Abstract
The organization of the eukaryotic cell into discrete membrane-bound organelles allows for the separation of incompatible biochemical processes, but the activities of these organelles must be coordinated. For example, lipid metabolism is distributed between the endoplasmic reticulum for lipid synthesis, lipid droplets for storage and transport, mitochondria and peroxisomes for β-oxidation, and lysosomes for lipid hydrolysis and recycling1,2,3,4,5. It is increasingly recognized that organelle contacts have a vital role in diverse cellular functions5,6,7,8. However, the spatial and temporal organization of organelles within the cell remains poorly characterized, as fluorescence imaging approaches are limited in the number of different labels that can be distinguished in a single image9. Here we present a systems-level analysis of the organelle interactome using a multispectral image acquisition method that overcomes the challenge of spectral overlap in the fluorescent protein palette. We used confocal and lattice light sheet10 instrumentation and an imaging informatics pipeline of five steps to achieve mapping of organelle numbers, volumes, speeds, positions and dynamic inter-organelle contacts in live cells from a monkey fibroblast cell line. We describe the frequency and locality of two-, three-, four- and five-way interactions among six different membrane-bound organelles (endoplasmic reticulum, Golgi, lysosome, peroxisome, mitochondria and lipid droplet) and show how these relationships change over time. We demonstrate that each organelle has a characteristic distribution and dispersion pattern in three-dimensional space and that there is a reproducible pattern of contacts among the six organelles, that is affected by microtubule and cell nutrient status. These live-cell confocal and lattice light sheet spectral imaging approaches are applicable to any cell system expressing multiple fluorescent probes, whether in normal conditions or when cells are exposed to disturbances such as drugs, pathogens or stress. This methodology thus offers a powerful descriptive tool and can be used to develop hypotheses about cellular organization and dynamics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Barbosa, A. D., Savage, D. B. & Siniossoglou, S. Lipid droplet-organelle interactions: emerging roles in lipid metabolism. Curr. Opin. Cell Biol. 35, 91–97 (2015)
Shai, N ., Schuldiner, M . & Zalckvar, E. No peroxisome is an island—peroxisome contact sites. Biochim. Biophys. Acta (2015)
Settembre, C., Fraldi, A., Medina, D. L. & Ballabio, A. Signals from the lysosome: a control centre for cellular clearance and energy metabolism. Nat. Rev. Mol. Cell Biol. 14, 283–296 (2013)
Tatsuta, T., Scharwey, M. & Langer, T. Mitochondrial lipid trafficking. Trends Cell Biol. 24, 44–52 (2014)
Phillips, M. J. & Voeltz, G. K. Structure and function of ER membrane contact sites with other organelles. Nat. Rev. Mol. Cell Biol. 17, 69–82 (2016)
Rowland, A. A., Chitwood, P. J., Phillips, M. J. & Voeltz, G. K. ER contact sites define the position and timing of endosome fission. Cell 159, 1027–1041 (2014)
Friedman, J. R. et al. ER tubules mark sites of mitochondrial division. Science 334, 358–362 (2011)
Wilfling, F. et al. Arf1/COPI machinery acts directly on lipid droplets and enables their connection to the ER for protein targeting. eLife 3, e01607 (2014)
Neher, R. & Neher, E. Optimizing imaging parameters for the separation of multiple labels in a fluorescence image. J. Microsc. 213, 46–62 (2004)
Chen, B.-C. et al. Lattice light-sheet microscopy: imaging molecules to embryos at high spatiotemporal resolution. Science 346, 1257998 (2014)
Herms, A. et al. AMPK activation promotes lipid droplet dispersion on detyrosinated microtubules to increase mitochondrial fatty acid oxidation. Nat. Commun. 6, 7176 (2015)
Friedman, J. R., Webster, B. M., Mastronarde, D. N., Verhey, K. J. & Voeltz, G. K. ER sliding dynamics and ER-mitochondrial contacts occur on acetylated microtubules. J. Cell Biol. 190, 363–375 (2010)
de Forges, H., Bouissou, A. & Perez, F. Interplay between microtubule dynamics and intracellular organization. Int. J. Biochem. Cell Biol. 44, 266–274 (2012)
