of −100 Å2 and density is shown at 5σ. e, Cryo-EM density for the decoding centre in structure III. The map was sharpened by applying a B-factor of −150 Å2 and density is shown at 4.5σ for the mRNA and the anticodon of A/T tRNA, G530, A1492, A1493 and A1913. f, Cryo-EM density for the G530 region in structure III. The map was sharpened by applying a B-factor of −150 Å2 and density is shown at 5σ. g, Cryo-EM density (grey mesh) showing the anti-conformation of G530 (yellow model) in structure II. The map was sharpened by applying a B-factor of −100 Å2 and density is shown at 5σ. h, i, Cryo-EM density showing the anti-conformation of G530 in structure III. The map was sharpened by applying a B-factor of −150 Å2 and density is shown at 5σ. j, Cryo-EM density of the decoding centre in structure I-nc. The map was not B-factor sharpened and density is shown at 3.75σ for mRNA, the anticodon of T tRNA, A1492, A1493 and A1913, or at 5σ for G530. k, Cryo-EM density for the decoding centre in structure II-nc. The map was sharpened by applying a B-factor of −25 Å2 and density is shown at 4.3σ for mRNA, the anticodon of tRNA, A1492, A1493 and A1913, or at 5.5σ for G530. l, Cryo-EM density of structure II-nc for the 30S shoulder including G530. The map was sharpened by applying a B-factor of −120 Å2 and density is shown at 3.5σ. m, Cryo-EM density for the decoding centre in structure III-nc. The map was sharpened by applying a B-factor of −50 Å2 and density is shown at 4.75σ for mRNA and the anticodon of tRNA, G530, A1492 and A1493, and at 4.5 for A1913. n, Cryo-EM density of structure III-nc for the 30S shoulder including G530. The map was sharpened by applying a B-factor of −100 Å2 and density is shown at 4.5σ. o, Cryo-EM density showing the syn-conformation of G530 in structure II-nc. The map was sharpened by applying a B-factor of −120 Å2 and density is shown at 3.7σ. p, q, Cryo-EM density showing the anti-conformation of G530 in structure III-nc. The map was sharpened by applying a B-factor of −120 Å2 and density is shown at 3.3σ. r, Nucleotide 34 of the anticodon stacks on C1054 in the cognate structure II. s, Cryo-EM density (grey mesh) for the cognate structure II. The map was sharpened by applying a B-factor of −100 Å2 and density is shown at 5σ. t, Cryo-EM density for the cognate structure III. The map was sharpened by applying a B-factor of −150 Å2 and density is shown at 5.5σ. u, Nucleotide 34 of the near-cognate anticodon in structure II-nc is shifted by approximately 2 Å from C1054, relative to its position in the cognate complex (shown in r). v, Cryo-EM density for the near-cognate structure II-nc. The map was sharpened by applying a B-factor of −120 Å2 and density is shown at 3.5σ. w, Cryo-EM density for the near-cognate structure III-nc. The map was sharpened by applying a B-factor of −50 Å2 and density is shown at 5.5σ. Modification of U34 of tRNALys to 5-methylaminomethyl-2-thiouridine (mnm5s2U34) is shown in u–w.
Extended Data Figure 7 Anchoring of EF-Tu to the 30S shoulder in structures I, II and III and to the SRL in structures III and III-nc.
a, Overview of structure III with boxes highlighting locations of EF-Tu contacts to 30S shoulder (dashed box) and to SRL (solid box). b, The contacts of EF-Tu with the 30S shoulder are similar among structures I (purple), II (grey) and III (red). c, Cryo-EM density for EF-Tu (red) and 16S rRNA (pale yellow) in structure I. The map was sharpened by applying a B-factor of −36 Å2 and is shown at 3σ. d, Cryo-EM density for EF-Tu and 16S rRNA in structure II. The map was sharpened by applying a B-factor of −75 Å2 and is shown at 3.5σ. e, Cryo-EM density for EF-Tu and 16S rRNA in structure III. The map was sharpened by applying a B-factor of −100 Å2 and is shown at 4.5σ. f, Cryo-EM density for EF-Tu (red) and the SRL of 23S rRNA (pale cyan) in structure III. The map was sharpened by applying a B-factor of −150 Å2 and is shown at 4.5σ. g, Cryo-EM density for EF-Tu and SRL of 23S rRNA in structure III-nc. The map was sharpened by applying a B-factor of −50 Å2 and is shown at 4.5σ.