Friedman, J. R., Dibenedetto, J. R., West, M., Rowland, A. A. & Voeltz, G. K. Endoplasmic reticulum-endosome contact increases as endosomes traffic and mature. Mol. Biol. Cell 24, 1030–1040 (2013)
Wait, E. et al. Visualization and correction of automated segmentation, tracking and lineaging from 5-D stem cell image sequences. BMC Bioinformatics 15, 328 (2014)
Rambold, A. S., Cohen, S. & Lippincott-Schwartz, J. Fatty acid trafficking in starved cells: regulation by lipid droplet lipolysis, autophagy, and mitochondrial fusion dynamics. Dev. Cell 32, 678–692 (2015)
Singh, R. et al. Autophagy regulates lipid metabolism. Nature 458, 1131–1135 (2009)
Ikonen, E. Cellular cholesterol trafficking and compartmentalization. Nat. Rev. Mol. Cell Biol. 9, 125–138 (2008)
Eisenberg-Bord, M., Shai, N., Schuldiner, M. & Bohnert, M. A tether is a tether is a tether: tethering at membrane contact sites. Dev. Cell 39, 395–409 (2016)
Patterson, G. H. & Lippincott-Schwartz, J. A photoactivatable GFP for selective photolabeling of proteins and cells. Science 297, 1873–1877 (2002)
Snapp, E. L., Sharma, A., Lippincott-Schwartz, J. & Hegde, R. S. Monitoring chaperone engagement of substrates in the endoplasmic reticulum of live cells. Proc. Natl Acad. Sci. USA 103, 6536–6541 (2006)
Kim, P. K., Mullen, R. T., Schumann, U. & Lippincott-Schwartz, J. The origin and maintenance of mammalian peroxisomes involves a de novo PEX16-dependent pathway from the ER. J. Cell Biol. 173, 521–532 (2006)
Jahr, W., Schmid, B., Schmied, C., Fahrbach, F. O. & Huisken, J. Hyperspectral light sheet microscopy. Nat. Commun. 6, 7990 (2015)
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 9, 62–66 (1979)
Jaqaman, K. et al. Robust single-particle tracking in live-cell time-lapse sequences. Nat. Methods 5, 695–702 (2008)
Gatta, A. T. & Levine, T. P. Piecing Together the Patchwork of Contact Sites. Trends Cell Biol. 27, 214–229 (2016)
Gonzales, R ., Woods, R . & Eddins, S. Digital Image Processing Using MATLAB, 2nd ed. (Gatesmark Publishing, 2009)
Winter, M., Mankowski, W., Wait, E., Temple, S. & Cohen, A. R. LEVER: software tools for segmentation, tracking and lineaging of proliferating cells. Bioinformatics 32, 3530–3531 (2016)
Cohen, A. R., Bjornsson, C. S., Temple, S., Banker, G. & Roysam, B. Automatic summarization of changes in biological image sequences using algorithmic information theory. IEEE Trans. Pattern Anal. Mach. Intell. 31, 1386–1403 (2009)
Tibshirani, R., Walther, G. & Hastie, T. Estimating the number of clusters in a data set via the gap statistic. J. R. Stat. Soc. Series B Stat. Methodol. 63, 411–423 (2001)
Garini, Y., Young, I. T. & McNamara, G. Spectral imaging: principles and applications. Cytometry A 69, 735–747 (2006)
Valm, A. M. et al. Systems-level analysis of microbial community organization through combinatorial labeling and spectral imaging. Proc. Natl Acad. Sci. USA 108, 4152–4157 (2011)
Valm, A. M., Oldenbourg, R. & Borisy, G. G. Multiplexed spectral imaging of 120 different fluorescent labels. PLoS One 11, e0158495 (2016)
Tsuriel, S., Gudes, S., Draft, R. W., Binshtok, A. M. & Lichtman, J. W. Multispectral labeling technique to map many neighboring axonal projections in the same tissue. Nat. Methods 12, 547–552 (2015)
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005)
Rodriguez, E. A. et al. A far-red fluorescent protein evolved from a cyanobacterial phycobiliprotein. Nat. Methods 13, 763–769 (2016)
Johnson, I. & Spence, M. T. Z. (eds) The Molecular Probes Handbook—A Guide to Fluorescent Probes and Labeling Technologies (Life Technologies, 2010)
Dickinson, M. E., Bearman, G., Tille, S., Lansford, R. & Fraser, S. E. Multi-spectral imaging and linear unmixing add a whole new dimension to laser scanning fluorescence microscopy. Biotechniques 31, 1272–1278 (2001)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012)
Acknowledgements
We are grateful to P. Sengupta and other members of the Lippincott-Schwartz laboratory for helpful discussions. This work was supported by the Intramural Research Program of the National Institutes of Health (A.M.V., S.C., and J.L.-S.) and the Howard Hughes Medical Institute (W.R.L., E.B., and J.L.-S.), by a Postdoctoral Research Associate (PRAT) Fellowship from the National Institute of General Medical Sciences to A.M.V. and by NIH grant R01AG041861 from the National Institute on Aging to E.W. and A.C.
Author information
Authors and Affiliations
Contributions
A.M.V., S.C., and J.L.-S. conceived and designed the study. A.M.V., S.C., and W.R.L performed the experiments. M.W.D. contributed novel reagents. A.M.V., S.C., J.M., E.W., U.H., and A.R.C. analysed the data. A.M.V., S.C., and J.L.-S. wrote the manuscript, with input from all co-authors. J.L.-S. and E.B. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S.-H. Shim and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Strategy for six-colour labelling, image acquisition and analysis (confocal).
Fluorescence spectral imaging has emerged as a technology that allows many different spectrally variant fluorescent markers to be distinguished in a single sample31. The most widely used approach for computational analysis of spectral images, called linear unmixing, involves a matrix inverse operation to find the best fit of known fluorophore spectra to that of the recorded spectrum at every pixel in a digital image9. Although this and other multispectral approaches have been used in commercial instruments to distinguish multiple combinations of organic dyes in fixed microbes32,33 and fixed neuronal tissue34, its application to multi-labelled cells and their quantitative analysis remains underdeveloped in live-cell experiments. a, Published emission spectra for the fluorophores used in confocal experiments: CFP35, EGFP35, YFP35, mOrange2 (ref. 35), mApple36 and BODIPY 665/676 (ref. 37). b, Schematic of the hardware used for six-colour confocal microscopy. The specimen was excited using three lasers simultaneously, by point-scanning illumination. Emitted light was collected by a linear array of detector elements after being dispersed by a reflective dispersion grating. c, To derive the values for the known fluorophore matrix, images of singly labelled cells were acquired at each wavelength and under the same acquisition conditions used to acquire images of six-colour-labelled cells. Intensity values centred at 512 nm and 591 nm were zero for all cells because these detector elements were blocked to prevent scattered laser excitation light from reaching the detector. d, Graphical representation of the unmixing matrix. The normalized intensity values at each wavelength range from 0 to 1. e, Close-up of a region of the cell shown in Fig. 1a. Scale bars, 5 μm. Micrographs are representative of 10 cells captured. f, Plots of mean pixel intensity values for all six fluorophores in every pixel in singly labelled cells that were segmented as foreground. Cells were singly labelled with LAMP1–CFP, Mito–EGFP, ss-YFP–KDEL, mOrange2–SKL, mApple–SiT, or BODIPY 665/676. n = 87,307 pixels from one cell (CFP), 5,933 pixels from one cell (EGFP), 84,127 pixels from one cell (YFP), 2,711 pixels from one cell (mOrange2), 11,804 pixels from one cell (mApple), 3,332 pixels from one cell (BODIPY 667/676). Error bars represent s.e.m. AU, arbitrary units. g, Imaging-informatics pipeline for quantitative analysis of organelle contacts. 26-channel micrographs of samples were subjected to pixel-based linear unmixing and spatial deconvolution algorithms, resulting in six-channel unmixed images. These images were segmented to generate features, and contacts between features (within 1 pixel, 97 nm) were analysed in single frames. Alternatively, globular organelles were tracked and their contacts with segmented features analysed over multiple frames. The pipeline is modular and involves five major components: pixel-based linear unmixing of raw image data; spatial deconvolution; segmentation of organelles to generate features; particle tracking of globular organelles over time; and integration of track data with segmented image data to identify organelle contacts between the labelled organelles. The first four modules are implemented in existing software packages, either commercially available (Zeiss Zen and Huygens software) or freely available (histogram-based segmentation algorithms and TrackMate plugin in ImageJ for particle tracking)38,39. For the final component of the pipeline we developed an image analysis program on the Mathematica platform (available for download at http://organelle-interactome.sourceforge.net) that identifies feature-based co-localization.