Extended Data Figure 8 Modifications of A37 in tRNAPhe and tRNALys, and magnesium ion coordination near G530.
a, Cryo-EM density for the codon–anticodon helix in structure II shows that the 2-methylthio moiety of 2-methylthio-N6-(2-isopentenyl)-adenosine at position 37 of tRNAPhe (ms2i6A37) stacks on U1 of the A-site codon. The map was sharpened by applying a B-factor of −75 Å2 and density is shown at 4.8σ. b, Cryo-EM density for the codon–anticodon helix in structure III shows that ms2i6A37 of tRNAPhe stacks on U1 similarly to that in structure II. The map was sharpened by applying a B-factor of −150 Å2 and density is shown at 6σ. c, Cryo-EM density for the codon–anticodon helix in structure III-nc shows that 6-threonylcarbamoyl adenosine at position 37 of tRNALys (t6A37) stacks on A1. The map was sharpened by applying a B-factor of −120 Å2 and density is shown at 4.5σ. d, Cryo-EM density for structure II shows the N6 modification of ms2i6A37 of tRNAPhe in close proximity to U33. The map was sharpened by applying a B-factor of −75 Å2 and density is shown at 4σ. e, Cryo-EM density for structure III shows the N6 modification of ms2i6A37 of tRNAPhe in close proximity to U33. The map was sharpened by applying a B-factor of −150 Å2 and density is shown at 4σ. f, Cryo-EM density for structure III-nc shows the N6 modification of t6A37 of tRNALys. The map was sharpened by applying a B-factor of −120 Å2 and density is shown at 3.5σ. g, In structure II, three magnesium ions (magenta) are coordinated (dotted lines) by G530 and codon–anticodon helix (in some instances, the coordination probably occurs via water molecules). Density for magnesium ions (mesh) was sharpened by applying a B-factor of −75 Å2, shown at 4σ. h, In structure III, the magnesium ions shift with G530. Density was sharpened by applying a B-factor of −150 Å2, shown at 4σ. i, In structure III-nc, three magnesium ions are seen at equivalent position to those in structure III. Density was sharpened by applying a B-factor of −120 Å2, shown at 3σ.
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion and Supplementary References. (PDF 735 kb)
Video 1: Animation showing the mechanism of tRNA discrimination by the ribosome, inferred from ensemble cryo-EM.
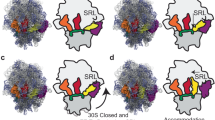
Rejection of a near-cognate tRNA (blue) is followed by acceptance of the cognate tRNA (green). Four scenes are shown: (1) A view of the complete 70S complex, as in Figure 1. Near-cognate ternary complex (EF-Tu is red) samples the A site of the open 30S subunit (Structures I-nc and II-nc), without inducing 30S subunit rearrangements. Cognate ternary complex samples the A site (I and II), resulting in the movement of the 30S shoulder (“30S closed”). This leads to tRNA stabilization and docking of EF-Tu at the SRL, resulting in a GTPase-activated state (III). (2) A close-up view of the 30S A site, showing the movement of the 30S shoulder upon stabilization of the cognate tRNA (III). (3) A close-up view of the decoding center. Near-cognate tRNA does not engage with G530, which remains in an OFF conformation (I-nc and II-nc). Cognate codon-anticodon helix induces G530 to switch to the SEMI-ON state (II; hydrogen bonds are shown by dotted lines). Stabilization of nucleotides A1492, A1493 and A1913 coincides with G530 movement into the ON state, latching the decoding center and shifting the 30S shoulder (III). (4) EF-Tu is at the 30S shoulder and separated from the SRL in the 30S-open states (I-nc, II-nc, I and II). EF-Tu docks at the SRL upon shoulder movement (III), activating catalytic residue H84 to hydrolyze GTP. EF-Tu rearrangement upon GTP hydrolysis (PDB: 5AFI) results in EF-Tu dissociation and accommodation of cognate tRNA into the 50S A site (not shown). (MOV 6892 kb)
Rights and permissions
About this article
Cite this article
Loveland, A., Demo, G., Grigorieff, N. et al. Ensemble cryo-EM elucidates the mechanism of translation fidelity. Nature 546, 113–117 (2017). https://doi.org/10.1038/nature22397
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22397
This article is cited by
-
Structures of the ribosome bound to EF-Tu–isoleucine tRNA elucidate the mechanism of AUG avoidance
Nature Structural & Molecular Biology (2024)
-
Geometric alignment of aminoacyl-tRNA relative to catalytic centers of the ribosome underpins accurate mRNA decoding
Nature Communications (2023)
-
Modulation of translational decoding by m6A modification of mRNA
Nature Communications (2023)
-
Quality control of protein synthesis in the early elongation stage
Nature Communications (2023)
-
Cryo-EM reveals dynamics of Tetrahymena group I intron self-splicing
Nature Catalysis (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.