Extended Data Figure 2 LD–organelle contact duration and dynamics.
a, Histograms showing the duration of LD–organelle contacts in time lapse images of a single cell, acquired and analysed as described in Fig. 1b. n = 480 LD contact events from one cell. b, All the LDs in a single cell were tracked, and their interorganelle contacts mapped with time. A blue line indicates that the LD was successfully tracked at the specified time point. Coloured lines indicate that the tracked LD was within 1 pixel (97 nm) of the following organelles at the specified time point: green, mitochondria; yellow, ER; red, peroxisome; cyan, lysosomes; magenta, Golgi. Tracks are sorted according to LD speed, from fastest to slowest. Only LDs that were tracked for at least 25 out of 60 frames are included. Boxes marked with stars indicate examples where a single LD contacts all five other organelles in the same image frame. Shown here are the contact maps for 38 randomly selected LDs from one cell.
Extended Data Figure 3 Cell-to-cell variation in the organelle interactome over time.
a, The absolute numbers of organelle contacts in each cell at a single time point are displayed as graphical half matrices. Each row in the matrix represents the number of organelle contacts with each target organelle (columns), and is colour-coded from 0 to maximum number of observed contacts in each cell. Each row of graphical matrices represents the organelle interactome in one cell and each column of graphical matrices represents the organelle interactome at a specific time point (0, 75, 150, 225 or 300 s). We performed an analysis of variance (ANOVA) in order to assess the variance in organelle–organelle contacts within cells over time. The results showed that the variance between cells is significantly larger than the variance within an individual cell across time (P < 1 × 10−37). b, Cluster analysis of the organelle contact data for all ten cells. The gap statistic was calculated for 1–9 hypothetical clusters (see ‘Statistics’ section), and no significant differences were found to separate the organelle associations for the ten cells. This suggested there is a reproducible and scalable pattern of organelle contacts despite cell-to-cell differences in the absolute numbers of organelles. n = 100 simulations, error bars represent s.e.m.
Extended Data Figure 4 Validation of six-colour labelling and organelle interaction measurements (confocal).
a, To test the effect of co-expressing all six labels on organelle properties, we compared the number and/or area of organelles in cells singly transfected with one organelle marker or incubated with BODIPY, with cells labelled with all six organelle markers. For LDs, peroxisomes, and lysosomes, mean cross-sectional area and number were measured. For Golgi, total cross-sectional area per cell was measured. For ER and mitochondria, the fraction of cell area occupied by these organelles was measured. Only LD number showed a significant difference between singly versus multiply labelled conditions. n = 20 cells for all six-labelled cells, n = 20 cells (BODIPY only), n = 14 cells (SKL only), n = 21 cells (LAMP-1 only), n = 19 cells (SiT only), n = 18 cells (ER only), n = 20 cells (Mito only). **P < 0.01 (unpaired, two-tailed t-test). Bar heights represent mean values and error bars represent s.e.m. b, Line graphs showing the fraction of LDs, peroxisomes or lysosomes contacting each of the other labelled organelles in one cell over time, measured discretely at 0, 75, 150, 225 and 300 s. The fraction of total LDs, peroxisomes or lysosomes contacting each of the other organelles remained constant over the course of imaging, consistent with minimal perturbation and phototoxicity during the imaging period. c, Line graphs showing the fraction of LDs, peroxisomes or lysosomes contacting each of the other labelled organelles in one cell (cell 1 in Extended Data Fig. 3a) after modulating the threshold value for all channels by a fixed percentage. Dashed lines represent a threshold modulated up or down by 20%. Ideal threshold = 100%. For all organelles except mitochondria, modulating the threshold up or down by up to 20% from the algorithmically determined optimal threshold value did not significantly alter the measured number of organelle contacts, suggesting that our organelle contact measurements are insensitive to small differences in threshold parameters. d, Examples of segmentation based on algorithmically determined, optimal intensity threshold values. Micrographs are representative of 10 cells captured. Scale bar, 10 μm.
Extended Data Figure 5 Effect of nocodazole on organelle contacts.
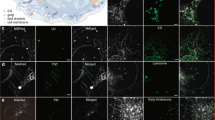
a, Micrographs of a COS-7 cell labelled as in Fig. 1a, except that instead of labelling Golgi, microtubules were labelled with mApple-MAP4-C10. i–iii, Enlargements of the regions outlined in the left panel. iii′, Region iii, without the ER displayed, for clarity. Lysosomes, mitochondria, the ER, peroxisomes, and LDs were all observed in close proximity to microtubules. Scale bars, 10 μm (left) and 2 μm (right). b, The same cell as in a, displaying only the microtubule channel, both before (left) and (after) treatment with 5 μM nocodazole for 1 h. Scale bar, 10 μm. Micrographs in a and b are representative of 20 cells captured. c, Network diagrams of untreated and nocodazole-treated cells. Untreated network is the same as in Fig. 1d. After nocodazole treatment, the ER remains the central node in the network. d, Comparison of object-based organelle contact analysis (bright) versus pixel-based organelle co-localization analysis (pastel). For the pixel-based analysis, a value of 1 indicates perfect co-localization, while a value of 0 indicates the organelles are never co-located. No statistical test was performed. e, Comparison of the effect of nocodazole treatment on organelle contacts when images were analysed using either an object-based or pixel-based co-localization analysis scheme. Red lines connecting the median values indicate that the median number of contacts decreased after nocodazole treatment. Shown are all organelle contact pairs that showed a statistically significant change in contact frequency when cells were treated with nocodazole (unpaired, two-tailed t-test). d, e, Object-based analysis data are the same as in Fig. 2a. n = 11 (nocodazole-treated) or n = 10 (untreated) cells from two experiments. The white line in the centre of each box represents the median value, the upper and lower edges of the boxes represent the 75th and 25th quantile of the data, respectively, and the upper and lower fences represent the 95% confidence level of the distribution.
Extended Data Figure 6 Effect of starvation or excess fatty acids on organelle contacts.
a, Box whisker plots showing the fraction of LDs contacting each of the other labelled compartments in cells grown in complete medium (CM, blue), Hank’s balanced salt solution (HBSS, red), or complete medium supplemented with 300 μM oleic acid (OA, green) for 18 h. *0.05 > P < 0.01, **P < 0.01 (unpaired, two-tailed t-test). The white line in the centre of each box represents the median value, the upper and lower edges of the boxes represent the 75th and 25th quantile of the data, respectively, and the upper and lower fences represent the 95% confidence level of the distribution. b, Network diagrams showing the organelle interactome in cells treated as described in a. a, b, Complete medium data are the same as control data shown in Figs 1d, 2a; n = 10 (complete medium), n = 15 (HBSS), or n = 14 (oleic acid) cells from two experiments.
Extended Data Figure 7 LLS spectral imaging and linear unmixing.
a, Schematic of the hardware used for six-colour light sheet microscopy. The specimen was excited using six lasers sequentially, by LLS illumination. Emitted light passed through a series of interference filters and was collected using a sCMOS camera. b, Plot of the emission intensity of the indicated fluorophores as a function of excitation wavelength, in images of singly labelled cells acquired as described in a. To identify fluorophores in the image data, we applied an excitation-side unmixing algorithm (see Image Acquisition and Unmixing). Our multispectral time-lapse LLS images consisted of upwards of 4.9 billion sets of six-colour-channel pixels (547 × 640 pixels per plane × 140 planes per cell × 100 time points per cell × 10 cells). Because the solution to the unmixing operation at every pixel is independent of every other pixel, we distributed the unmixing operation over 32 cores of a computer workstation. c, Plots of mean pixel intensity values for all six fluorophores in every pixel in singly labelled cells that were segmented as foreground. Cells were singly labelled with CFP–SKL, Mito–EGFP, ss-YFP–KDEL, mApple–SiT, Texas Red dextran, or BODIPY 665/676. The error in LLS unmixing is higher than for confocal (see Extended Data Fig. 1f) as expected and is due partly to the fact that only six channels of spectral information were used to unmix the overlapping spectra. n = 149 pixels (CFP), n = 3,910 pixels (EGFP), n = 9,180 pixels (YFP), n = 1,549 pixels (mApple), n = 806 pixels (Texas Red Dextran), n = 3,248 pixels (BODIPY 667/676). Error bars represent s.e.m. d, Tilted volume rendering of the same cell shown in Fig. 3a. Scale bar, 10 μm. e, Zoomed, segmented images from the cell shown in d. The left panel does not include the ER channel while the right panel does (transparent yellow). Scale bar, 5 μm. Micrographs in d and e are representative of 10 cells captured.
Extended Data Figure 8 Validation of organelle interaction measurements (LLS).
a, Box whisker plots showing the median fraction of LDs, peroxisomes or lysosomes making contact with each of the other labelled compartments in data obtained using confocal (bright) or LLS (pastel) microscopy. Confocal data are the same as in Fig. 2a. n = 10 cells (confocal), n = 10 cells (LLS). No statistical test was performed. The similarity in measurements from LLS and confocal images is likely because the globular organelles that we examined are smaller than the depth of focus of the confocal microscope, ensuring that all their inter-organelle interactions were detected even in the confocal images. b, Line graphs showing the fraction of LDs, peroxisomes or lysosomes contacting each of the other labelled organelles in one cell measured over time at discrete points: 0, 174, 358, 541, 725 and 908 s. c, Line graphs showing the fraction of LDs, peroxisomes or lysosomes contacting each of the other labelled organelles in one cell after modulating the threshold value for all channels by a fixed percentage. Dashed lines represent a threshold modulated by 20%. d, Examples of segmentation performed using the ideal threshold (that is, 100%) in c. Scale bar, 2 μm. Micrographs are representative of 10 cells.
Extended Data Figure 9 Comparison of object- versus pixel-based analysis (LLS).
a, Comparison of object-based organelle contact analysis (bright) versus pixel-based organelle co-localization analysis (pastel). Object-based analysis data are the same as LLS data in Extended Data Fig. 8a. For the pixel-based analysis, a value of 1 indicates perfect co-localization, a value of 0 indicates the organelles are never co-located. No statistical test was performed. b, Half matrix showing pixel-based co-localization analysis for all the labelled organelle pairs, including those that were not included in the object-based analysis. a, b, n = 10 cells. The white line in the centre of each box represents the median value, the upper and lower edges of the boxes represent the 75th and 25th quantile of the data, respectively, and the upper and lower fences represent the 95% confidence level of the distribution.
Supplementary information
Video 1: Point-scanning confocal, 6-colour time-lapse images
COS-7 cells expressing fusion proteins targeted to the lysosomes (cyan), mitochondria (green), ER (yellow), peroxisomes (red), and Golgi (magenta), and labelled with BODIPY 665/676 to stain LDs (blue) were imaged as described in Fig. 1a. Images were acquired every 5 s, for a total of 60 frames (5 min). In frames 21-40, the ER channel (yellow) is omitted so that the other channels can be seen more clearly. Video plays at a rate of 6 frames per s. Scale bar, 10 µm. (MP4 3009 kb)
Video 2: Time-lapse images of tracked LDs
Videos of the tracked LDs (outlined in white) shown in Fig. 2b. Confocal images were acquired every 5 s, for a total of 60 frames (5 min). Video plays at a rate of 6 frames per s. Scale bar, 5 µm. (MP4 654 kb)
Video 3: Time-lapse images of a cell treated with nocodazole.
A COS-7 cell expressing fusion proteins targeted to the lysosomes (cyan), mitochondria (green), ER (yellow), peroxisomes (red), and Golgi (magenta), and labelled with BODIPY 665/676 to stain LDs (blue), was incubated on ice for 2 min and treated with 5 μM nocodazole for 1 h. After the nocodazole treatment, confocal images were acquired every 5 s, for a total of 60 frames (5 min). In frames 21-40, the ER channel (yellow) is omitted so that the other channels can be seen more clearly. Video plays at a rate of 6 frames per s. Scale bar, 10 µm. (MP4 3046 kb)
Video 4: Lattice light sheet, 6-colour time-lapse images
Volume rendering of COS-7 cells expressing fusion proteins targeted to the peroxisomes (cyan), mitochondria (green), ER (yellow), and Golgi (red), and labelled with Texas Red dextran (lysosomes, magenta) and BODIPY 665/676 (LDs, blue), imaged as described in Fig. 3a. Image stacks of 140 planes were acquired every 9.2 seconds, for a total of 100 frames (15.3 min). Video plays at a rate of 6 frames per s. Scale bar, 10 µm. (MP4 7091 kb)
Video 5: Organelle dispersion through the cytoplasm over time
Volume rendering of 6 organelles in a COS-7 cell. Voxels are colour-coded according to the time that they were last occupied by the organelle from blue to red. (MP4 26227 kb)
Video 6: Montage of mitochondria-organelle contacts in time lapse, lattice light sheet images
Volume rendering of mitochondria (magenta) in COS-7 cells expressing fusion proteins targeting 3 other organelles, and labelled with Texas Red dextran and BODIPY 665/676. Contacts between mitochondria and other organelles are coloured green. Image stacks of 140 planes were acquired every 9.2 seconds, for a total of 100 frames (15.3 min). Scale bar, 5 µm. (MP4 3894 kb)
Video 7: Mitochondria-organelle contacts in time lapse, lattice light sheet images
Volume rendering of mitochondria in COS-7 cells expressing fusion proteins targeting 3 other organelles, and labelled with Texas Red dextran and BODIPY 665/676. Contacts between mitochondria and other organelles are coloured yellow (ER), cyan (peroxisomes) red (Golgi), magenta (lysosomes) and blue (LDs). Image stacks of 140 planes were acquired every 9.2 seconds, for a total of 100 frames (15.3 min). Scale bar, 10 µm. (MP4 4295 kb)
Video 8: Mitochondria-ER metaorganelle contacts with other organelles
Volume rendering of mitochondria-ER contacts in COS-7 cells expressing fusion proteins targeting 2 other organelles, and labelled with Texas Red dextran and BODIPY 665/676. Contacts between mitochondria-ER metaorganelle and other organelles are coloured cyan (peroxisomes) red (Golgi), magenta (lysosomes) and blue (LDs). Image stacks of 140 planes were acquired every 9.2 seconds, for a total of 100 frames (15.3 min). Scale bar, 10 µm. (MP4 2085 kb)
Rights and permissions
About this article
Cite this article
Valm, A., Cohen, S., Legant, W. et al. Applying systems-level spectral imaging and analysis to reveal the organelle interactome. Nature 546, 162–167 (2017). https://doi.org/10.1038/nature22369
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22369
This article is cited by
-
Organellar electrophysiology: DNA nanodevices charging at the unmeasured
BMC Biology (2024)
-
Orientation-invariant autoencoders learn robust representations for shape profiling of cells and organelles
Nature Communications (2024)
-
Drosophila TMEM63 and mouse TMEM63A are lysosomal mechanosensory ion channels
Nature Cell Biology (2024)
-
Dissecting organelle interdependence
Nature Cell Biology (2024)
-
Lipid droplets in pathogen infection and host immunity
Acta Pharmacologica Sinica (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.